Abstract
Purpose
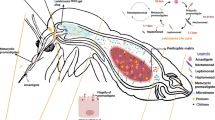
Sand flies are the only proven vectors of leishmaniases, a tropical neglected disease endemic in at least 92 countries. Vector–parasite interactions play a significant role in vector-borne disease transmission. There are various bottlenecks to Leishmania colonization of the sand fly midgut. Such bottlenecks include the production of innate immune-related molecules, digestive proteases, parasite impermeable peritrophic membrane, and resident gut microbiota. These barriers determine the parasite load transmitted and, consequently, the disease outcome in mammalian host. Therefore, it is important to understand the molecular responses of both sand fly and Leishmania during infection.
Method
Here, we reviewed the published literature on sand fly–Leishmania interactions bringing together earlier and current findings to highlight new developments and research gaps in the field.
Conclusion
Recent research studies on sand fly–Leishmania interaction have revealed contrasting observations to past studies. However, how Leishmania parasites evade the sand fly immune response still needs further research. Sand fly response to Leishmania infection can be best understood by analyzing its tissue transcriptome. Better characterization of the role of midgut components could be a game changer in development of transmission-blocking strategies for leishmaniasis.



Similar content being viewed by others
References
Maurício I (2018) Leishmania taxonomy. In: Bruschi F, Gradoni L (eds) The leishmaniases: old neglected tropical disease, 1st edn. Springer, pp 17–20 (Retrieved 13 Mar 2021)
Gossage S, Rogers M, Bates P (2003) Two separate growth phases during the development of Leishmania in sand flies: implications for understanding the life cycle. Int J Parasitol 33(10):1027–1034. https://doi.org/10.1016/s0020-7519(03)00142-5
Rogers M, Bates P (2007) Leishmania manipulation of sand fly feeding behavior results in enhanced transmission. PLoS Pathog 3(6):e91. https://doi.org/10.1371/journal.ppat.0030091
Dvorak V, Shaw J, Volf P (2018) Parasite biology: the vectors. In: Bruschi F, Gradoni L (eds) The leishmaniases: old neglected tropical diseases, 1st edn. Springer, Cham, pp 31–40 (Retrieved 13 March 2021)
WHO (2021) Leishmaniasis. Who.int. Retrieved 14 March 2021, from https://www.who.int/westernpacific/health-topics/leishmaniasis.
Bates P, Depaquit J, Galati E, Kamhawi S, Maroli M, McDowell M et al (2015) Recent advances in phlebotomine sand fly research related to leishmaniasis control. Parasites Vectors 8(1):131. https://doi.org/10.1186/s13071-015-0712-x
Killick-Kendrick R (1990) Phlebotomine vectors of the leishmaniases: a review. Med Vet Entomol 4(1):1–24. https://doi.org/10.1111/j.1365-2915.1990.tb00255.x
Oliveira F, Jochim R, Valenzuela J, Kamhawi S (2009) Sand flies, Leishmania, and transcriptome-borne solutions. Parasitol Int 58(1):1–5. https://doi.org/10.1016/j.parint.2008.07.004
Bates P (2007) Transmission of Leishmania metacyclic promastigotes by phlebotomine sand flies. Int J Parasitol 37(10):1097–1106
Wilson R, Bates M, Dostalova A, Jecna L, Dillon R, Volf P, Bates P (2010) Stage-specific adhesion of Leishmania promastigotes to sand fly midguts assessed using an improved comparative binding assay. PLoS Negl Trop Dis 4(9):e816. https://doi.org/10.1371/journal.pntd.0000816
Louradour I, Monteiro C, Inbar E, Ghosh K, Merkhofer R, Lawyer P et al (2017) The midgut microbiota plays an essential role in sand fly vector competence for Leishmania major. Cell Microbiol 19(10):e12755. https://doi.org/10.1111/cmi.12755
Campolina T, Villegas L, Monteiro C, Pimenta P, Secundino N (2020) Tripartite interactions: Leishmania, microbiota and Lutzomyia longipalpis. PLoS Negl Trop Dis 14(10):e0008666. https://doi.org/10.1371/journal.pntd.0008666
Kimblin N, Peters N, Debrabant A, Secundino N, Egen J, Lawyer P et al (2008) Quantification of the infectious dose of Leishmania major transmitted to the skin by single sand flies. Proc Natl Acad Sci 105(29):10125–10130. https://doi.org/10.1073/pnas.0802331105
Stamper L, Patrick R, Fay M, Lawyer P, Elnaiem D, Secundino N et al (2011) Infection parameters in the sand fly vector that predict transmission of Leishmania major. PLoS Negl Trop Dis 5(8):e1288. https://doi.org/10.1371/journal.pntd.0001288
Maia C, Seblova V, Sadlova J, Votypka J, Volf P (2011) Experimental transmission of Leishmania infantum by two major vectors: a comparison between a viscerotropic and a dermotropic strain. PLoS Negl Trop Dis 5(6):e1181. https://doi.org/10.1371/journal.pntd.0001181
Secundino N, de Freitas V, Monteiro C, Pires A, David B, Pimenta P (2012) The transmission of Leishmania infantum chagasi by the bite of the Lutzomyia longipalpis to two different vertebrates. Parasites Vectors. https://doi.org/10.1186/1756-3305-5-20
Yu X, Zhu Y, Ma C, Fabrick J, Kanost M (2002) Pattern recognition proteins in Manduca sexta plasma. Insect Biochem Mol Biol 32(10):1287–1293. https://doi.org/10.1016/s0965-1748(02)00091-7
Pal S, Wu LP (2009) Lessons from the fly: pattern recognition in Drosophila melanogaster. In: Kishore U (ed) Target pattern recognition in innate immunity. Advances in experimental medicine and biology, vol 653. Springer, New York NY, pp 45–46
Ramalho-Ortigão M, Jochim R, Anderson J, Lawyer P, Pham V, Kamhawi S, Valenzuela J (2007) Exploring the midgut transcriptome of Phlebotomus papatasi: comparative analysis of expression profiles of sugar-fed, blood-fed and Leishmania major-infected sand flies. BMC Genomics 8(1):300. https://doi.org/10.1186/1471-2164-8-300
Jochim R, Teixeira C, Laughinghouse A, Mu J, Oliveira F, Gomes R et al (2008) The midgut transcriptome of Lutzomyia longipalpis: comparative analysis of cDNA libraries from sugar-fed, blood-fed, post-digested and Leishmania infantum chagasi-infected sand flies. BMC Genomics 9(1):15. https://doi.org/10.1186/1471-2164-9-15
Abrudan J, Ramalho-Ortigão M, O’Neil S, Stayback G, Wadsworth M, Bernard M et al (2013) The characterization of the Phlebotomus papatasi transcriptome. Insect Mol Biol 22(2):211–232. https://doi.org/10.1111/imb.12015
Sloan M, Sadlova J, Lestinova T, Sanders M, Cotton J, Volf P, Ligoxygakis P (2021) The Phlebotomus papatasi systemic transcriptional response to trypanosomatid-contaminated blood does not differ from the non-infected blood meal. Parasites Vectors. https://doi.org/10.1186/s13071-020-04498-0
Boulanger N, Lowenberger C, Volf P, Ursic R, Sigutova L, Sabatier L et al (2004) Characterization of a defensin from the sand fly Phlebotomus duboscqi induced by challenge with bacteria or the protozoan parasite Leishmania major. Infect Immun 72(12):7140–7146. https://doi.org/10.1128/iai.72.12.7140-7146.2004
Hajmová M, Chang K, Kolli B, Volf P (2004) Down-regulation of gp63 in Leishmania amazonensis reduces its early development in Lutzomyia longipalpis. Microbes Infect 6(7):646–649. https://doi.org/10.1016/j.micinf.2004.03.003
Secundino N, Kimblin N, Peters N, Lawyer P, Capul A, Beverley S et al (2010) Proteophosphoglycan confers resistance of Leishmania major to midgut digestive enzymes induced by blood feeding in vector sand flies. Cell Microbiol 12(7):906–918. https://doi.org/10.1111/j.1462-5822.2010.01439.x
Evans J, Aronstein K, Chen Y, Hetru C, Imler J, Jiang H et al (2006) Immune pathways and defense mechanisms in honey bees Apis mellifera. Insect Mol Biol 15(5):645–656. https://doi.org/10.1111/j.1365-2583.2006.00682.x
Tanaka H, Ishibashi J, Fujita K, Nakajima Y, Sagisaka A, Tomimoto K et al (2008) A genome-wide analysis of genes and gene families involved in innate immunity of Bombyx mori. Insect Biochem Mol Biol 38(12):1087–1110. https://doi.org/10.1016/j.ibmb.2008.09.001
Clayton A, Dong Y, Dimopoulos G (2014) The Anopheles innate immune system in the defense against malaria infection. J Innate Immun 6(2):169–181. https://doi.org/10.1159/000353602
Klowden M (2007) Physiological systems in insects, 2nd edn. Elsevier/Academic Press, pp 378–382
Chapman R, Simpson S, Douglas A (2013) The insects, 5th edn. Cambridge University Press, pp 48–55 (64–65, 121–125)
Nation J (2015) Insect physiology and biochemistry, 3rd edn. Chapman and Hall/CRC, pp 45–50 (437-444)
Rosales C (2017) Cellular and molecular mechanisms of insect immunity. Insect Physiol Ecol. https://doi.org/10.5772/67107
Di-Blasi T, Telleria E, Marques C, Couto R, Silva-Neves M, Jancarova M et al (2019) Lutzomyia longipalpis TGF-β has a role in Leishmania infantum chagasi survival in the vector. Front Cell Infect Microbiol. https://doi.org/10.3389/fcimb.2019.00071
Telleria E, Azevedo-Brito D, Kykalová B, Tinoco-Nunes B, Pitaluga A, Volf P, Traub-Csekö Y (2021) Leishmania infantum infection modulates the Jak-STAT pathway in Lutzomyia longipalpis LL5 embryonic cells and adult females, and affects parasite growth in the sand fly. Front Trop Dis. https://doi.org/10.3389/fitd.2021.747820
Louradour I, Ghosh K, Inbar E, Sacks D (2019) CRISPR/Cas9 mutagenesis in Phlebotomus papatasi: the immune deficiency pathway impacts vector competence for Leishmania major. MBio. https://doi.org/10.1128/mbio.01941-19
Volf P, Kiewegova A, Nemec A (2002) Bacterial colonization in the gut of Phlebotomus duboscqi (Diptera: Psychodidae): transstadial passage and the role of female diet. Folia Parasitol 49(1):73–77. https://doi.org/10.14411/fp.2002.014
Telleria E, Sant’Anna M, Alkurbi M, Pitaluga A, Dillon R, Traub-Csekö Y (2013) Bacterial feeding, Leishmania infection and distinct infection routes induce differential defensin expression in Lutzomyia longipalpis. Parasites Vectors 6(1):12. https://doi.org/10.1186/1756-3305-6-12
Kykalová B, Tichá L, Volf P, Loza Telleria E (2021) Phlebotomus papatasi antimicrobial peptides in larvae and females and a gut-specific defensin upregulated by Leishmania major Infection. Microorganisms 9(11):2307. https://doi.org/10.3390/microorganisms9112307
Nimmo D, Ham P, Ward R, Maingon R (1997) The sand fly Lutzomyia longipalpis shows specific humoral responses to bacterial challenge. Med Vet Entomol 11(4):324–328. https://doi.org/10.1111/j.1365-2915.1997.tb00417.x
Dillon R, Ivens A, Churcher C, Holroyd N, Quail M, Rogers M et al (2006) Analysis of ESTs from Lutzomyia longipalpis sand flies and their contribution toward understanding the insect–parasite relationship. Genomics 88(6):831–840. https://doi.org/10.1016/j.ygeno.2006.06.011
Pitaluga A, Beteille V, Lobo A, Ortigão-Farias J, Dávila A, Souza A et al (2009) EST sequencing of blood-fed and Leishmania-infected midgut of Lutzomyia longipalpis, the principal visceral leishmaniasis vector in the Americas. Mol Genet Genom 282(3):307–317. https://doi.org/10.1007/s00438-009-0466-2
Dostálová A, Votýpka J, Favreau A, Barbian K, Volf P, Valenzuela J, Jochim R (2011) The midgut transcriptome of Phlebotomus (Larroussius) perniciosus, a vector of Leishmania infantum: comparison of sugar fed and blood fed sand flies. BMC Genom. https://doi.org/10.1186/1471-2164-12-223
Coutinho-Abreu I, Serafim T, Meneses C, Kamhawi S, Oliveira F, Valenzuela J (2020) Leishmania infection induces a limited differential gene expression in the sand fly midgut. BMC Genom. https://doi.org/10.1186/s12864-020-07025-8
Dong Y, Aguilar R, Xi Z, Warr E, Mongin E, Dimopoulos G (2006) Anopheles gambiae immune responses to human and rodent Plasmodium parasite species. PLoS Pathog 2(6):e52. https://doi.org/10.1371/journal.ppat.0020052
Bahia A, Kubota M, Tempone A, Pinheiro W, Tadei W, Secundino N et al (2010) Anopheles aquasalis infected by Plasmodium vivax displays unique gene expression profiles when compared to other malaria vectors and plasmodia. PLoS ONE 5(3):e9795. https://doi.org/10.1371/journal.pone.0009795
Smagghe G, Goodman C, Stanley D (2009) Insect cell culture and applications to research and pest management. In Vitro Cell Dev Biol Anim 45(3–4):93–105. https://doi.org/10.1007/s11626-009-9181-x
Tinoco-Nunes B, Telleria E, da Silva-Neves M, Marques C, Azevedo- Brito D, Pitaluga A, Traub-Csekö Y (2016) The sand fly Lutzomyia longipalpis LL5 embryonic cell line has active Toll and Imd path- ways and shows immune responses to bacteria, yeast and Leishmania. Parasites Vectors. https://doi.org/10.1186/s13071-016-1507-4
da Silva Gonçalves D, Iturbe-Ormaetxe I, Martins-da-Silva A, Telleria E, Rocha M, Traub-Csekö Y et al (2019) Wolbachia introduction into Lutzomyia longipalpis (Diptera: Psychodidae) cell lines and its effects on immune-related gene expression and interaction with Leishmania infantum. Parasites Vectors. https://doi.org/10.1186/s13071-018-3227-4
Telleria E, Tinoco-Nunes B, Leštinová T, de Avellar L, Tempone A, Pitaluga A et al (2021) Lutzomyia longipalpis antimicrobial peptides: differential expression during development and potential involvement in vector interaction with microbiota and Leishmania. Microorganisms 9(6):1271. https://doi.org/10.3390/microorganisms9061271
Jing T, Qi F, Wang Z (2020) Most dominant roles of insect gut bacteria: digestion, detoxification, or essential nutrient provision. Microbiome. https://doi.org/10.1186/s40168-020-00823-y
Omondi ZN, Demir S (2020) Bacteria composition and diversity in the gut of sand fly: impact on Leishmania and sand fly development. Int J Trop Insect Sci 41(1):25–32. https://doi.org/10.1007/s42690-020-00184-x
Telleria E, Sant’Anna M, Ortigão-Farias J, Pitaluga A, Dillon V, Bates P et al (2012) Caspar-like gene depletion reduces Leishmania infection in sand fly host Lutzomyia longipalpis. J Biol Chem 287(16):12985–12993. https://doi.org/10.1074/jbc.m111.331561
Bang I (2019) JAK/STAT signaling in insect innate immunity. Entomol Res 49(8):339–353. https://doi.org/10.1111/1748-5967.12384
Gupta L, Molina-Cruz A, Kumar S, Rodrigues J, Dixit R, Zamora R, Barillas-Mury C (2009) The STAT pathway mediates late-phase immunity against Plasmodium in the mosquito Anopheles gambiae. Cell Host Microbe 5(5):498–507. https://doi.org/10.1016/j.chom.2009.04.003
Bahia A, Kubota M, Tempone A, Araújo H, Guedes B, Orfanó A et al (2011) The JAK-STAT pathway controls Plasmodium vivax load in early stages of Anopheles aquasalis infection. PLoS Negl Trop Dis 5(11):e1317. https://doi.org/10.1371/journal.pntd.0001317
Lehane M (1997) Peritrophic matrix structure and function. Annu Rev Entomol 42(1):525–550. https://doi.org/10.1146/annurev.ento.42.1.525
Hegedus D, Erlandson M, Gillott C, Toprak U (2009) New insights into peritrophic matrix synthesis, architecture, and function. Annu Rev Entomol 54(1):285–302. https://doi.org/10.1146/annurev.ento.54.110807.090559
Blackburn K, Wallbanks K, Molyneux D, Lavin D, Winstanley S (1988) The peritrophic membrane of the female sand fly Phlebotomus papatasi. Ann Trop Med Parasitol 82(6):613–619. https://doi.org/10.1080/00034983.1988.11812297
Devenport M, Alvarenga P, Shao L, Fujioka H, Bianconi M, Oliveira P, Jacobs-Lorena M (2006) Identification of the Aedes aegypti peritrophic matrix protein AeIMUCI as a heme-binding protein. Biochemistry 45(31):9540–9549. https://doi.org/10.1021/bi0605991
Pimenta P, Modi G, Pereira S, Shahabuddin M, Sacks D (1997) A novel role for the peritrophic matrix in protecting Leishmania from the hydrolytic activities of the sand fly midgut. Parasitology 115(4):359–369. https://doi.org/10.1017/s0031182097001510
Schlein Y, Schnur L, Jacobson R (1990) Released glycoconjugate of indigenous Leishmania major enhances survival of a foreign L. major in Phlebotomus papatasi. Trans R Soc Trop Med Hyg 84(3):353–355. https://doi.org/10.1016/0035-9203(90)90315-6
Rogers M, Hajmová M, Joshi M, Sadlova J, Dwyer D, Volf P, Bates P (2008) Leishmania chitinase facilitates colonization of sand fly vectors and enhances transmission to mice. Cell Microbiol 10(6):1363–1372. https://doi.org/10.1111/j.1462-5822.2008.01132.x
Sádlová J, Volf P (2009) Peritrophic matrix of Phlebotomus duboscqi and its kinetics during Leishmania major development. Cell Tissue Res 337(2):313–325. https://doi.org/10.1007/s00441-009-0802-1
Pruzinova K, Sadlova J, Seblova V, Homola M, Votypka J, Volf P (2015) Comparison of bloodmeal digestion and the peritrophic matrix in four sand fly species differing in susceptibility to Leishmania donovani. PLoS ONE 10(6):e0128203. https://doi.org/10.1371/journal.pone.0128203
Sadlova J, Dvorak V, Seblova V, Warburg A, Votypka J, Volf P (2013) Sergentomyia schwetzi is not a competent vector for Leishmania donovani and other Leishmania species pathogenic to humans. Parasit Vectors 6:186–196
Sadlova J, Homola M, Myskova J, Jancarova M, Volf P (2018) Refractoriness of Sergentomyia schwetzi to Leishmania spp. is mediated by the peritrophic matrix. PLOS Negl Trop Dis 12(4):e0006382
Telleria E, Araújo A, Secundino N, d’Avila-Levy C, Traub-Csekö Y (2010) Trypsin-like serine proteases in Lutzomyia longipalpis – expression, activity and possible modulation by Leishmania infantum chagasi. PLoS ONE 5(5):e10697. https://doi.org/10.1371/journal.pone.0010697
Cao X, Gulati M, Jiang H (2017) Serine protease-related proteins in the malaria mosquito, Anopheles gambiae. Insect Biochem Mol Biol 88:48–62. https://doi.org/10.1016/j.ibmb.2017.07.008
Ramalho-Ortigao M, Saraiva E, Traub-Csekö Y (2010) Sand fly-Leishmania interactions: long relationships are not necessarily easy. Open Parasitol J 4(1):195–204. https://doi.org/10.2174/1874421401004010195
Schlein Y, Romano H (1986) Leishmania major and L. donovani: effects on proteolytic enzymes of Phlebotomus papatasi (Diptera, Psychodidae). Exp Parasitol 62(3):376–380. https://doi.org/10.1016/0014-4894(86)90045-7
Dillon RJ, Lane RP (1993) Influence of Leishmania infection on blood- meal digestion in the sandflies Phlebotomus papatasi and P. langeroni. Parasitol Res 79:492–496
Lawyer P, Githure J, Roberts C, Anjili C, Mebrahtu Y, Ngumbi P et al (1990) Development of Leishmania major in Phlebotomus duboscqi and Sergentomyia schwetzi (Diptera: Psychodidae). Am J Trop Med Hyg 43(1):31–43. https://doi.org/10.4269/ajtmh.1990.43.31
Sant’Anna M, Diaz-Albiter H, Mubaraki M, Dillon R, Bates P (2009) Inhibition of trypsin expression in Lutzomyia longipalpis using RNAi enhances the survival of Leishmania. Parasit Vectors 2(1):62. https://doi.org/10.1186/1756-3305-2-62
Verma S, Das S, Mandal A, Ansari M, Kumari S, Mansuri R, Kumar A, Singh R, Saini S, Abhishek K, Kumar V, Sahoo G, Das P (2017) Role of inhibitors of serine peptidases in protecting Leishmania donovani against the hydrolytic peptidases of sand fly midgut. Parasites Vectors. https://doi.org/10.1186/s13071-017-2239-9
Sacks DL et al (2000) The role of phosphoglycans in Leishmania- sand fly interactions. Proc Natl Acad Sci USA 97:406–411
Dostalova A, Volf P (2012) Leishmania development in sand flies: parasite-vector interactions overview. Parasites Vectors 5:276. https://doi.org/10.1186/1756-3305-5-276
Moraes C, Lucena S, Moreira B, Brazil R, Gontijo N, Genta F (2012) Relationship between digestive enzymes and food habit of Lutzomyia longipalpis (Diptera: Psychodidae) larvae: characterization of carbohydrases and digestion of microorganisms. J Insect Physiol 58(8):1136–1145. https://doi.org/10.1016/j.jinsphys.2012.05.015
Peterkova-Koci K, Robles-Murguia M, Ramalho-Ortigao M, Zurek L (2012) Significance of bacteria in oviposition and larval development of the sand fly Lutzomyia longipalpis. Parasit Vectors 5(1):145. https://doi.org/10.1186/1756-3305-5-145
Diaz-Albiter H, Sant’Anna M, Genta F, Dillon R (2012) Reactive oxygen species-mediated immunity against Leishmania mexicana and Serratia marcescens in the Phlebotomine sand fly Lutzomyia longipalpis. J Biol Chem 287(28):23995–24003. https://doi.org/10.1074/jbc.m112.376095
Sant’Anna M, Diaz-Albiter H, Aguiar-Martins K, Al Salem W, Cavalcante R, Dillon V et al (2014) Colonization resistance in the sand fly gut: Leishmania protects Lutzomyia longipalpis from bacterial infection. Parasites Vectors 7(1):329. https://doi.org/10.1186/1756-3305-7-329
Marayati B, Schal C, Ponnusamy L, Apperson C, Rowland T, Wasserberg G (2015) Attraction and oviposition preferences of Phlebotomus papatasi (Diptera: Psychodidae), vector of old-world cutaneous leishmaniasis, to larval rearing media. Parasites Vectors. https://doi.org/10.1186/s13071-015-1261-z
Kelly P, Bahr S, Serafim T, Ajami N, Petrosino J, Meneses C et al (2017) The gut microbiome of the vector Lutzomyia longipalpis is essential for survival of Leishmania infantum. MBio 8(1):e01121-e1216. https://doi.org/10.1128/mbio.01121-16
Telleria EL, Martins-da-Silva A, Tempone AJ, Traub-Csekö YM (2018) Leishmania, microbiota and sand fly immunity. Parasitology 145(10):1336–1353. https://doi.org/10.1017/s0031182018001014
Hurwitz I, Hillesland H, Fieck A, Das P, Durvasula R (2011) The paratransgenic sand fly: a platform for control of Leishmania transmission. Parasites Vectors. https://doi.org/10.1186/1756-3305-4-82
Wu S, Liao C, Pan R, Juang J (2012) Infection-induced intestinal oxidative stress triggers organ-to-organ immunological communication in Drosophila. Cell Host Microbe 11(4):410–417. https://doi.org/10.1016/j.chom.2012.03.004
Kamhawi S (2006) Phlebotomine sand flies and Leishmania parasites: friends or foes? Trends Parasitol 22(9):439–445. https://doi.org/10.1016/j.pt.2006.06.012
Volf P, Myskova J (2007) Sand flies and Leishmania: specific versus permissive vectors. Trends Parasitol 23(3):91–92. https://doi.org/10.1016/j.pt.2006.12.010
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Omondi, Z.N., Arserim, S.K., Töz, S. et al. Host–Parasite Interactions: Regulation of Leishmania Infection in Sand Fly. Acta Parasit. 67, 606–618 (2022). https://doi.org/10.1007/s11686-022-00519-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11686-022-00519-3