Abstract
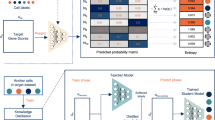
The rapid accumulation of large-scale single-cell RNA-seq datasets from multiple institutions presents remarkable opportunities for automatically cell annotations through integrative analyses. However, the privacy issue has existed but being ignored, since we are limited to access and utilize all the reference datasets distributed in different institutions globally due to the prohibited data transmission across institutions by data regulation laws. To this end, we present scPrivacy, which is the first and generalized automatically single-cell type identification prototype to facilitate single cell annotations in a data privacy-preserving collaboration manner. We evaluated scPrivacy on a comprehensive set of publicly available benchmark datasets for single-cell type identification to stimulate the scenario that the reference datasets are rapidly generated and distributed in multiple institutions, while they are prohibited to be integrated directly or exposed to each other due to the data privacy regulations, demonstrating its effectiveness, time efficiency and robustness for privacy-preserving integration of multiple institutional datasets in single cell annotations.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Data availability
The 27 single-cell type identification benchmark datasets and 15 patients datasets were curated from five studies including three tissues: peripheral blood mononuclear cells (PBMCs) (Ding et al., 2019; Mereu et al., 2020; Ren et al., 2021), the brain (Tasic et al., 2016; Tasic et al., 2018) and the pancreas (Baron et al., 2016; Muraro et al., 2016; Segerstolpe et al., 2016; Xin et al., 2016) (Tables S1 and S12 in Supporting Information). The four pancreas datasets (Baron et al., 2016; Muraro et al., 2016; Segerstolpe et al., 2016; Xin et al., 2016) and one of the brain datasets (Tasic et al., 2018) were collected in previous work of scmap (Kiselev et al., 2018) (https://hemberg-lab.github.io/scRNA.seq.datasets), and the other three brain datasets and seven datasets in “PBMC-Ding” (Ding et al., 2019) were curated from the following benchmark study (Abdelaal et al., 2019) (https://doi.org/10.5281/zenodo.3357167). The 12 datasets in “PBMC-Mereu” (Mereu et al., 2020) were collected from GSE133549, and the corresponding RData file can be downloaded in https://www.dropbox.com/s/i8mwmyymchx8mn8/sce.all_classified.technologies.RData?dl=0. All these datasets were converted into Bioconductor SingleCellExperiment (http://bioconductor.org/packages/SingleCellExperiment) class objects. The 15 datasets of COVID-19 patients were collected from GSE158055 (Table S12 in Supporting Information).
References
Abdelaal, T., Michielsen, L., Cats, D., Hoogduin, D., Mei, H., Reinders, M. J.T., and Mahfouz, A. (2019). A comparison of automatic cell identification methods for single-cell RNA sequencing data. Genome Biol 20, 194.
Acar, A., Aksu, H., Uluagac, A.S., and Conti, M. (2019). A survey on homomorphic encryption schemes. ACM Comput Surv 51, 1–35.
Aran, D., Looney, A.P., Liu, L., Wu, E., Fong, V., Hsu, A., Chak, S., Naikawadi, R.P., Wolters, P.J., Abate, A.R., et al. (2019). Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat Immunol 20, 163–172.
Baron, M., Veres, A., Wolock, S.L., Faust, A.L., Gaujoux, R., Vetere, A., Ryu, J.H., Wagner, B.K., Shen-Orr, S.S., Klein, A.M., et al. (2016). A single-cell transcriptomic map of the human and mouse pancreas reveals inter- and intra-cell population structure. Cell Syst 3, 346–360.e4.
Benefield, H., Ashkanazi, G., and Rozensky, R.H. (2006). Communication and records: hippa issues when working in health care settings. Prof Psychol-Res Pract 37, 273–277.
Byrd, J.B., Greene, A.C., Prasad, D.V., Jiang, X., and Greene, C.S. (2020). Responsible, practical genomic data sharing that accelerates research. Nat Rev Genet 21, 615–629.
Chen, S., Luo, Y., Gao, H., Li, F., Chen, Y., Li, J., You, R., Hao, M., Bian, H., Xi, X., et al. (2022a). hECA: the cell-centric assembly of a cell atlas. iScience 25, 104318.
Chen, S., Luo, Y., Gao, H., Li, F., Li, J., Chen, Y., You, R., Lv, H., Hua, K., Jiang, R., et al. (2022b). Toward a unified information framework for cell atlas assembly. Natl Sci Rev 9, nwab179.
Chen, S., Xue, D., Chuai, G., Yang, Q., and Liu, Q. (2021). FL-QSAR: a federated learning-based QSAR prototype for collaborative drug discovery. Bioinformatics 36, 5492–5498.
Ding, J., Adiconis, X., Simmons, S.K., Kowalczyk, M.S., Hession, C.C., Marjanovic, N.D., Hughes, T.K., Wadsworth, M.H., Burks, T., Nguyen, L.T., Kwon, J.Y.H., Barak, B., Ge, W., Kedaigle, A.J., Carroll, S., Li, S., Hacohen, N., Rozenblatt-Rosen, O., Shalek, A.K., Villani, A.-C., Regev, A., and Levin, J.Z. (2019). Systematic comparative analysis of single cell RNA-sequencing methods. bioRxiv, 632216.
Domínguez Conde, C., Xu, C., Jarvis, L.B., Rainbow, D.B., Wells, S.B., Gomes, T., Howlett, S.K., Suchanek, O., Polanski, K., King, H.W., et al. (2022). Cross-tissue immune cell analysis reveals tissue-specific features in humans. Science 376, eabl5197.
Duan, B., Chen, S., Chen, X., Zhu, C., Tang, C., Wang, S., Gao, Y., Fu, S., and Liu, Q. (2021). Integrating multiple references for single-cell assignment. Nucl Acids Res 49, e80.
Duan, B., Zhu, C., Chuai, G., Tang, C., Chen, X., Chen, S., Fu, S., Li, G., and Liu, Q. (2020). Learning for single-cell assignment. Sci Adv 6, eabd0855.
Elmentaite, R., Ross, A.D.B., Roberts, K., James, K.R., Ortmann, D., Gomes, T., Nayak, K., Tuck, L., Pritchard, S., Bayraktar, O.A., et al. (2020). Single-cell sequencing of developing human gut reveals transcriptional links to childhood crohn’s disease. Dev Cell 55, 771–783.e5.
Eraslan, G., Drokhlyansky, E., Anand, S., Fiskin, E., Subramanian, A., Slyper, M., Wang, J., Van Wittenberghe, N., Rouhana, J.M., Waldman, J., et al. (2022). Single-nucleus cross-tissue molecular reference maps toward understanding disease gene function. Science 376, eabl4290.
Guan, Y.N., Li, Y., Roosan, M., and Jing, Q. (2021). Single-cell transcriptomics of murine mural cells reveals cellular heterogeneity. Sci China Life Sci 64, 1077–1086.
Halamka, J.D., and Tripathi, M. (2017). The HITECH era in retrospect. N Engl J Med 377, 907–909.
Jiang, H., Zhang, H., and Zhang, X. (2021). Single-cell genomic profile-based analysis of tissue differentiation in colorectal cancer. Sci China Life Sci 64, 1311–1325.
Kiselev, V.Y., Yiu, A., and Hemberg, M. (2018). scmap: projection of single-cell RNA-seq data across data sets. Nat Methods 15, 359–362.
Li, C., Liu, B., Kang, B., Liu, Z., Liu, Y., Chen, C., Ren, X., and Zhang, Z. (2020). SciBet as a portable and fast single cell type identifier. Nat Commun 11, 1818.
Liu, J., Li, J., Wang, H., and Yan, J. (2020). Application of deep learning in genomics. Sci China Life Sci 63, 1860–1878.
Liu, Z., and Zhang, Z. (2022). Mapping cell types across human tissues. Science 376, 695–696.
Lotfollahi, M., Naghipourfar, M., Luecken, M.D., Khajavi, M., Büttner, M., Wagenstetter, M., Avsec, Ž., Gayoso, A., Yosef, N., Interlandi, M., et al. (2022). Mapping single-cell data to reference atlases by transfer learning. Nat Biotechnol 40, 121–130.
Ma, F., and Pellegrini, M. (2020). ACTINN: automated identification of cell types in single cell RNA sequencing. Bioinformatics 36, 533–538.
McKeen, F., Alexandrovich, I., Anati, I., Caspi, D., Johnson, S., Leslie-Hurd, R., and Rozas, C. (2016). Intel® Software Guard Extensions (Intel® SGX) Support for Dynamic Memory Management Inside an Enclave. In Proceedings of the Hardware and Architectural Support for Security and Privacy 2016 on — HASP 2016, pp. 1–9.
McMahan, H.B., Moore, E., Ramage, D., and Hampson, S. (2016). Communication-efficient learning of deep networks from decentralized data. arXiv preprint.
Mereu, E., Lafzi, A., Moutinho, C., Ziegenhain, C., McCarthy, D.J., Álvarez-Varela, A., Batlle, E., Sagar, E., Grün, D., Lau, J.K., et al. (2020). Benchmarking single-cell RNA-sequencing protocols for cell atlas projects. Nat Biotechnol 38, 747–755.
Muraro, M.J., Dharmadhikari, G., Grün, D., Groen, N., Dielen, T., Jansen, E., van Gurp, L., Engelse, M.A., Carlotti, F., de Koning, E.J.P., et al. (2016). A single-cell transcriptome atlas of the human pancreas. Cell Syst 3, 385–394.e3.
Papatheodorou, I., Moreno, P., Manning, J., Fuentes, A.M.P., George, N., Fexova, S., Fonseca, N.A., Füllgrabe, A., Green, M., Huang, N., et al. (2019). Expression Atlas update: from tissues to single cells. Nucl Acids Res 48, D77–D83.
Paszke, A., Gross, S., Massa, F., Lerer, A., Bradbury, J., Chanan, G., Killeen, T., Lin, Z., Gimelshein, N., and Antiga, L. (2019). PyTorch: an imperative style, high-performance deep learning library. Paper presented at: Advances in Neural Information Processing Systems. (New York: ACM), pp. 8026–8037.
Plass, M., Solana, J., Wolf, F.A., Ayoub, S., Misios, A., Glažar, P., Obermayer, B., Theis, F.J., Kocks, C., and Rajewsky, N. (2018). Cell type atlas and lineage tree of a whole complex animal by single-cell transcriptomics. Science 360.
Politou, E., Alepis, E., and Patsakis, C. (2018). Forgetting personal data and revoking consent under the GDPR: Challenges and proposed solutions. J Cybersecur 4.
Regev, A., Teichmann, S.A., Lander, E.S., Amit, I., Benoist, C., Birney, E., Bodenmiller, B., Campbell, P., Carninci, P., Clatworthy, M., et al. (2017). The human cell atlas. eLife 6, e27041.
Ren, X., Wen, W., Fan, X., Hou, W., Su, B., Cai, P., Li, J., Liu, Y., Tang, F., Zhang, F., et al. (2021). COVID-19 immune features revealed by a large-scale single-cell transcriptome atlas. Cell 184, 1895–1913.e19.
Rozenblatt-Rosen, O., Regev, A., Oberdoerffer, P., Nawy, T., Hupalowska, A., Rood, J.E., Ashenberg, O., Cerami, E., Coffey, R.J., Demir, E., et al. (2020). The human tumor atlas network: charting tumor transitions across space and time at single-cell resolution. Cell 181, 236–249.
Saldanha, O.L., Quirke, P., West, N.P., James, J.A., Loughrey, M.B., Grabsch, H.I., Salto-Tellez, M., Alwers, E., Cifci, D., Ghaffari Laleh, N., et al. (2022). Swarm learning for decentralized artificial intelligence in cancer histopathology. Nat Med 28, 1232–1239.
Segerstolpe, Å., Palasantza, A., Eliasson, P., Andersson, E.M., Andréasson, A.C., Sun, X., Picelli, S., Sabirsh, A., Clausen, M., Bjursell, M.K., et al. (2016). Single-cell transcriptome profiling of human pancreatic islets in health and type 2 diabetes. Cell Metab 24, 593–607.
Snyder, M.P., Lin, S., Posgai, A., Atkinson, M., Regev, A., Rood, J., Rozenblatt-Rosen, O., Gaffney, L., Hupalowska, A., Satija, R., et al. (2019). The human body at cellular resolution: the NIH Human Biomolecular Atlas Program. Nature 574, 187–192.
Sohn, K. (2016). Improved deep metric learning with multi-class N-pair loss objective. Adv Neur In 29.
Stuart, T., Butler, A., Hoffman, P., Hafemeister, C., Papalexi, E., Mauck Iii, W.M., Hao, Y., Stoeckius, M., Smibert, P., and Satija, R. (2019). Comprehensive integration of single-cell data. Cell 177, 1888–1902. e21.
Suo, C., Dann, E., Goh, I., Jardine, L., Kleshchevnikov, V., Park, J.E., Botting, R.A., Stephenson, E., Engelbert, J., Tuong, Z.K., et al. (2022). Mapping the developing human immune system across organs. Science 376.
Jones, R.C., Karkanias, J., Krasnow, M.A., Pisco, A.O., Quake, S.R., Salzman, J., Yosef, N., Bulthaup, B., Brown, P., Harper, W., et al. (2022). The Tabula Sapiens: a multiple-organ, single-cell transcriptomic atlas of humans. Science 376, eabl4896.
Tasic, B., Menon, V., Nguyen, T.N., Kim, T.K., Jarsky, T., Yao, Z., Levi, B., Gray, L.T., Sorensen, S.A., Dolbeare, T., et al. (2016). Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat Neurosci 19, 335–346.
Tasic, B., Yao, Z., Graybuck, L.T., Smith, K.A., Nguyen, T.N., Bertagnolli, D., Goldy, J., Garren, E., Economo, M.N., Viswanathan, S., et al. (2018). Shared and distinct transcriptomic cell types across neocortical areas. Nature 563, 72–78.
Travaglini, K.J., Nabhan, A.N., Penland, L., Sinha, R., Gillich, A., Sit, R. V., Chang, S., Conley, S.D., Mori, Y., Seita, J., et al. (2020). A molecular cell atlas of the human lung from single-cell RNA sequencing. Nature 587, 619–625.
Warnat-Herresthal, S., Schultze, H., Shastry, K.L., Manamohan, S., Mukherjee, S., Garg, V., Sarveswara, R., Händler, K., Pickkers, P., Aziz, N.A., et al. (2021). Swarm learning for decentralized and confidential clinical machine learning. Nature 594, 265–270.
Winnubst, J., and Arber, S. (2021). A census of cell types in the brain’s motor cortex. Nature 598, 33–34.
Xie, X., Cheng, X., Wang, G., Zhang, B., Liu, M., Chen, L., Cheng, H., Hao, S., Zhou, J., Zhu, P., et al. (2021). Single-cell transcriptomes of peripheral blood cells indicate and elucidate severity of COVID-19. Sci China Life Sci 64, 1634–1644.
Xin, Y., Kim, J., Okamoto, H., Ni, M., Wei, Y., Adler, C., Murphy, A.J., Yancopoulos, G.D., Lin, C., and Gromada, J. (2016). RNA sequencing of single human islet cells reveals type 2 diabetes genes. Cell Metab 24, 608–615.
Yang, Q., Liu, Y., Chen, T., and Tong, Y. (2019). Federated machine learning: Concept and applications. ACM Transactions on Intelligent Systems and Technology (TIST) 10, 1–19.
Yao, A.C. (1982). Protocols for secure computations. In: Proceedings of the 23rd Annual IEEE Symposium on Foundations of Computer Science.
Zhang, Y., and Yang, Q. (2018). An overview of multi-task learning. Natl Sci Rev 5, 30–43.
Zhao, Y., Wang, T., Liu, Z., Ke, Y., Li, R., Chen, H., You, Y., Wu, G., Cao, S., Du, Z., et al. (2022). Single-cell transcriptomics of immune cells in lymph nodes reveals their composition and alterations in functional dynamics during the early stages of bubonic plague. Sci China Life Sci, doi: https://doi.org/10.1007/s11427-021-2119-5.
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2021YFF1200900, 2021YFF1201200), the National Natural Science Foundation of China (31970638, 61572361), the Shanghai Artificial Intelligence Technology Standard Project (19DZ2200900), the Shanghai Shuguang Scholars Project, WeBank Scholars Project and the Fundamental Research Funds for the Central Universities.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Compliance and ethics
The author(s) declare that they have no conflict of interest.
Code availability
scPrivacy is developed as a python package for simulations, which is available at https://github.com/bm2-lab/scPrivacy.
Rights and permissions
About this article
Cite this article
Chen, S., Duan, B., Zhu, C. et al. Privacy-preserving integration of multiple institutional data for single-cell type identification with scPrivacy. Sci. China Life Sci. 66, 1183–1195 (2023). https://doi.org/10.1007/s11427-022-2224-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-022-2224-4