Abstract
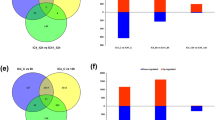
Sweetpotato (Ipomoea batatas [L.] Lam) is an important starch crop that ensures food and nutrition security in the era of climate change. Sweetpotato is tolerant to environmental stresses such as drought, high temperature, and high salt, and therefore, is well adapted to marginal lands; however, it is relatively vulnerable to flooding stress, which severely reduces its yield and commercial value. To understand the flooding stress response of sweetpotato, we performed comparative transcriptome analysis of the leaves of two sweetpotato cultivars with contrasting flooding stress tolerance levels: Yeonjami (YJM; flooding tolerant) and Jeonmi (JM; flooding sensitive). Both cultivars were partially submerged in water for 0, 0.5, and 3 days. RNA-seq data of both cultivars revealed 14,229 differentially expressed genes (DEGs), which were categorized into seven clusters and six groups, based on the expression pattern of co-expressed DEGs and expression duration of DEGs, respectively. Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses showed that DEGs of distinguishing functions between the two cultivars were involved in plant hormone signaling, carbohydrate transport, and mitogen-activated protein kinase (MAPK) signaling. Based on these results, we predict that YJM promotes adventitious growth, whereas JM exhibits shoot elongation under flooding stress. The expression levels of several key candidate genes involved in flooding tolerance correlated well with the comparative transcriptomics data. Overall, this study provides further insights into the molecular mechanism of flooding stress response in sweetpotato, and reveals candidate genes that could be used for developing new flooding tolerant sweetpotato cultivars.





Similar content being viewed by others
Abbreviations
- ABA:
-
Abscisic acid
- ACC:
-
1-Aminocyclopropane-1-carboxylic acid
- ADH:
-
Alcohol dehydrogenase
- ARF:
-
ADP-ribosylation factor
- CTR1:
-
CONSTITUTIVE TRIPLE RESPONSE 1
- DEG:
-
Differentially expressed gene
- EIN3:
-
Ethylene-insensitive protein 3
- ERF:
-
Ethylene-responsive transcription factor
- ETR:
-
Ethylene receptor
- FDR:
-
False discovery rate
- GA:
-
Gibberellic acid
- GO:
-
Gene Ontology
- JM:
-
Jeonmi
- KO:
-
KEGG Orthology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- LDH:
-
Lactate dehydrogenase
- MAPK:
-
Mitogen-activated protein kinase
- nr:
-
Non-redundant
- NumReads:
-
Number of reads per transcript
- qRT-PCR:
-
Quantitative real-time PCR
- RNA-seq:
-
RNA sequencing
- TOM:
-
Topological overlap matrix
- YJM:
-
Yeonjami
References
Azuma T, Hirano T, Deki Y, Uchida N, Yasuda T, Yamaguchi T (1995) Involvement of the decrease in levels of abscisic acid in the internodal elongation of submerged floating rice. J Plant Physiol 146:323–328
Banga M, Slaa EJ, Blom CW, Voesenek LA (1996) Ethylene biosynthesis and accumulation under drained and submerged conditions (a comparative study of two Rumex species). Plant Physiol 112:229–237
Blom CWPM, Voesenek LACJ (1996) Flooding: the survival strategies of plants. Trends Ecol Evol 11:290–295
Bovell-Benjamin AC (2007) Sweet potato: a review of its past, present, and future role in human nutrition. Adv Nutr 52:1–59
Burrows WJ, Carr DJ (1969) Effects of flooding the root system of sunflower plants on the cytokin in content in the xylem sap. Physiol Plant 22:1105–1112
Bushnell B (2014) BBMap: a fast, accurate, splice-aware aligner (No. LBNL-7065E). Lawrence Berkeley National Lab. (LBNL), Berkeley
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinform 10:421
Chen H, Qualls RG, Blank RR (2005) Effect of soil flooding on photosynthesis, carbohydrate partitioning and nutrient uptake in the invasive exotic Lepidium latifolium. Aquat Bot 82:250–268
Chen SP, Lin W, Chen X, Huang YH, Chang SC, Lo HS, Lu HH, Yeh KW (2016) Sweet potato NAC transcription factor, IbNAC1, up regulates sporamin gene expression by binding the SWRE motif against mechanical wounding and herbivore attack. Plant J 86:234–248
Christianson JA, Llewellyn DJ, Dennis ES, Wilson IW (2010) Global gene expression responses to waterlogging in roots and leaves of cotton (Gossypium hirsutum L.). Plant Cell Physiol 51:21–37
Cruz-Mendívil A, López-Valenzuela JA, Calderón-Vázquez CL, Vega-García MO, Reyes-Moreno C, Valdez-Ortiz A (2015) Transcriptional changes associated with chilling tolerance and susceptibility in ‘Micro-Tom’ tomato fruit using RNA-Seq. Biol Technol 99:141–151
Drew MC, Jackson MB, Giffard SC (1979) Ethylene-promoted adventitious rooting and development of cortical air spaces (aerenchyma) in roots may be adaptive responses to flooding in Zea mays L. Planta 147:83–88
Eguchi T, Yoshida S (2007) Effects of gas exchange inhibition and hypoxia on tuberous root morphogenesis in sweetpotato (Ipomoea batatas (L.) Lam.). Environ Control Biol 45:103–111
Ellis MH, Dennis ES, Peacock WJ (1999) Arabidopsis roots and shoots have different mechanisms for hypoxic stress tolerance. Plant Physiol 119:57–64
Else MA, Davies WJ, Malone M, Jackson MB (1995) A negative hydraulic message from oxygen-deficient roots of tomato plants? (influence of soil flooding on leaf water potential, leaf expansion, and synchrony between stomatal conductance and root hydraulic conductivity). Plant Physiol 109:1017–1024
FAO (2018). www.fao.org/news/story/en/item/1106977. Accessed 23 Nov 2020
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, Heger A, Hetherington K, Holm L, Mistry J, Sonnhammer EL, Tate J, Punta M (2014) Pfam: the protein families database. Nucleic Acids Res 42(D1):D222–D230
Fukao T, Bailey-Serres J (2004) Plant responses to hypoxia–is survival a balancing act? Trends Plant Sci 9:449–456
Gibbs J, Greenway H (2003) Mechanisms of anoxia tolerance in plants. I. Growth, survival and anaerobic catabolism. Funct Plant Biol 30:1–47
Hoffmann-Benning S, Kende H (1992) On the role of abscisic acid and gibberellin in the regulation of growth in rice. Plant Physiol 99:1156–1161
Hunter S, Apweiler R, Attwood TK, Bairoch A, Bateman A, Binns D, Bork P, Das U, Daugherty L, Duquenne L, Finn RD, Gough J, Haft D, Hulo N, Kahn D, Kelly E, Laugraud A, Letunic I, Lonsdale D, Lopez R, Madera M, Maslen J, McAnulla C, McDowall J, Mistry J, Mitchell A, Mulder N, Natale D, Orengo C, Quinn AF, Selengut JD, Sigrist CJA, Thimma M, Thomas PD, Valentin F, Wilson D, Wu CH, Yeats C (2009) InterPro: the integrative protein signature database. Nucleic Acids Res 37(Suppl 1):D211–D215
Hwang SY, VanToai TT (1991) Abscisic acid induces anaerobiosis tolerance in corn. Plant Physiol 97:593–597
Ismond KP, Dolferus R, De Pauw M, Dennis ES, Good AG (2003) Enhanced low oxygen survival in Arabidopsis through increased metabolic flux in the fermentative pathway. Plant Physiol 132:1292–1302
Jackson MB (2008) Ethylene-promoted elongation: an adaptation to submergence stress. Ann Bot 101:229–248
Ji CY, Chung WH, Kim HS, Jung WY, Kang L, Jeong JC, Kwak SS (2017) Transcriptome profiling of sweetpotato tuberous roots during low temperature storage. Plant Physiol Biochem 112:97–108
Ji CY, Bian X, Lee CJ, Kim HS, Kim SE, Park SC, Xie Y, Guo X, Kwak SS (2019) De novo transcriptome sequencing and gene expression profiling of sweet potato leaves during low temperature stress and recovery. Gene 700:23–30
Jiang X, Jianjun H, Wang Y (2004) Sweetpotato processing and product research and development at the Sichuan Academy of Agricultural Sciences. In: Fuglie KO, Hermann M (eds) Sweetpotato post-harvest research and development in China. Proceedings of an International Workshop held in Chengdu, Sichuan, PR China, November 7–8, 2001, International Potato Center (CIP), Bogor, Indonesia.
Jones P, Binns D, Chang HY, Fraser M, Li W, McAnulla C, McWilliam H, Maslen J, Mitchell A, Nuka G, Pesseat S, Quinn AF, Sangrador-Vegas A, Scheremetjew M, Yong SY, Lopez R, Hunter S (2014) InterProScan 5: genome-scale protein function classification. Bioinformatics 30:1236–1240
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Kang L, Park SC, Ji CY, Kim HS, Lee HS, Kwak SS (2017) Metabolic engineering of carotenoids in transgenic sweetpotato. Breed Sci 67:27–34
Kende H (1993) Ethylene biosynthesis. Annu Rev Plant Biol 44:283–307
Kim HS, Wang W, Kang L, Kim SE, Lee CJ, Park SC, Park WS, Ahn MJ, Kwak SS (2020) Metabolic engineering of low molecular weight antioxidants in sweetpotato. Plant Biotechnol Rep 14:193–205
Kwak SS (2019) Biotechnology of the sweetpotato: ensuring global food and nutrition security in the face of climate change. Plant Cell Rep 38:1361–1363
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9:559
Lin KHR, Tsou CC, Hwang SY, Chen LFO, Lo HF (2006) Paclobutrazol pre-treatment enhanced flooding tolerance of sweet potato. J Plant Physiol 163:750–760
Lorbiecke R, Sauter M (1999) Adventitious root growth and cell-cycle induction in deepwater rice. Plant Physiol 119:21–30
Luo W, Brouwer C (2013) Pathview: an R/Bioconductor package for pathway-based data integration and visualization. Bioinformatics 29:1830–1831
O’Leary NA, Wright MW, Brister JR, Ciufo S, Haddad D, McVeigh R, Rajput B, Robbertse B, Smith-White B, Ako-Adjei D, Astashyn A, Badretdin A, Bao Y, Blinkova O, Brover V, Chetvernin V, Choi J, Cox E, Ermolaeva O, Farrell CM, Goldfarb T, Gupta T, Haft D, Hatcher E, Hlavina W, Joardar VS, Kodali VK, Li W, Maglott D, Masterson P, McGarvey KM, Murphy MR, O’Neill K, Pujar S, Rangwala SH, Rausch D, Riddick LD, Schoch C, Shkeda A, Storz SS, Sun H, Thibaud-Nissen F, Tolstoy I, Tully RE, Vatsan AR, Wallin C, Webb D, Wu W, Landrum MJ, Kimchi A, Tatusova T, DiCuccio M, Kitts P, Murphy TD, Pruitt KD (2016) Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res 44(D1):D733–D745
Olson DC, Oetiker JH, Yang SF (1995) Analysis of LE-ACS3, a 1-aminocyclopropane-1-carboxylic acid synthase gene expressed during flooding in the roots of tomato plants. J Biol Chem 270:14056–14061
Pagnussat GC, Lanteri ML, Lombardo MC, Lamattina L (2004) Nitric oxide mediates the indole acetic acid induction activation of a mitogen-activated protein kinase cascade involved in adventitious root development. Plant Physiol 135:279–286
Park SU, Lee CJ, Kim SE, Lim YH, Lee HU, Nam SS, Kim HS, Kwak SS (2020) Selection of flooding stress tolerant sweetpotato cultivars based on biochemical and phenotypic characterization. Plant Physiol Biochem 155:243–251
Park SC, Kim YH, Ji CY, Park S, Jeong JC, Lee HS, Kwak SS (2012) Stable internal reference genes for the normalization of real-time PCR in different sweetpotato cultivars subjected to abiotic stress conditions. PLoS ONE 7:e51502
Patro R, Duggal G, Love MI, Irizarry RA, Kingsford C (2017) Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods 14:417–419
Pierik R, Tholen D, Poorter H, Visser EJ, Voesenek LA (2006) The Janus face of ethylene: growth inhibition and stimulation. Trends Plant Sci 11:176–183
Ponnamperuma FN (1972) The chemistry of submerged soils. Adv Agron 24:29–96
Puig CP, Dagar A, Ibanez CM, Singh V, Crisosto CH, Friedman H, Lurie S, Granell A (2015) Pre-symptomatic transcriptome changes during cold storage of chilling sensitive and resistant peach cultivars to elucidate chilling injury mechanisms. BMC Genet 16:245
Ridge I (1987) Ethylene and growth control in amphibious plants. In: Crawford RMM (ed) Plant life in aquatic and amphibious habitats. Oxford University Press, Oxford, pp 53–76
Ritchie ME, Phipson B, Wu DI, Hu Y, Law CW, Shi W, Smyth GK (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43:e47–e47
Roberts W, Russo V (1991) Time of flooding and cultivar affect sweetpotato yield. HortScience 26:1473–1474
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140
Rout GR, Sahoo S (2015) Role of iron in plant growth and metabolism. Rev Agric Sci 3:1–24
Sairam RK, Kumutha D, Ezhilmathi K, Deshmukh PS, Srivastava GC (2008) Physiology and biochemistry of waterlogging tolerance in plants. Biol Plant 52:401
Sasidharan R, Voesenek LA (2015) Ethylene-mediated acclimations to flooding stress. Plant Physiol 169:3–12
Setter TL, Waters I (2003) Review of prospects for germplasm improvement for waterlogging tolerance in wheat, barley and oats. Plant Soil 253:1–34
Shiu OY, Oetiker JH, Yip WK, Yang SF (1998) The promoter of LE-ACS7, an early flooding-induced 1-aminocyclopropane-1-carboxylate synthase gene of the tomato, is tagged by a Sol3 transposon. Proc Natl Acad Sci USA 95:10334–10339
Sinha AK, Jaggi M, Raghuram B, Tuteja N (2011) Mitogen-activated protein kinase signaling in plants under abiotic stress. Plant Signal Behav 6:196–203
Sivanandhan G, Arun M, Mayavan S, Rajesh M, Jeyaraj M, Dev GK, Manickavasagam M, Selvaraj N, Ganapathi A (2012) Optimization of elicitation conditions with methyl jasmonate and salicylic acid to improve the productivity of withanolides in the adventitious root culture of Withania somnifera (L.) Dunal. Appl Biochem Biotechnol 168:681–696
Smulders MJ, Horton RF (1991) Ethylene promotes elongation growth and auxin promotes radial growth in Ranunculus sceleratus petioles. Plant Physiol 96:806–811
Smyth GK, Speed T (2003) Normalization of cDNA microarray data. Methods 31:265–273
Summers JE, Jackson MB (1996) Anaerobic promotion of stem extension in Potamogeton pectinatus. Roles for carbon dioxide, acidification and hormones. Physiol Plant 96:615–622
Sun J, Nishiyama T, Shimizu K, Kadota K (2013) TCC: an R package for comparing tag count data with robust normalization strategies. BMC Bioinform 14:219
Takahashi H, Xiaohua Q, Shimamura S, Yanagawa A, Hiraga S, Nakazono M (2018) Sucrose supply from leaves is required for aerenchymatous phellem formation in hypocotyl of soybean under waterlogged conditions. Ann Bot 121:723–732
Unger IM, Kennedy AC, Muzika RM (2009) Flooding effects on soil microbial communities. Agric Ecosyst Environ Appl Soil Ecol 42:1–8
UniProt Consortium (2019) UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res 47(D1):D506–D515
Voesenek LACJ, Colmer TD, Pierik R, Millenaar FF, Peeters AJM (2006) How plants cope with complete submergence. New Phytol 170:213–226
Yang C, Lu X, Ma B, Chen SY, Zhang JS (2015) Ethylene signaling in rice and Arabidopsis: conserved and diverged aspects. Mol Plant 8:495–505
Yang J, Moeinzadeh MH, Kuhl H, Helmuth J, Xiao P, Haas S, Liu G, Zheng J, Sun Z, Fan W, Deng G, Wang H, Hu F, Zhao S, Fernie AR, Boerno S, Timmermann B, Zhang P, Deng G (2017) Haplotype-resolved sweet potato genome traces back its hexaploidization history. Nat Plants 3:696–703
Yu G, Wang LG, Han Y, He QY (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. Omics 16:284–287
Yu KW, Gao W, Hahn EJ, Paek KY (2002) Jasmonic acid improves ginsenoside accumulation in adventitious root culture of Panax ginseng CA Meyer. Biochem Eng J 11:211–215
Zhang P, Lyu D, Jia L, He J, Qin S (2017) Physiological and de novo transcriptome analysis of the fermentation mechanism of Cerasus sachalinensis roots in response to short-term waterlogging. BMC Genom 18:649
Zhu A, Li W, Ye J, Sun X, Ding Y, Cheng Y, Deng X (2011) Microarray expression profiling of postharvest Ponkan mandarin (Citrus reticulata) fruit under cold storage reveals regulatory gene candidates and implications on soluble sugars metabolism. J Integr Plant Biol 53:358–374
Acknowledgements
This work was supported by the Korea Institute of Planning and Evaluation for Technology in Food, Agriculture, Forestry (IPET) through the Agri-Bio Industry Technology Development Program funded by the Ministry of Agriculture, Food and Rural Affairs (MAFRA) (118038-3) and the KRIBB initiative program.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Park, SU., Kim, YH., Lee, CJ. et al. Comparative transcriptome profiling of two sweetpotato cultivars with contrasting flooding stress tolerance levels. Plant Biotechnol Rep 14, 743–756 (2020). https://doi.org/10.1007/s11816-020-00650-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-020-00650-5