Abstract
Alzheimer’s disease (AD) is a complex genetic disorder. To date, published data have reported conflicting results on the role of CD33 rs3865444 polymorphism in AD. The present study aimed at evaluating the effect of rs3865444 on AD in a large cohort of Greek native patients with AD. We also conducted a meta-analysis by pooling information from different studies on the same topic. Patients with AD (n = 327) and healthy controls (n = 327) were analyzed and genotyped for rs3865444. Single locus analyses were run to explore possible associations between CD33 rs3865444 polymorphism and AD. Our analysis yielded no significant interaction between AD and the CD33 rs3865444 polymorphism. The lack of interaction between the two variables persisted even after a pooled meta-analysis of 8 studies (with 13 datasets), with 4015 AD cases and 7981 controls. The overall results do not support the hypothesis that CD33 rs3865444 polymorphism increases the risk of AD. The results also suggest that the identification of functional variants in CD33 that are indisputably correlated with AD may be an important factor to investigate in future genetic screening studies.


Similar content being viewed by others
References
Aziz AL et al (2017) Difference in imaging biomarkers of neurodegeneration between early and late-onset amnestic Alzheimer’s disease. Neurobiol Aging 54:22–30. https://doi.org/10.1016/j.neurobiolaging.2017.02.010
Ballard C, Gauthier S, Corbett A, Brayne C, Aarsland D, Jones E (2011) Alzheimer’s disease lancet (London, England). Lancet 377:1019–1031. https://doi.org/10.1016/s0140-6736(10)61349-9
Banna GL et al (2017) Lactobacillus rhamnosus GG: an overview to explore the rationale of its use in cancer. Front Pharmacol 8:603. https://doi.org/10.3389/fphar.2017.00603
Bao J, Wang XJ, Mao ZF (2016) Associations between genetic variants in 19p13 and 19q13 regions and susceptibility to Alzheimer disease: a meta-analysis Medical science monitor : international medical journal of experimental and clinical research. Med Sci Monit 22:234–243
Bradshaw EM et al (2013) CD33 Alzheimer’s disease locus: altered monocyte function and amyloid biology nature. Nat Neurosci 16:848–850. https://doi.org/10.1038/nn.3435
Cacace R, Sleegers K, Van Broeckhoven C (2016) Molecular genetics of early-onset Alzheimer’s disease revisited Alzheimer’s & dementia : the journal of the Alzheimer’s Association. Alzheimers Dement 12:733–748. https://doi.org/10.1016/j.jalz.2016.01.012
Candido S et al (2019) The analysis of miRNA expression profiling datasets reveals inverse microRNA patterns in glioblastoma and Alzheimer’s disease oncology reports. Oncol Rep 42:911–922. https://doi.org/10.3892/or.2019.7215
Carmona S, Zahs K, Wu E, Dakin K, Bras J, Guerreiro R (2018) The role of TREM2 in Alzheimer’s disease and other neurodegenerative disorders the lancet neurology. Lancet Neurol 17:721–730. https://doi.org/10.1016/s1474-4422(18)30232-1
Carrasquillo MM et al (2011) Replication of EPHA1 and CD33 associations with late-onset Alzheimer's disease: a multi-centre case-control study. Mol Neurodegener 6:54. https://doi.org/10.1186/1750-1326-6-54
Carrasquillo MM et al (2014) Late-onset Alzheimer disease genetic variants in posterior cortical atrophy and posterior AD. Neurology 82:1455–1462. https://doi.org/10.1212/wnl.0000000000000335
Chung SJ, Lee JH, Kim SY, You S, Kim MJ, Lee JY, Koh J (2013) Association of GWAS top hits with late-onset Alzheimer disease in Korean population. Alzheimer Dis Assoc Disord 27:250–257. https://doi.org/10.1097/WAD.0b013e31826d7281
Cuyvers E, Sleegers K (2016) Genetic variations underlying Alzheimer’s disease: evidence from genome-wide association studies and beyond. Lancet Neurol 15:857–868. https://doi.org/10.1016/S1474-4422(16)00127-7
Dabke K, Hendrick G, Devkota S (2019) The gut microbiome and metabolic syndrome. J Clin Invest 129:4050–4057. https://doi.org/10.1172/jci129194
Dardiotis E et al (2015) Effect of angiotensin-converting enzyme tag single nucleotide polymorphisms on the outcome of patients with traumatic brain injury. Pharmacogenet Genomics 25:485–490. https://doi.org/10.1097/fpc.0000000000000161
Dardiotis E et al (2018a) The role of microRNAs in patients with amyotrophic lateral sclerosis, J Mol Neurosci : MN. 66:617–628. https://doi.org/10.1007/s12031-018-1204-1
Dardiotis E et al (2018b) Genetic polymorphisms in amyotrophic lateral sclerosis: evidence for implication in detoxification pathways of environmental toxicants. Environ Int 116:122–135. https://doi.org/10.1016/j.envint.2018.04.008
Dardiotis E et al (2018c) H. pylori and Parkinson’s disease: meta-analyses including clinical severity. Clin Neurol Neurosurg 175:16–24. https://doi.org/10.1016/j.clineuro.2018.09.039
Dardiotis E, Siokas V, Moza S, Kosmidis MH, Vogiatzi C, Aloizou AM, Geronikola N, Ntanasi E, Zalonis I, Yannakoulia M, Scarmeas N, Hadjigeorgiou GM (2019) Pesticide exposure and cognitive function: results from the Hellenic Longitudinal Investigation of Aging and Diet (HELIAD). Environ Res 177:108632. https://doi.org/10.1016/j.envres.2019.108632
Deng YL et al (2012) The prevalence of CD33 and MS4A6A variant in Chinese Han population with Alzheimer’s disease. Hum Genet 131:1245–1249. https://doi.org/10.1007/s00439-012-1154-6
DerSimonian R, Laird N (1986) Meta-analysis in clinical trials. Control Clin Trials 7:177–188
Dos Santos LR et al (2017) Validating GWAS variants from microglial genes implicated in Alzheimer’s disease. J Mol Neurosci: MN 62:215–221. https://doi.org/10.1007/s12031-017-0928-7
Falzone L et al (2016) Computational identification of microRNAs associated to both epithelial to mesenchymal transition and NGAL/MMP-9 pathways in bladder cancer. Oncotarget 7:72758–72766. https://doi.org/10.18632/oncotarget.11805
Falzone L, Scola L, Zanghi A, Biondi A, Di Cataldo A, Libra M, Candido S (2018) Integrated analysis of colorectal cancer microRNA datasets: identification of microRNAs associated with tumor development. Aging 10:1000–1014. https://doi.org/10.18632/aging.101444
Falzone L et al. (2019a) Identification of novel microRNAs and their diagnostic and prognostic significance in oral cancer. Cancers 11(5). https://doi.org/10.3390/cancers11050610
Falzone L et al (2019b) Prognostic significance of deregulated microRNAs in uveal melanomas. Mol Med Rep 19:2599–2610. https://doi.org/10.3892/mmr.2019.9949
Griciuc A et al (2013) Alzheimer’s disease risk gene CD33 inhibits microglial uptake of amyloid beta. Neuron 78:631–643. https://doi.org/10.1016/j.neuron.2013.04.014
Gubandru M et al (2013) Alzheimer’s disease treated patients showed different patterns for oxidative stress and inflammation markers food and chemical toxicology: an international journal published for the British industrial biological research association. Food Chem Toxicol 61:209–214. https://doi.org/10.1016/j.fct.2013.07.013
Hafsi S et al (2016) Correlation between the overexpression of Yin Yang 1 and the expression levels of miRNAs in Burkitt’s lymphoma: a computational study. Oncol Lett 11:1021–1025. https://doi.org/10.3892/ol.2015.4031
Hollingworth P et al (2011) Common variants at ABCA7, MS4A6A/MS4A4E, EPHA1, CD33 and CD2AP are associated with Alzheimer’s disease. Nature Genet 43:429–435. https://doi.org/10.1038/ng.803
Jafarian Z, Saliminejad K, Kamali K, Ohadi M, Kowsari A, Nasehi L, Khorram Khorshid HR (2018) Association of glutathione S-transferases M1, P1 and T1 variations and risk of late-onset. Alzheimers Dis Neurol Res 40:41–44. https://doi.org/10.1080/01616412.2017.1390902
Jiang T, Yu JT, Hu N, Tan MS, Zhu XC, Tan L (2014) CD33 in Alzheimer’s disease. Mol Neurobiol 49:529–535. https://doi.org/10.1007/s12035-013-8536-1
Jiang C, Li G, Huang P, Liu Z, Zhao B (2017) The gut microbiota and Alzheimer’s disease. J Alzheimers Dis: JAD 58:1–15. https://doi.org/10.3233/jad-161141
Jiang YT, Li HY, Cao XP, Tan L (2018) Meta-analysis of the association between CD33 and Alzheimer’s disease. Ann Translat Med 6:169. https://doi.org/10.21037/atm.2018.04.21
Jiao B et al (2015) Polygenic analysis of late-onset Alzheimer’s disease from Mainland China. PloS One 10:e0144898. https://doi.org/10.1371/journal.pone.0144898
Karch CM, Goate AM (2015) Alzheimer’s disease risk genes and mechanisms of disease pathogenesis. Biol Psychiatry 77:43–51. https://doi.org/10.1016/j.biopsych.2014.05.006
Lambert JC et al (2013) Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet 45:1452–1458. https://doi.org/10.1038/ng.2802
Li X, Shen N, Zhang S, Liu J, Jiang Q, Liao M, Feng R, Zhang L, Wang G, Ma G, Zhou H, Chen Z, Jiang Y, Zhao B, Li K, Liu G (2015) CD33 rs3865444 polymorphism contributes to Alzheimer’s disease susceptibility in Chinese, European, and North American populations. Mol Neurobiol 52:414–421. https://doi.org/10.1007/s12035-014-8880-9
Liberati A et al. (2009) The PRISMA Statement for Reporting Systematic Reviews and Meta-Analyses of Studies That Evaluate Health Care Interventions: Explanation and Elaboration. PLoS Med 6(7):e1000100. https://doi.org/10.1016/j.jclinepi.2009.06.006
Lin P-I, Vance JM, Pericak-Vance MA, Martin ER (2007) No gene is an island: the flip-flop phenomenon. Am J Hum Genet 80:531–538
Liu C-C, Liu C-C, Kanekiyo T, Xu H, Bu G (2013) Apolipoprotein E and Alzheimer disease: risk, mechanisms and therapy. Nat Rev Neurol 9:106–118. https://doi.org/10.1038/nrneurol.2012.263
Liu S, Wu Y, Liu X, Zhou J, Wang Z, He Z, Huang Z (2017) Lack of association between MTHFR A1298C variant and Alzheimer’s disease: evidence from a systematic review and cumulative meta-analysis. Neurol Res 39:426–434. https://doi.org/10.1080/01616412.2017.1297340
Logue MW et al (2011) A comprehensive genetic association study of Alzheimer disease in African Americans. Arch Neurol 68:1569–1579. https://doi.org/10.1001/archneurol.2011.646
Ma J et al (2014) Association study of TREM2 polymorphism rs75932628 with late-onset Alzheimer’s disease in Chinese Han population. Neurol Res 36:894–896. https://doi.org/10.1179/1743132814y.0000000376
Malik M et al (2015) Genetics of CD33 in Alzheimer’s disease and acute myeloid leukemia. Hum Mol Genet 24:3557–3570. https://doi.org/10.1093/hmg/ddv092
Mantel N, Haenszel W (1959) Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst 22:719–748
Mao YF, Guo ZY, Pu JL, Chen YX, Zhang BR (2015) Association of CD33 and MS4A cluster variants with Alzheimer’s disease in East Asian populations. Neurosci Lett 609:235–239. https://doi.org/10.1016/j.neulet.2015.10.007
McKhann G, Drachman D, Folstein M, Katzman R, Price D, Stadlan EM (1984) Clinical diagnosis of Alzheimer’s disease: report of the NINCDS-ADRDA work group under the auspices of department of health and human services task force on Alzheimer’s disease. Neurol 34:939–944. https://doi.org/10.1212/wnl.34.7.939
Miyashita A, Koike A, Jun G, Wang LS, Takahashi S, Matsubara E, Kawarabayashi T, Shoji M, Tomita N, Arai H, Asada T, Harigaya Y, Ikeda M, Amari M, Hanyu H, Higuchi S, Ikeuchi T, Nishizawa M, Suga M, Kawase Y, Akatsu H, Kosaka K, Yamamoto T, Imagawa M, Hamaguchi T, Yamada M, Morihara T, Takeda M, Takao T, Nakata K, Fujisawa Y, Sasaki K, Watanabe K, Nakashima K, Urakami K, Ooya T, Takahashi M, Yuzuriha T, Serikawa K, Yoshimoto S, Nakagawa R, Kim JW, Ki CS, Won HH, Na DL, Seo SW, Mook-Jung I, Alzheimer Disease Genetics Consortium, St George-Hyslop P, Mayeux R, Haines JL, Pericak-Vance MA, Yoshida M, Nishida N, Tokunaga K, Yamamoto K, Tsuji S, Kanazawa I, Ihara Y, Schellenberg GD, Farrer LA, Kuwano R (2013) SORL1 is genetically associated with late-onset Alzheimer’s disease in Japanese, Koreans and Caucasians. PloS One 8:e58618. https://doi.org/10.1371/journal.pone.0058618
Moreno DJ, Ruiz S, Rios A, Lopera F, Ostos H, Via M, Bedoya G (2017) Association of GWAS top genes with late-onset Alzheimer’s disease in Colombian population. Am J Alzheimers Dis Other Demen 32:27–35. https://doi.org/10.1177/1533317516679303
Moreno-Grau S et al (2018) Genome-wide significant risk factors on chromosome 19 and the APOE locus. Oncotarget 9:24590–24600. https://doi.org/10.18632/oncotarget.25083
Naj AC et al (2011) Common variants at MS4A4/MS4A6E, CD2AP, CD33 and EPHA1 are associated with late-onset Alzheimer’s disease. Nat Genet 43:436–441. https://doi.org/10.1038/ng.801
Omoumi A, Fok A, Greenwood T, Sadovnick AD, Feldman HH, Hsiung GY (2014) Evaluation of late-onset Alzheimer disease genetic susceptibility risks in a Canadian population. Neurobiol Aging 35(936):e935–e912. https://doi.org/10.1016/j.neurobiolaging.2013.09.025
Panegyres PK, Chen H-Y (2013) Differences between early and late onset Alzheimer’s disease. Am J Neurodegener Dis 2:300–306
Panoutsopoulou K, Tachmazidou I, Zeggini E (2013) In search of low-frequency and rare variants affecting complex traits. Hum Mol Genet 22:R16–R21. https://doi.org/10.1093/hmg/ddt376
Razgonova MP et al (2019) Panax ginseng components and the pathogenesis of Alzheimer’s disease (review). Mol Med Rep 19:2975–2998. https://doi.org/10.3892/mmr.2019.9972
Reitz C, Brayne C, Mayeux R (2011) Epidemiology of Alzheimer disease. Nat Rev Neurol 7:137–152. https://doi.org/10.1038/nrneurol.2011.2
Ruszkiewicz JA, Tinkov AA, Skalny AV, Siokas V, Dardiotis E, Tsatsakis A, Bowman AB, da Rocha JBT, Aschner M (2019) Brain diseases in changing climate. Environ Res 177:108637. https://doi.org/10.1016/j.envres.2019.108637
Scheltens P, Blennow K, Breteler MMB, de Strooper B, Frisoni GB, Salloway S, Van der Flier WM (2016) Alzheimer’s disease. Lancet 388:505–517. https://doi.org/10.1016/S0140-6736(15)01124-1
Serrano-Pozo A, Frosch MP, Masliah E, Hyman BT (2011) Neuropathological alterations in Alzheimer disease. Cold Spring Harb Perspect Med 1:a006189. https://doi.org/10.1101/cshperspect.a006189
Siokas V et al (2017) The role of TOR1A polymorphisms in dystonia: a systematic review and meta-analysis. PloS One 12:e0169934. https://doi.org/10.1371/journal.pone.0169934
Siokas V et al (2019a) BDNF rs6265 (Val66Met) polymorphism as a risk factor for blepharospasm. Neuromol Med 21:68–74. https://doi.org/10.1007/s12017-018-8519-5
Siokas V et al (2019b) Lack of association of the rs11655081 ARSG gene with blepharospasm. J Mol Neurosci 67:472–476. https://doi.org/10.1007/s12031-018-1255-3
Skol AD, Scott LJ, Abecasis GR, Boehnke M (2006) Joint analysis is more efficient than replication-based analysis for two-stage genome-wide association studies. Nat Genet 38:209–213. https://doi.org/10.1038/ng1706
Sole X, Guino E, Valls J, Iniesta R, Moreno V (2006) SNPStats: a web tool for the analysis of association studies bioinformatics (Oxford, England). Bioinformatics 22:1928–1929. https://doi.org/10.1093/bioinformatics/btl268
Stamati P, Siokas V, Aloizou AM, Karampinis E, Arseniou S, Rakitskii VN, Tsatsakis A, Spandidos DA, Gozes I, Mitsias PD, Bogdanos DP, Hadjigeorgiou GM, Dardiotis E (2019) Does SCFD1 rs10139154 polymorphism decrease Alzheimer’s disease risk? J Mol Neurosci 69:343–350. https://doi.org/10.1007/s12031-019-01363-3
Sun Q, Xie N, Tang B, Li R, Shen Y (2017) Alzheimer’s disease: from genetic variants to the distinct pathological mechanisms. Front Mol Neurosci 10:319. https://doi.org/10.3389/fnmol.2017.00319
Tan L et al (2013) Association of GWAS-linked loci with late-onset Alzheimer’s disease in a northern Han Chinese population. Alzheimers Demen: JAlzheimers Assoc 9:546–553. https://doi.org/10.1016/j.jalz.2012.08.007
Theuns J et al (2014) Global investigation and meta-analysis of the C9orf72 (G4C2)n repeat in Parkinson disease. Neurol 83:1906–1913. https://doi.org/10.1212/wnl.0000000000001012
Van Cauwenberghe C, Van Broeckhoven C, Sleegers K (2016) The genetic landscape of Alzheimer disease: clinical implications and perspectives. Genet Med: Official J Am College Med Genet 18:421–430. https://doi.org/10.1038/gim.2015.117
Vivarelli S et al (2019) Gut microbiota and cancer: from pathogenesis to therapy. Cancers 11. https://doi.org/10.3390/cancers11010038
Walker DG, Whetzel AM, Serrano G, Sue LI, Beach TG, Lue LF (2015) Association of CD33 polymorphism rs3865444 with Alzheimer’s disease pathology and CD33 expression in human cerebral cortex. Neurobiol Aging 36:571–582. https://doi.org/10.1016/j.neurobiolaging.2014.09.023
Wattmo C, Wallin ÅK (2017) Early- versus late-onset Alzheimer disease: long-term functional outcomes, nursing home placement, and risk factors for rate of progression. Dement Geriatr Cogn Dis Extra 7:172–187
Funding
This study was supported in part by a research grant from the Research Committee of the University of Thessaly, Greece (code: 5287). MA was supported in part by the National Institute of Environmental Health Sciences (R01ES07331, R01ES10563, and R01ES020852).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical Approval
The research protocol was approved by the Ethics Committee of the University General Hospital of Larissa.
Informed Consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary file 1
The complete search algorithm. (DOCX 12 kb)
Supplementary file 2
PRISMA 2009 Checklist. (DOCX 56 kb)
Supplementary file 3
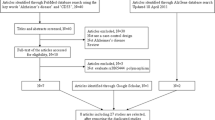
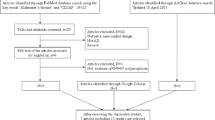
Flow chart presenting the selection of eligible studies (DOC 63.5 KB)
Rights and permissions
About this article
Cite this article
Siokas, V., Aslanidou, P., Aloizou, AM. et al. Does the CD33 rs3865444 Polymorphism Confer Susceptibility to Alzheimer’s Disease?. J Mol Neurosci 70, 851–860 (2020). https://doi.org/10.1007/s12031-020-01507-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-020-01507-w