Abstract
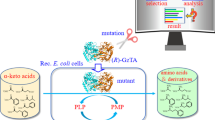
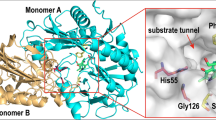
ω-Transaminase (ω-TA) is a promising biocatalyst for the synthesis of chiral amines. In this study, a ω-TA derived from Vitreoscilla stercoraria DSM 513 (VsTA) was heterologous expressed in recombinant E. coli cells and applied to reduce 4′-(trifluoromethyl)acetophenone (TAP) to (S)-1-[4-(trifluoromethyl)phenyl]ethylamine ((S)-TPE), a pharmaceutical intermediate of chiral amine. Aimed to a more efficient synthesis of (S)-TPE, VsTA was further engineered via a semi-rational strategy. Compared to wild-type VsTA, the obtained R411A variant exhibited 2.39 times higher activity towards TAP and enhanced catalytic activities towards other prochiral aromatic ketones. Additionally, better thermal stability for R411A variant was observed with 25.4% and 16.3% increase in half-life at 30 °C and 40 °C, respectively. Structure-guided analysis revealed that the activity improvement of R411A variant was attributed to the introduction of residue A411, which is responsible for the increase in the hydrophobicity of substrate tunnel and the alleviation of steric hindrance, thereby facilitating the accessibility of hydrophobic substrate TAP to the active center of VsTA. This study provides an efficient strategy for the engineering of ω-TA based on semi-rational approach and has the potential for the molecular modification of other biocatalysts.
Graphical Abstract









Similar content being viewed by others
Data Availability
All data generated or analyzed during this study are included in this published article and its supplementary information file.
References
Kelly, S. A., Pohle, S., Wharry, S., Mix, S., Allen, C. C. R., Moody, T. S., & Gilmore, B. F. (2018). Application of ω-transaminases in the pharmaceutical industry. Chemical Reviews, 118, 349–367. https://doi.org/10.1021/acs.chemrev.7b00437
Savile, C. K., Janey, J. M., Mundorff, E. C., Moore, J. C., Tam, S., Jarvis, W. R., Colbeck, J. C., Krebber, A., Fleitz, F. J., Brands, J., Devine, P. N., Huisman, G. W., & Hughes, G. J. (2010). Biocatalytic asymmetric synthesis of chiral amines from ketones applied to sitagliptin manufacture. Science, 329, 305–309. https://doi.org/10.1126/science.1188934
Novick, S. J., Dellas, N., Garcia, R., Ching, C., Bautista, A., Homan, D., Alvizo, O., Entwistle, D., Kleinbeck, F., Schlama, T., & Ruch, T. (2021). Engineering an amine transaminase for the efficient production of a chiral sacubitril precursor. ACS Catalysis, 11, 3762–3770. https://doi.org/10.1021/acscatal.0c05450
Schober, M., MacDermaid, C., Ollis, A. A., Chang, S., Khan, D., Hosford, J., Latham, J., Ihnken, L. A. F., Brown, M. J. B., Fuerst, D., Sanganee, M. J., & Roiban, G. D. (2019). Chiral synthesis of LSD1 inhibitor GSK2879552 enabled by directed evolution of an imine reductase. Nature Catalysis, 2, 909–915. https://doi.org/10.1038/s41929-019-0341-4
Cabré, A., Verdaguer, X., & Riera, A. (2022). Recent advances in the enantioselective synthesis of chiral amines via transition metal-catalyzed asymmetric hydrogenation. Chemical Reviews, 122, 269–339. https://doi.org/10.1021/acs.chemrev.1c00496
Kohls, H., Steffen-Munsberg, F., & Höhne, M. (2014). Recent achievements in developing the biocatalytic toolbox for chiral amine synthesis. Current Opinion in Chemical Biology, 19, 180–192. https://doi.org/10.1016/j.cbpa.2014.02.021
Patil, M. D., Grogan, G., Bommarius, A., & Yun, H. (2018). Oxidoreductase-catalyzed synthesis of chiral amines. ACS Catalysis, 8, 10985–11015. https://doi.org/10.1021/acscatal.8b02924
Fuchs, M., Farnberger, J. E., & Kroutil, W. (2015). The industrial age of biocatalytic transamination. European Journal of Organic Chemistry, 2015, 6965–6982. https://doi.org/10.1002/ejoc.201500852
Ferrandi, E. E., & Monti, D. (2018). Amine transaminases in chiral amines synthesis: Recent advances and challenges. World Journal of Microbiology and Biotechnology, 34, 13. https://doi.org/10.1007/s11274-017-2395-2
Gomm, A., & O’Reilly, E. (2018). Transaminases for chiral amine synthesis. Current Opinion in Chemical Biology, 43, 106–112. https://doi.org/10.1016/j.cbpa.2017.12.007
Ghislieri, D., & Turner, N. J. (2014). Biocatalytic approaches to the synthesis of enantiomerically pure chiral amines. Topics in Catalysis, 57, 284–300. https://doi.org/10.1007/s11244-013-0184-1
Guo, F., & Berglund, P. (2017). Transaminase biocatalysis: Optimization and application. Green Chemistry, 19, 333–360. https://doi.org/10.1039/c6gc02328b
Li, F., Du, Y., Liang, Y., Wei, Y., Zheng, Y., & Yu, H. (2023). Redesigning an (R)-selective transaminase for the efficient synthesis of pharmaceutical N-heterocyclic amines. ACS Catalysis, 13, 422–432. https://doi.org/10.1021/acscatal.2c05177
Slabu, I., Galman, J. L., Lloyd, R. C., & Turner, N. J. (2017). Discovery, engineering, and synthetic application of transaminase biocatalysts. ACS Catalysis, 7, 8263–8284. https://doi.org/10.1021/acscatal.7b02686
Qu, G., Li, A., Acevedo-Rocha, C. G., Sun, Z., & Reetz, M. T. (2020). The crucial role of methodology development in directed evolution of selective enzymes. Angewandte Chemie- International Edition, 59, 13204–13231. https://doi.org/10.1002/anie.201901491
Morley, K. L., & Kazlauskas, R. J. (2005). Improving enzyme properties: When are closer mutations better? Trends in Biotechnology, 23, 231–237. https://doi.org/10.1016/j.tibtech.2005.03.005
Shin, J. S., & Kim, B. G. (2002). Exploring the active site of amine:Pyruvate aminotransferase on the basis of the substrate structure-reactivity relationship: How the enzyme controls substrate specificity and stereoselectivity. Journal of Organic Chemistry, 67, 2848–2853. https://doi.org/10.1021/jo016115i
Meng, Q., Ramírez-Palacios, C., Capra, N., Hooghwinkel, M. E., Thallmair, S., Rozeboom, H. J., Thunnissen, A. M. W. H., Wijma, H. J., Marrink, S. J., & Janssen, D. B. (2021). Computational redesign of an ω-transaminase from Pseudomonas jessenii for asymmetric synthesis of enantiopure bulky amines. ACS Catalysis, 11, 10733–10747. https://doi.org/10.1021/acscatal.1c02053
Nobili, A., Steffen-Munsberg, F., Kohls, H., Trentin, I., Schulzke, C., Höhne, M., & Bornscheuer, U. T. (2015). Engineering the active site of the amine transaminase from Vibrio fluvialis for the asymmetric synthesis of aryl-alkyl amines and amino alcohols. ChemCatChem, 7, 757–760. https://doi.org/10.1002/cctc.201403010
Land, H., Ruggieri, F., Szekrenyi, A., Fessner, W. D., & Berglund, P. (2020). Engineering the active site of an (S)-selective amine transaminase for acceptance of doubly bulky primary amines. Advanced Synthesis and Catalysis, 362, 812–821. https://doi.org/10.1002/adsc.201901252
Xie, Y., Xu, F., Yang, L., Liu, H., Xu, X., Wang, H., & Wei, D. (2021). Engineering the large pocket of an (S)-selective transaminase for asymmetric synthesis of (S)-1-amino-1-phenylpropane. Catalysis Science and Technology, 11, 2461–2470. https://doi.org/10.1039/d0cy02426k
Ma, Y., Jiao, X., Wang, Z., Mu, H., Sun, K., Li, X., Zhao, T., Liu, X., & Zhang, N. (2022). Engineering a transaminase for the efficient synthesis of a key intermediate for rimegepant. Organic Process Research and Development, 26, 1971–1977. https://doi.org/10.1021/acs.oprd.1c00376
Jia, D. X., Peng, C., Li, J. L., Wang, F., Liu, Z. Q., & Zheng, Y. G. (2021). Redesign of (R)-omega-transaminase and its application for synthesizing amino acids with bulky side chain. Applied Biochemistry and Biotechnology, 193, 3624–3640. https://doi.org/10.1007/s12010-021-03616-7
Wang, Y., Feng, J., Dong, W., Chen, X., Yao, P., Wu, Q., & Zhu, D. (2021). Improving catalytic activity and reversing enantio-specificity of ω-transaminase by semi-rational engineering en route to chiral bulky β-amino esters. ChemCatChem, 13, 3396–3400. https://doi.org/10.1002/cctc.202100503
Han, R., Cao, X., Fang, H., Zhou, J., & Ni, Y. (2021). Structure-based engineering of ω-transaminase for enhanced catalytic efficiency toward (R)-(+)-1-(1-naphthyl)ethylamine synthesis. Molecular Catalysis, 502, 111368. https://doi.org/10.1016/j.mcat.2020.111368
Cheng, F., Chen, X. L., Li, M. Y., Zhang, X. J., Jia, D. X., Wang, Y. J., Liu, Z. Q., & Zheng, Y. G. (2020). Creation of a robust and R-selective ω-amine transaminase for the asymmetric synthesis of sitagliptin intermediate on a kilogram scale. Enzyme and Microbial Technology, 141, 109655. https://doi.org/10.1016/j.enzmictec.2020.109655
Yang, X., Wu, T., Phipps, R. J., & Toste, F. D. (2015). Advances in catalytic enantioselective fluorination, mono-, di-, and trifluoromethylation, and trifluoromethylthiolation reactions. Chemical Reviews, 115, 826–870. https://doi.org/10.1021/cr500277b
Zhou, Y., Wang, J., Gu, Z., Wang, S., Zhu, W., Acenã, J. L., Soloshonok, V. A., Izawa, K., & Liu, H. (2016). Next generation of fluorine-containing pharmaceuticals, compounds currently in phase II-III clinical trials of major pharmaceutical companies: New structural trends and therapeutic areas. Chemical Reviews, 116, 422–518. https://doi.org/10.1021/acs.chemrev.5b00392
Cardiac sarcomere inhibitors. (2022). US Patent No 11414424 B2. Available from: https://ppubs.uspto.gov/dirsearch-public/print/downloadPdf/11414424. Accessed 18 Feb 2024
Indole carboxamide derivative and pharmaceutical composition containing same. (2023). US Patent No 20230000821 A1. Available from: https://ppubs.uspto.gov/dirsearch-public/print/downloadPdf/20230000821. Accessed 18 Feb 2024
Waterhouse, A., Bertoni, M., Bienert, S., Studer, G., Tauriello, G., Gumienny, R., Heer, F. T., Beer, T. A. P., Rempfer, C., Bordoli, L., Lepore, R., & Schwede, T. (2018). SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Research, 46, W296–W303. https://doi.org/10.1093/nar/gky427
Laskowski, R. A., MacArthur, M. W., Moss, D. S., & Thornton, J. M. (1993). PROCHECK: A program to check the stereochemical quality of protein structures. Journal of Applied Crystallography, 26, 283–291. https://doi.org/10.1107/s0021889892009944
Colovos, C., & Yeates, T. O. (1993). Verification of protein structures: Patterns of nonbonded atomic interactions. Protein Science, 2, 1511–1519. https://doi.org/10.1002/pro.5560020916
Forli, S., Huey, R., Pique, M. E., Sanner, M. F., Goodsell, D. S., & Olson, A. J. (2016). Computational protein-ligand docking and virtual drug screening with the AutoDock suite. Nature Protocols, 11, 905–919. https://doi.org/10.1038/nprot.2016.051
Morrison, K. L., & Weiss, G. A. (2001). Combinatorial alanine-scanning. Current Opinion in Chemical Biology, 5, 302–307. https://doi.org/10.1016/S1367-5931(00)00206-4
Chovancova, E., Pavelka, A., Benes, P., Strnad, O., Brezovsky, J., Kozlikova, B., Gora, A., Sustr, V., Klvana, M., Medek, P., Biedermannova, L., Sochor, J., & Damborsky, J. (2012). CAVER 30: A tool for the analysis of transport pathways in dynamic protein structures. PLoS Computational Biology, 8, e1002708. https://doi.org/10.1371/journal.pcbi.1002708
Xiang, C., Ao, Y. F., Höhne, M., & Bornscheuer, U. T. (2022). Shifting the pH optima of (R)-selective transaminases by protein engineering. International Journal of Molecular Sciences, 23, 15347. https://doi.org/10.3390/ijms232315347
Kelefiotis-Stratidakis, P., Tyrikos-Ergas, T., & Pavlidis, I. V. (2019). The challenge of using isopropylamine as an amine donor in transaminase catalysed reactions. Organic and Biomolecular Chemistry, 17, 1634–1642. https://doi.org/10.1039/c8ob02342e
Contente, M. L., Planchestainer, M., Molinari, F., & Paradisi, F. (2016). Stereoelectronic effects in the reaction of aromatic substrates catalysed by: Halomonas elongata transaminase and its mutants. Organic and Biomolecular Chemistry, 14, 9306–9311. https://doi.org/10.1039/c6ob01629d
Rodrigues, C. J. C., Ferrer, M., & de Carvalho, C. C. C. R. (2021). ω-Transaminase-mediated asymmetric synthesis of (S)-1-(4-trifluoromethylphenyl)ethylamine. Catalysts, 11, 307. https://doi.org/10.3390/catal11030307
López-Iglesias, M., González-Martínez, D., Rodríguez-Mata, M., Gotor, V., Busto, E., Kroutil, W., & Gotor-Fernández, V. (2017). Asymmetric biocatalytic synthesis of fluorinated pyridines through transesterification or transamination: Computational insights into the reactivity of transaminases. Advanced Synthesis & Catalysis, 359, 279–291. https://doi.org/10.1002/adsc.201600835
Funding
This work was financially supported by grants from the National Key Research and Development Program of China (2022YFA0911804), the National Natural Science Foundation of China (No. 21676250), and the Zhejiang Province Natural Science Foundation of China (LY22B060010).
Author information
Authors and Affiliations
Contributions
ZD: visualization, writing—original draft, data curation, conceptualization, writing—review and editing. YW: data curation, investigation. DS: data curation. XS: writing—review and editing. PW: conceptualization, resources, project administration, funding acquisition, supervision.
Corresponding author
Ethics declarations
Ethics Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for Publication
All authors have reviewed the manuscript and have agreed to submit it to the journal. We have taken full account of the intellectual property protection issues associated with this work, and there are no barriers to publication in terms of intellectual property rights.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Zhi-Wen Duan and Yao-Wu Wang are co-first authors.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Duan, ZW., Wang, YW., Shen, DD. et al. Engineered the Active Site of ω-Transaminase for Enhanced Asymmetric Synthesis Towards (S)-1-[4-(Trifluoromethyl)phenyl]ethylamine. Appl Biochem Biotechnol (2024). https://doi.org/10.1007/s12010-024-04886-7
Accepted:
Published:
DOI: https://doi.org/10.1007/s12010-024-04886-7