Abstract
Introduction
According to the Chinese Health Commission guidelines, coronavirus disease 2019 (COVID-19) severity is classified as mild, moderate, severe, or critical. The mortality rate of COVID-19 is higher among patients with severe and critical diseases; therefore, early identification of COVID-19 prevents disease progression and improves patient survival. Computed tomography (CT) radiomics, as a machine learning method, provides an objective and mathematical evaluation of COVID-19 pneumonia. As CT-based radiomics research has recently focused on COVID-19 diagnosis and severity analysis, this meta-analysis aimed to investigate the predictive power of a CT-based radiomics model in determining COVID-19 severity.
Materials and methods
This study followed the diagnostic version of PRISMA guidelines. PubMed, Embase databases and the Cochrane Central Register of Controlled Trials, and the Cochrane Database of Systematic Reviews were searched to identify relevant articles in the meta-analysis from inception until July 16, 2021. The sensitivity and specificity were analyzed using forest plots. The overall predictive power was calculated using the summary receiver operating characteristic curve. The bias was evaluated using a funnel plot. The quality of the included literature was assessed using the radiomics quality score and quality assessment of diagnostic accuracy studies tool.
Results
The radiomics quality scores ranged from 7 to 16 (achievable score: 2212 8 to 36). The pooled sensitivity and specificity were 0.800 (95% confidence interval [CI] 0.662–0.891) and 0.874 (95% CI 0.773–0.934), respectively. The pooled area under the receiver operating characteristic curve was 0.908. The quality assessment tool showed favorable results.
Conclusion
This meta-analysis demonstrated that CT-based radiomics models might be helpful for predicting the severity of COVID-19 pneumonia.
Similar content being viewed by others
Introduction
Coronavirus disease 2019 (COVID-19) is a pandemic [1]. COVID-19 has spread worldwide and has led to millions of deaths. According to the Chinese Health Commission (CHC) guidelines, COVID-19 severity is classified as mild, moderate, severe, or critical [2]. The Chinese Center for Disease Control and Prevention reported that 81% of COVID-19 cases were non-severe, and the remaining 19% were severe or critical [3]. Existing epidemiological studies suggest that the mortality rate of patients with severe COVID-19 is more than ten times higher than that of patients with non-severe COVID-19 [4]. To treat patients with COVID-19, early identification of severe cases directly influences treatment and prevents clinical deterioration. Similarly, early identification and management of patients with severe COVID-19 prevent disease progression and improve survival [5].
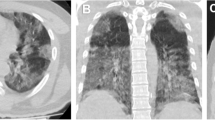
According to recent experience, abnormal findings on lung imaging appear before clinical symptoms develop, which highlights the importance of lung imaging in screening for COVID-19 pneumonia [6]. Computed tomography (CT) is helpful for COVID-19 diagnosis and in assessing COVID-19 pneumonia progression [7, 8]. The typical findings on chest CT imagery for patients with COVID-19 are ground-glass opacities and bilateral lung consolidations with peripheral involvement [9]. However, the evaluation of these conventional textures varies among radiologists and is often subjective.
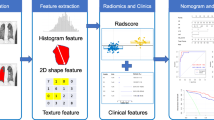
Computed tomography radiomics, a non-invasive developing machine learning technology, can extract histograms, shapes, or textural features from images. In addition, artificial intelligence can further quantify textural information using mathematical analysis; therefore, abnormal lesions on CT images can be evaluated precisely and objectively using radiomics. Recently, CT-based radiomics has been widely used for tumor diagnosis, cancer treatment, and prognosis assessment [10, 11].
In previous studies on COVID-19, machine learning CT-based radiomics has been shown to help diagnose and differentiate COVID-19 pneumonia from pneumonia caused by other pathogens [12,13,14]. Additionally, CT-based radiomics reportedly predicts the severity and outcome of COVID-19 pulmonary opacities [15]. However, the mechanism between COVID-19 pneumonia severity, pulmonary opacities, and clinical manifestations has not been well addressed, and a detailed meta-analysis using CT-based radiomics has not been performed. Therefore, this study aimed to investigate whether CT-based radiomics models can predict COVID-19 pneumonia severity.
Materials and methods
Study protocol and literature search
This study followed the diagnostic version of PRISMA guidelines [16]. Two investigators searched PubMed, Embase, the Cochrane Central Register of Controlled Trials and the Cochrane Database of Systematic Reviews for articles published between the inception of the databases until July 16, 2021. The keywords used were as follows: (“COVID-19” OR “severe acute respiratory coronavirus-2[SARS-CoV-2]”) AND (“radiomics” OR “textural”) AND (“computed tomography” OR “CT”).
Literature selection criteria
The inclusion criteria were as follows:
-
1.
Studies using shape- and texture-based radiomics to predict COVID-19 severity.
-
2.
Studies wherein COVID-19 severity was defined according to the CHC guidelines.
-
3.
Studies with full text available.
-
4.
Studies published in the English language.
In contrast, the exclusion criteria were as follows:
-
1.
Studies wherein radiomics was not used to predict the severity of COVID-19.
-
2.
Conference posters or papers for which only the abstract was available.
COVID-19 pneumonia severity classification
According to the CHC guidelines, COVID-19 illness is classified according to disease severity [4]. Patients with COVID-19 pneumonia included in this study were classified into those with non-severe disease (non-SVD) and those with severe disease (SVD). Patients who met any of the following criteria were included in the SVD group: (1) respiratory rate ≥ 30 times per minute, (2) oxygen saturation ≤ 93% by finger oximetry at resting status, (3) partial pressure of oxygen in arterial blood (PaO2)/fraction of inspired oxygen (FiO2) ≤ 300 mmHg), (4) patients with > 50% lesion progression on chest imaging over 1–2 days, (5) respiratory failure and assisted ventilation requirement; (6) shock, or (7) organ failure that required admission to the intensive care unit (ICU).
Data collection
We extracted the true-positive, false-positive, false-negative, and true-negative rates from the literature. The radiomics model with the highest area under the receiver operating characteristic curve (AUC) within the articles was used for extraction. Some studies used bootstrapping or cross-validation; therefore, the resulting values were not integers that could be used for extraction. For simplicity, we rounded the figures used in the calculations. Additionally, we extracted other information from the literature, including the author details, publication year, nation, number of patients, and further information.
Statistical analysis
The pooled sensitivity and specificity of the included radiomics studies were determined using statistical analysis. The pooled results are presented as forest plots. The overall predictive power was calculated by creating a summary receiver operating characteristic (SROC) curve. We evaluated the heterogeneity of the included literature by visually investigating the SROC curve [17]. The analysis was conducted using the R language [18], R package (Mada [19] and Meta [20]), and R studio [21].
Bias and study quality assessment
The publication bias was evaluated using a funnel plot. The quality of the included studies was assessed using the radiomics quality score (RQS) [22] and quality assessment of diagnostic accuracy studies (QUADAS-2) tool [23]. The RQS assessment investigated 16 components, which resulted in a score ranging from − 8 to 36, defined as 0% and 100%, respectively. The QUADAS-2 tool, which assesses seven components, was used to evaluate the risk of bias and applicability concerns. Two authors independently scored the RQS and QUADAS-2 tools. If a discrepancy was observed, the final score was discussed by the two authors to reach consensus.
Results
We retrieved a total of 682 articles. After removing duplicates, 118 articles were selected for evaluation. After screening for eligibility based on titles and abstracts, 12 articles were retrieved for complete evaluation. Four studies were excluded from the analysis as follows: one observational study [24], which used a repetitive patient population, one observational study [15], which used pulmonary opacities on chest images to predict disease severity, and two observational studies [25, 26], which used other severity assessment protocols to predict disease outcome. Finally, eight articles were used for qualitative analysis [27,28,29,30,31,32,33,34]. Only seven reports were included in the meta-analysis as a study by Li et al. [34] was excluded because only patients with severe COVID-19 were included in the report. A flowchart of the literature review is shown in Fig. 1. The details of the selected studies are presented in Table 1.
Pooled analysis of the included studies
Seven studies comprising 1460 patients with COVID-19 were included in this meta-analysis. The forest plot of pooled sensitivity was 0.800 (95% confidence interval [CI] = 0.662–0.891), as shown in Fig. 2. The forest plot of pooled specificity was 0.874 (95% CI = 0.773–0.934), as shown in Fig. 3. The pooled AUC was 0.908, and the SROC curve is shown in Fig. 4. We identified the heterogeneity within the included studies by visually investigating the SROC curve.
SROC, summary receiver operating characteristic curve; conf. region, 95% confidence region for the SROC curve.
Radiomics quality score of the included studies
The radiomics quality scores of the included studies are presented in Table 2. The radiomics quality scores ranged from 7 to 16. After a detailed evaluation of each RQS component by two authors, all included studies presented their image protocols, feature reduction performance, discrimination statistics reports, a comparison of the results to the gold standard, and potential clinical utility.
Qualities assessment of the selected literature
The QUADAS-2 tool was used to evaluate the literature. All studies had at least five out of seven low-risk bias assessment points. The results are presented in Fig. 5.
Publication bias assessment of the included studies
The funnel plot is shown in Fig. 6. As the number of included studies was less than 10, we cannot conclude whether a publication bias exists.
Review of the radiomics and clinical features used in the included studies
As stated by the IEEE International Symposium on Biomedical Imaging, there are many types of texture features, including first-order texture features, shape-based texture features, gray-level distance-zone matrix texture features, gray-level size-zone matrix texture features, neighborhood gray-tone difference matrix texture features, neighboring gray-level dependence matrix texture features, gray-level run-length matrix texture features, and gray-level co-occurrence matrix texture features [35]. The types of textural features used in the included studies are listed in Table 3. Four studies used shape-based radiomics features, six studies used first-order radiomics features, and five studies used second-order radiomics features.
Review of the prediction algorithms used in the included studies
Three selected studies used the least absolute shrinkage and selection operator (LASSO). One of the included studies used the XGBclassifier. Two of the studies used the random forest method. The other two studies used logistic regression, and the details of the prediction algorithms are listed in Table 4.
Discussion
Our meta-analysis revealed that CT-based radiomics could be used to predict the severity of COVID-19 pneumonia. In other CT-based radiomics studies, different COVID-19 pneumonia severity protocols could predict the severity of COVID-19 pneumonia [25, 26]. The management of COVID-19 pneumonia depends on disease severity [38, 39]. Therefore, early prediction of severe COVID-19 pneumonia before clinical deterioration using CT-based radiomics may aid in providing early management for these patients and reduce mortality [5, 40].
Our study included 1460 patients. The pooled sensitivity and specificity were 0.800 (95% CI = 0.662–0.891) and 0.874 (95% CI = 0.773–0.934), respectively. The pooled AUC was quite high at 0.908, indicating that radiomics is a promising tool for predicting the severity of COVID-19 pneumonia. The heterogeneity within the included studies may be attributed to the properties of radiomics features. As a previous study implied, radiomics features could be influenced by the calculation kernel, tumor delineation variability, technical settings of the CT scan, and software used to produce radiomics features [41]. This meta-analysis pooled results from various studies with different settings, thus providing robust results.
The RQS assessment resulted in a score ranging from −8 to 36, defined as 0% and 100%, respectively. The RQS values of the included literature ranged from seven to 16; thus, the highest RQS in the selected studies was only 40%. A previous meta-analysis also found a maximum RQS score of 16 for CT-based texture features used to differentiate between COVID-19 and viral pneumonia [14]. Compared with this study, a low RQS score makes it challenging to conduct a high-quality radiomics study in current research settings.
In contrast, the QUADAS-2 tool showed a favorable quality assessment of the selected studies. The risk of bias was primarily low in the selected studies, except for the patient selection bias. The patient selection bias was unclear or high because the selected studies were retrospective, and the patients were not randomly enrolled. The concern of applicability rating was low because the patient and index test interpretations were suitable for our review of the selected studies.
The types of radiomics features used in the selected studies should be discussed. While six studies assessed first-order features, five studies assessed second-order features, either alone or in combination with other features. Second-order features have been widely used in radiomics models for cancer patients, as they measure the heterogeneity within the region of interest. Hence, future studies investigating the molecular mechanisms associated with second-order radiomics features are warranted to deepen the understanding of COVID-19.
The algorithms used significantly varied between the selected studies. The most frequently used algorithm was the LASSO. The LASSO algorithm is a logistic regression-based algorithm that adds a regularization term to reduce the effect of noise on prediction. Another study used the XGBclassifier, a tree-based prediction algorithm that starts with a weak classifier and subsequently boosts to a stronger classifier [42]. Two of the included studies used the random forest method, another tree-based classifier, which starts with a robust classifier and reaches the final prediction result by voting [43]. The other two studies used traditional logistic regression models.
This meta-analysis had some limitations. First, the articles selected for this meta-analysis were retrospective. Second, the study protocols for each article were conducted in China, which can be attributed to our use of the CHC guidelines for COVID-19 pneumonia severity classification. Third, as this meta-analysis focused on predicting COVID-19 pneumonia severity using a CT-based radiomics learning model, the patients’ clinical data and disease course spectrum were not analyzed further. Although CT-based radiomics models were helpful for predicting COVID-19 pneumonia severity, the equivalence of pneumonia severity prediction to the prognosis and mortality prediction was not investigated in this meta-analysis. Therefore, future prospective and multicenter research should be performed to verify the effectiveness of radiomics in predicting COVID-19 pneumonia severity.
Conclusions
Our meta-analysis demonstrated that CT-based radiomics feature models might be powerful tools for predicting the severity of COVID-19 pneumonia.
References
Cucinotta D, Vanelli M (2020) WHO declares COVID-19 a pandemic. Acta Biomed 91(1):157–160
National Health Commission & National Administration of Traditional Chinese Medicine (2020) Diagnosis and treatment protocol for novel coronavirus pneumonia (Trial Version 7). Chin Med J 133(09):1087–1095
Wu Z, McGoogan JM (2020) Characteristics of and important lessons from the coronavirus disease 2019 (COVID-19) outbreak in china: summary of a report of 72 314 cases from the chinese center for disease control and prevention. JAMA 323(13):1239–1242
Mahendra M, Nuchin A, Kumar R, Shreedhar S, Mahesh PA (2021) Predictors of mortality in patients with severe COVID-19 pneumonia - a retrospective study. Adv Respir Med 89(2):135–144
Hashmi HAS, Asif HM (2020) Early detection and assessment of Covid-19. Front Med (Lausanne) 7:311
Pan Y, Guan H, Zhou S, Wang Y, Li Q, Zhu T, Hu Q, Xia L (2020) Initial CT findings and temporal changes in patients with the novel coronavirus pneumonia (2019-nCoV): a study of 63 patients in Wuhan. China Eur Radiol 30:3306–3309
Xiong Y, Sun D, Liu Y, Fan Y, Zhao L, Li X et al (2020) Clinical and high-resolution CT features of the COVID-19 infection: comparison of the initial and follow-up changes. Invest Radiol 55(6):332–339
Ai T, Yang Z, Hou H, Zhan C, Chen C, Lv W et al (2020) Correlation of chest CT and RT-PCR testing for coronavirus disease 2019 (COVID-19) in China: a report of 1014 cases. Radiology 296(2):E32–E40
Bao C, Liu X, Zhang H, Li Y, Liu J (2020) Coronavirus disease 2019 (COVID-19) CT findings: a systematic review and meta-analysis. J Am Coll Radiol 17(6):701–709
Liu Z, Wang S, Dong D, Wei J, Fang C, Zhou X et al (2019) The applications of radiomics in precision diagnosis and treatment of oncology: opportunities and challenges. Theranostics 9(5):1303–1322
Ursprung S, Beer L, Bruining A, Woitek R, Stewart GD, Gallagher FA et al (2020) Radiomics of computed tomography and magnetic resonance imaging in renal cell carcinoma-a systematic review and meta-analysis. Eur Radiol 30(6):3558–3566
Gülbay M, Özbay BO, Mendi BAR, Baştuğ A, Bodur H (2021) A CT radiomics analysis of COVID-19-related ground-glass opacities and consolidation: Is it valuable in a differential diagnosis with other atypical pneumonias? PLoS One 16(3):e0246582
Jin C, Chen W, Cao Y, Xu Z, Tan Z, Zhang X et al (2020) Development and evaluation of an artificial intelligence system for COVID-19 diagnosis. Nat Commun 11(1):5088
Kao YS, Lin KT (2021) A meta-analysis of computerized tomography-based radiomics for the diagnosis of COVID-19 and viral pneumonia. Diagnost (Basel) 11(6):991
Homayounieh F, Babaei R, Karimi Mobin H, Arru CD, Sharifian M, Mohseni I et al (2020) Computed tomography radiomics can predict disease severity and outcome in coronavirus disease 2019 pneumonia. J Comput Assist Tomogr 44(5):640–646
McInnes MD, Moher D, Thombs BD, McGrath TA, Bossuyt PM, Clifford T et al (2018) Preferred reporting items for a systematic review and meta-analysis of diagnostic test accuracy studies: the PRISMA-DTA statement. Jama 319(4):388–396
Lee J, Kim KW, Choi SH, Huh J, Park SH (2015) Systematic review and meta-analysis of studies evaluating diagnostic test accuracy: a practical review for clinical researchers-part II. Statistical methods of meta-analysis. Korean J Radiol 16(6):1188–1896
Haka R, Gentleman R (1996) R: a language for data analysis and graphics. J Comput Gr Statist 5(3):299–314
Doebler P, Holling H (2015) Meta-analysis of diagnostic accuracy with mada. R Packag 1:15
Schwarzer GJRN (2007) meta: an R package for meta-analysis. R News 7(3):40–45
Allaire JJB (2012) RStudio: integrated development environment for R. Boston MA 770(394):165–171
Lambin P, Leijenaar RTH, Deist TM, Peerlings J, de Jong EEC, van Timmeren J et al (2017) Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol 14(12):749–762
Whiting PF, Rutjes AW, Westwood ME, Mallett S, Deeks JJ, Reitsma JB et al (2011) QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med 155(8):529–536. https://doi.org/10.7326/0003-4819-155-8-201110180-00009
Tan HB, Xiong F, Jiang YL, Huang WC, Wang Y, Li HH et al (2020) The study of automatic machine learning base on radiomics of non-focus area in the first chest CT of different clinical types of COVID-19 pneumonia. Sci Rep 10(1):18926
Purkayastha S, Xiao Y, Jiao Z, Thepumnoeysuk R, Halsey K, Wu J et al (2021) Machine learning-based prediction of COVID-19 severity and progression to critical illness using CT imaging and clinical data. Korean J Radiol 22(7):1213–1224
Kang M, Hong KS, Chikontwe P, Luna M, Jang JG, Park J et al (2021) Quantitative assessment of chest CT patterns in COVID-19 and bacterial pneumonia patients: a deep learning perspective. J Korean Med Sci 36(5):e46
Xie Z, Sun H, Wang J, Xu H, Li S, Zhao C et al (2021) A novel CT-based radiomics in the distinction of severity of coronavirus disease 2019 (COVID-19) pneumonia. BMC Infect Dis 21(1):608
Li L, Wang L, Zeng F, Peng G, Ke Z, Liu H et al (2021) Development and multicenter validation of a CT-based radiomics signature for predicting severe COVID-19 pneumonia. Eur Radiol 31:7901–7912
Wang J, Xu Z, Cheng G, Hu Q, He L, Xia J et al (2020) Development and validation of a radiomics nomogram for identification of severity of patients with COVID-19. https://doi.org/10.21203/rs.3.rs-54606/v1
Xiong F, Wang Y, You T, Li HH, Fu TT, Tan H et al (2021) The clinical classification of patients with COVID-19 pneumonia was predicted by Radiomics using chest CT. Medicine (Baltimore) 100(12):e25307
Wei W, Hu XW, Cheng Q, Zhao YM, Ge YQ (2020) Identification of common and severe COVID-19: the value of CT texture analysis and correlation with clinical characteristics. Eur Radiol 30(12):6788–6796
Cai W, Liu T, Xue X, Luo G, Wang X, Shen Y et al (2020) CT Quantification and machine-learning models for assessment of disease severity and prognosis of COVID-19 patients. Acad Radiol 27(12):1665–1678
Tang Z, Zhao W, Xie X, Zhong Z, Shi F, Ma T et al (2021) Severity assessment of COVID-19 using CT image features and laboratory indices. Phys Med Biol 66(3):035015
Li C, Dong D, Li L, Gong W, Li X, Bai Y et al (2020) Classification of severe and critical covid-19 using deep learning and radiomics. IEEE J Biomed Health Inform 24(12):3585–3594
Zwanenburg A, Vallières M, Abdalah MA, Aerts H, Andrearczyk V, Apte A et al (2020) The image biomarker standardization initiative: standardized quantitative radiomics for high-throughput image-based phenotyping. Radiology 295(2):328–338
Chang YC, Yu CJ, Chang SC, Galvin JR, Liu HM, Hsiao CH et al (2005) Pulmonary sequelae in convalescent patients after severe acute respiratory syndrome: evaluation with thin-section CT. Radiology 236(3):1067–1075
Pan F, Ye T, Sun P, Gui S, Liang B, Li L et al (2020) Time course of lung changes at chest CT during recovery from coronavirus disease 2019 (COVID-19). Radiology 295(3):715–721
COVID-19 Treatment Guidelines. (2021) Available from: COVID-19 Treatment Guidelines Panel. Coronavirus Disease 2019 (COVID-19) Treatment Guidelines. National Institutes of Health. Available at https://www.covid19treatmentguidelines.nih.gov
Control CFD (2020) Prevention, Interim clinical guidance for management of patients with confirmed coronavirus disease (COVID-19). Available at https://stacks.cdc.gov/view/cdc/89980
Li T (2020) Diagnosis and clinical management of severe acute respiratory syndrome Coronavirus 2 (SARS-CoV-2) infection: an operational recommendation of Peking Union Medical College Hospital (V20). Emerg Microbes Infect 9(1):582–585
Park CM (2019) Can artificial intelligence fix the reproducibility problem of radiomics? Radiology 292(2):374–375
Chen T, Xgboost CG (2016) A scalable tree boosting system. In: Proceedings of the 22nd ACM SIGKDD international conference on knowledge discovery and data mining. pp 785–794
Breiman L (2001) Random Forests 45(1):5–32. https://link.springer.com/article/10.1023/A:1010933404324
Funding
This research did not receive external funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Human and animal rights
The study data were extracted from the included papers. The research was conducted in accordance with the 1964 Helsinki declaration and its amendments.
Informed consent
This meta-analysis was performed using data extracted from published papers. Informed consent was obtained from included papers.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kao, YS., Lin, KT. A meta-analysis of the diagnostic test accuracy of CT-based radiomics for the prediction of COVID-19 severity. Radiol med 127, 754–762 (2022). https://doi.org/10.1007/s11547-022-01510-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11547-022-01510-8