Abstract
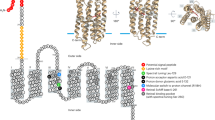
The breakthrough of environmental genomics of marine microbes has revealed the existence of eubacterial rhodopsin in the sea, named proteorhodopsin (PR), which can take light to produce bio-energy for cell metabolism. Gene and protein sequence analysis and laser flash-induced photolysis experiments have validated the function of PR as light-driven proton-pump. During the pumping process, light energy is transformed into chemical gradient potential across plasma inner-membrane, the potential energy is then used to synthesize ATP. The finding of PR actually brings to light a novel pathway of sunlight utilization existing in heterotrophic eubacteria in contrast to the well-known chlorophyll-dependent photosynthesis in the sea. Since the group of PR-bearing bacteria is one of the numerically richest microorganisms on the Earth, accounting for 13% of the total in sea surface water, and with averaged cellular PR molecules of 2.5×104, PR-bearing bacteria are a key component not to be ignored in energy metabolism and carbon cycling in the sea. Based on the understanding of current literature and our own investigation on PR in the China seas which indicated a ubiquitous presence and high diversity of PR in all the marine environments, we propose a conceptual model of energy flow and carbon cycling driven by both pigment-dependent and-independent biological utilization of light in the ocean.
Similar content being viewed by others
References
Fenchel T. Marine bugs and carbon flux. Science, 2001, 292: 2444–2445
Béjà O, Koonin E V, Aravind L, et al. Bacterial rhodopsin: evidence for a new type of phototrophy. Science, 2000, 289: 1902–1906
Friedrich T, Geibel S, Kalmbach R, et al. Proteorhodopsin is a light-driven proton pump with variable vectoriality. J Mol Biol, 2002, 321: 821–838
Sabehi G, Loy A, Jung K-H, et al. New insights into metabolic properties of marine bacteria encoding proteorhodopsins. PLoS Boil, 2005, 3(8): e273
Spudich J L, Yang C-S, Jung K-H, et al. Retinylidene proteins: Structures and functions from archaea to humans. Annu Rev Cell Dev Biol, 2000, 16: 365–392
Haupts U, Tittor J, Oesterhelt D. Closing in on bacteriorhodopsin: Progress in understanding the molecule. Annu Rev Biophys Biomol Struct, 1999, 28: 367–399
Bogomolni R A, Stoeckenius W, Szundi I, et al. Removal of transducer HtrI allows electrogenic proton translocation by sensory rhodopsin I. Proc Natl Acad Sci USA, 1994, 91: 10188–10192
Bieszke J A, Braun E L, Bean L E, et al. The nop-1 gene of Neurospora crassa encodes a seven transmembrane helix retinal-binding protein homologous to archaeal rhodopsins. Proc Natl Acad Sci USA, 1999, 96: 8034–8039
Sineshchekov O A, Jung K H, Spudich J L. Two rhodopsins mediate phototaxis to low-and high-intensity light in Chlamydomonas reinhardtii. Proc Natl Acad Sci USA, 2002, 99: 8689–8694
Jung K H, Trivedi V D, Spudich J L. Demonstration of a sensory rhodopsin in eubacteria. Mol Microbiol, 2003, 47: 1513–1522
Spudich J L, Jung K-H. Microbial rhodopsins: Phylogenetic and functional diversity. In: Briggs W R, Spudich J L, eds. Handbook of Photosensory Receptors. Weinheim: Wiley-VCH, 2005, 1–23
Waschuk S A, Bezerra A G Jr, Shi L, et al. Leptosphaeria rhodopsin: Bacteriorhodopsin-like proton pump from a eukaryote. Proc Natl Acad Sci USA, 2005, 102: 6879–6883
Béjà O, Koonin E V, Aravind L, et al. Construction and analysis of bacterial artificial chromosome libraries from a marine microbial assemblage. Environ Microbiol, 2000, 2: 516–529
Váró G, Brown L S, Lakatos M, et al. Characterization of the photochemical reaction cycle of proteorhodopsin. Biophysi, 2003, 84: 1202–1207
Béjà O, Aravind L, Eugene V, et al. Proteorhodopsin phototrophy in the ocean. Nature, 2001, 411: 786–789
Giovannoni S J, Bibbs L, Cho J C, et al. Proteorhodopsin in the ubiquitous marine bacterium SAR11. Nature, 2005, 438: 82–85
Sabehi G, Massana R, Bielawski J P, et al. Novel proteorhodopsin variants from the Mediterranean and Red Seas. Environ Microbiol, 2003, 5: 812–849
Bielawski J P, Dunn K A, Sabehi G. Darwinian adaptation of proteorhodopsin to different light intensities in the marine environment. Proc Natl Acad Sci USA, 2004, 101: 14824–14829
Venter J C, Remington K, Heidelberg J F, et al. Environmental genome shotgun sequencing of the Sargasso Sea. Science, 2004, 306: 66–74
Kyndt J, Meyer T E, Cusanovich M A. Photoactive yellow protein bacteriophytochrome, and sensory rhodopsin in purple phototrophic bacteria. Photochem Photobiol Sci, 2004, 3: 519–530
Torre J R, Christianson L M, Béjà O, et al. Proteorhodopsin genes are distributed in divergent marine bacterial taxa. Proc Natl Acad Sci USA, 2003, 100: 12830–12835
Sabehi G, Béjà O, Suzuki M T, et al. Different SAR86 subgroups harbour divergent proteorhodopsins. Environ Microbiol, 2004, 6: 903–910
Rappé M S, Connon S A, Vergin K L, et al. Cultivation of the ubiquitous SAR11 marine bacterioplanton clade. Nature, 2002, 418: 630–633
Giovannoni S J, Tripp H J, Givan S, et al. Genome streamlining in a cosmopolitan oceanic bacterium. Science, 2005, 309(5738): 1242–1245
Man D, Wang W W, Sabehi G, et al. Diversification and spectral tuning in marine proteorhodopsins. EMBO J, 2003, 22: 1725–1731
Wang W W, Sineshchekov O A, Spudich E N, et al. Spectroscopic and photochemical characterization of a deep ocean proteorhodopsin. J Biol Chem, 2003, 278: 33985–33991
Man-Aharonovich D, Sabehi G, Sineshchekov O A, et al. Characterization of RS29, a blue-green proteorhodopsin variant from the Red sea. Photochem Photobiol Sci, 2004, 3: 459–462
Hoff W D, Jung K H, Spudich J L. Molecular mechanism of photosignaling by archaeal sensory rhodopsins. Annu Rev Biophys Biomol Struct, 1997, 26: 223–258
Haupts U, Haupts C, Oesterhelt D. The photoreceptor sensory rhodopsin I as a two-photon-driven proton pump. Proc Natl Acad Sci USA, 1995, 92: 3834–3838
Michel H, Oesterhelt D. Electrochemical proton gradient across the cell membrane of Halobacterium halobium: Effect of N,N9-dicyclohexylcarbodiimide, relation to intracellular adenosine triphosphate, adenosine diphosphate, and phosphate concentration, and influenceof the potassium gradient. Biochemistry, 1980, 19: 4607–4614
Tsutomu M, Koichi K. Quantum conversion and image detection by a bacteriorhodopsin-based artificial photoreceptor. Science, 1992, 255: 342–344
Kennedy D. Breakthrough of the year. Science, 2004, 306: 2001
Jiao N Z, Sieracki M E, Zhang Y, et al. Aerobic anoxygenic phototrophic bacteria and their roles in marine ecosystems. Chin Sci Bull, 48(11): 1064–1068
Jiao N Z, Zhang Y, Chen Y. Time series observation based Infrared epifluorescence microscopic approach for accurate enumeration of bacteriochlorophyll containing microbes in Marine Environments. J Microbiol Methods, 2006, 65(3): 428–439
Hampp N, Oesterhelt D. Bacteriorhodopsin and its potential in technical applications. In: Niemeyer C, Mirkin C, eds. Nano-biotechnology. Weinheim: Wiley-VCH-Verlag, 2004, 146–167
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Jiao, N., Feng, F. & Wei, B. Proteorhodopsin—A new path for biological utilization of light energy in the sea. CHINESE SCI BULL 51, 889–896 (2006). https://doi.org/10.1007/s11434-008-0889-x
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s11434-008-0889-x