Abstract
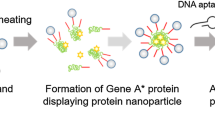
An anticancer drug delivery system consisting of DNA nanoparticles synthesized by rolling circle amplification (RCA) was developed for prostate cancer membrane antigen (PSMA) targeted cancer therapy. The template of RCA was a DNA oligodeoxynucleotide coded with PSMA-targeted aptamer, drug-loading domain, primer binding site and pH-sensitive spacer. Anticancer drug doxorubicin, as the model drug, was loaded into the drug-loading domain (multiple GC-pair sequences) of the DNA nanoparticles by intercalation. Due to the integrated pH-sensitive spacers in the nanoparticles, in an acidic environment, the cumulative release of doxorubicin was far more than the cumulative release of the drug in the normal physiological environment. In cell uptake experiments, treated with doxorubicin loaded DNA nanoparticles, PSMA-positive C4-2 cells could take up more doxorubicin than PSMA-null PC-3 cells. The prepared DNA nanoparticles showed the potential as drug delivery system for PSMA targeting prostate cancer therapy.

Similar content being viewed by others
References
Chen W, Zheng R, Baade P D, Zhang S, Zeng H, Bray F, Jemal A, Yu X Q, He J. Cancer statistics in China, 2015. CA: A Cancer Journal for Clinicians, 2016, 66(2): 115–132
Billingham M E, Bristow M R, Glatstein E, Mason J W, Masek M A, Daniels J R. Adriamycin cardiotoxicity: Endomyocardial biopsy evidence of enhancement by irradiation. American Journal of Surgical Pathology, 1977, 1(1): 17–23
Brannon-Peppas L, Blanchette J O. Nanoparticle and targeted systems for cancer therapy. Advanced Drug Delivery Reviews, 2004, 56(11): 1649–1659
Ghosh A, Heston W D. Tumor target prostate specific membrane antigen (PSMA) and its regulation in prostate cancer. Journal of Cellular Biochemistry, 2004, 91(3): 528–539
Farokhzad O C, Jon S, Khademhosseini A, Tran T N, Lavan D A, Langer R. Nanoparticle-aptamer bioconjugates: A new approach for targeting prostate cancer cells. Cancer Research, 2004, 64(21): 7668–7672
Kanwar J, Roy K, Maremanda N, Subramanian K, Veedu R, Bawa R. Nucleic acid-based aptamers: Applications, development and clinical trials. Current Medicinal Chemistry, 2015, 22(21): 2539–2557
Jia R, Wang T, Jiang Q, Wang Z, Song C, Ding B. Self-assembled DNA nanostructures for drug delivery. Chinese Journal of Chemistry, 2016, 34(3): 265–272
Zhu G, Niu G, Chen X. Aptamer-drug conjugates. Bioconjugate Chemistry, 2015, 26(11): 2186–2197
Stoltenburg R, Reinemann C, Strehlitz B. SELEX—a (r)evolutionary method to generate high-affinity nucleic acid ligands. Biomolecular Engineering, 2007, 24(4): 381–403
Zhu H, Li J, Zhang X B, Ye M, Tan W. Nucleic acid aptamer—mediated drug delivery for targeted cancer therapy. ChemMedChem, 2015, 10(1): 39–45
Keefe A D, Pai S, Ellington A. Aptamers as therapeutics. Nature Reviews. Drug Discovery, 2010, 9(7): 537–550
Nimjee S M, Rusconi C P, Sullenger B A. Aptamers: An emerging class of therapeutics. Annual Review of Medicine, 2005, 56(1): 555–583
Palchetti I, Mascini M. Nucleic acid biosensors for environmental pollution monitoring. Analyst (London), 2008, 133(7): 846–854
Lupold S E, Hicke B J, Lin Y, Coffey D S. Identification and characterization of nuclease-stabilized RNA molecules that bind human prostate cancer cells via the prostate-specific membrane antigen. Cancer Research, 2002, 62(14): 4029–4033
Dhar S, Gu F X, Langer R, Farokhzad O C, Lippard S J. Targeted delivery of cisplatin to prostate cancer cells by aptamer functionalized Pt(IV) prodrug-PLGA-PEG nanoparticles. Proceedings of the National Academy of Sciences of the United States of America, 2008, 105(45): 17356–17361
Bagalkot V, Farokhzad O C, Langer R, Jon S. An aptamerdoxorubicin physical conjugate as a novel targeted drug-delivery platform. Angewandte Chemie International Edition, 2006, 45(48): 8149–8152
Lee I H, An S, Yu M K, Kwon H K, Im S H. Targeted chemoimmunotherapy using drug-loaded aptamer-dendrimer bioconjugates. Journal of Controlled Release, 2011, 155(3): 435–441
Tan L, Neoh K G, Kang E T, Choe W S, Su X. PEGylated anti- MUC1 aptamer-doxorubicin complex for targeted drug delivery to MCF7 breast cancer cells. Macromolecular Bioscience, 2011, 11 (10): 1331–1335
Boyacioglu O, Stuart C H, Kulik G, Gmeiner W H. Dimeric DNA aptamer complexes for high-capacity-targeted drug delivery using pH-sensitive covalent linkages. Mol Therapy-Nucleic Acids, 2013, 2(1): e107
Stuart C H, Singh R, Smith T L, D’Agostino R Jr, Caudell D, Balaji K C, Gmeiner W H. Prostate-specific membrane antigen-targeted liposomes specifically deliver the Zn2+ chelator TPEN inducing oxidative stress in prostate cancer cells. Nanomedicine (London), 2016, 11(10): 1207–1222
Fire A, Xu S Q. Rolling replication of short DNA circles. Proceedings of the National Academy of Sciences of the United States of America, 1995, 92(10): 4641–4645
Zhao W, Ali M M, Brook M A, Li Y. Rolling circle amplification: Applications in nanotechnology and biodetection with functional nucleic acids. Angewandte Chemie International Edition, 2008, 47 (34): 6330–6337
Ali M M, Li F, Zhang Z, Zhang K, Kang D K, Ankrum J A, Le X C, Zhao W. Rolling circle amplification: A versatile tool for chemical biology, materials science and medicine. Chemical Society Reviews, 2014, 43(10): 3324–3341
Roh Y H, Lee J B, Shopsowitz K E, Dreaden E C, Morton S W, Poon Z, Hong J, Yamin I, Bonner D K, Hammond P T. Layer-bylayer assembled antisense DNA microsponge particles for efficient delivery of cancer therapeutics. ACS Nano, 2014, 8(10): 9767–9780
Lee H Y, Jeong H, Jung I Y, Jang B, Seo Y C, Lee H, Lee H. DhITACT: DNA hydrogel formation by isothermal amplification of complementary target in fluidic channels. Advanced Materials, 2015, 27(23): 3513–3517
Hamblin G D, Carneiro K M, Fakhoury J F, Bujold K E, Sleiman H F. Rolling circle amplification-templated DNA nanotubes show increased stability and cell penetration ability. Journal of the American Chemical Society, 2012, 134(6): 2888–2891
Mei L, Zhu G, Qiu L, Wu C, Chen H, Liang H, Cansiz S, Lv Y, Zhang X, TanW. Self-assembled multifunctional DNA nanoflowers for the circumvention of multidrug resistance in targeted anticancer drug delivery. Nano Research, 2015, 8(11): 3447–3460
Lizardi PM, Huang X, Zhu Z, Brayward P, Thomas D C,Ward D C. Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nature Genetics, 1998, 19(3): 225–232
Am Hong C, Jang B, Jeong E H, Jeong H, Lee H. Self-assembled DNA nanostructures prepared by rolling circle amplification for the delivery of siRNA conjugates. Chemical Communications, 2014, 50 (86): 13049–13051
Lv Y, Hu R, Zhu G, Zhang X, Mei L, Liu Q, Qiu L, Wu C, Tan W. Preparation and biomedical applications of programmable and multifunctional DNA nanoflowers. Nature Protocols, 2015, 10(10): 1508–1524
Zhang L, Zhu G, Mei L, Wu C, Qiu L, Cui C, Liu Y, Teng I T, Tan W. Self-assembled DNA immuno nanoflowers as multivalent CpG nanoagents. ACS Applied Materials & Interfaces, 2015, 7(43): 24069–24074
Sun W, Jiang T, Lu Y, Reiff M, Mo R, Gu Z. Cocoon-like selfdegradable DNA nanoclew for anticancer drug delivery. Journal of the American Chemical Society, 2014, 136(42): 14722–14725
Malloy A. Count, size and visualize nanoparticles. Materials Today, 2011, 14(4): 170–173
Zhu G, Hu R, Zhao Z, Chen Z, Zhang X, TanW. Noncanonical selfassembly of multifunctional DNA nanoflowers for biomedical applications. Journal of the American Chemical Society, 2013, 135 (44): 16438–16445
Acknowledgements
This work was supported in part by National Key Research and Development Plan (2016YFE0119200) and the National Natural Science Foundation of China (Grant Nos. 81402856, A3 project-81361140344 and 21402143). This research was also partially sponsored by Tianjin Municipal Science and Technology Commission (15JCYBJC28700 and 15JCQNJC13600) and National Students’ Innovation and Entrepreneurship Training Program (201510062008).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Zhang, P., Ye, J., Liu, E. et al. Aptamer-coded DNA nanoparticles for targeted doxorubicin delivery using pH-sensitive spacer. Front. Chem. Sci. Eng. 11, 529–536 (2017). https://doi.org/10.1007/s11705-017-1645-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11705-017-1645-z