Abstract
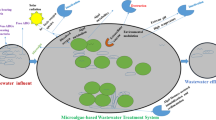
Antibiotics have become a concern in the aquatic environments owing to the potential development of bacterial resistances. Thus, this study evaluated the removal of cephalexin (CEP) and erythromycin (ERY) from a local wastewater treatment plant (WWTP) effluent, mediated by microalgae-bacteria consortium. Likewise, the removal of correlated antibiotics resistance genes blaTEM and ermB was also assessed. The incubation results showed that the added concentrations of selected antibiotics did not restrain the consortium growth. Moreover, CEP and ERY were almost completely removed after the cultivation period, reaching total removals of 96.54% and 92.38%, respectively. The symbiotic interaction between microalgae and bacteria plays a role in the kinetics removal of CEP and ERY. The abundance of blaTEM and ermB was reduced by 0.56 and 1.75 logs, respectively. Lastly, our results suggest that technology based on natural microalgae-bacteria consortium could be a potential alternative to improve the quality of WWTP effluents.





Similar content being viewed by others
References
APHA (2012) Standard methods for examination of water and wastewater. American Public Health Association/American Water Works Association/Water Environment Federation, Washington, DC, USA. Water Environmental Federation
da Silva Rodrigues DA, da Cunha CCRF, Freitas MG, de Barros ALC, e Castro PBN, Pereira AR, de Queiroz Silva S, da Fonseca Santiago A, Afonso RJCF (2020) Biodegradation of sulfamethoxazole by microalgae-bacteria consortium in wastewater treatment plant effluents. Science of The Total Environment 749:141441. https://doi.org/10.1016/j.scitotenv.2020.141441
Fan C, He J (2011) Proliferation of antibiotic resistance genes in microbial consortia of sequencing batch reactors (SBRs) upon exposure to trace erythromycin or erythromycin-H2O. Water research 45:3098–3106. https://doi.org/10.1016/j.watres.2011.03.025
Ferrando L, Matamoros V (2020) Attenuation of nitrates, antibiotics and pesticides from groundwater using immobilised microalgae-based systems. Science of The Total Environment 703:134740. https://doi.org/10.1016/j.scitotenv.2019.134740
Ferris MJ, Muyzer G, Ward DM (1996) Denaturing gradient gel electrophoresis profiles of 16S rRNA-defined populations inhabiting a hot spring microbial mat community. Applied and Environmental Microbiology 62:340–346
Furlan JPR, Dos Santos LDR, Moretto JAS, Ramos MS, Gallo IFL, Alves GAD, Paulelli AC, de Souza Rocha CC, Cesila CA, Gallimberti M (2020) Occurrence and abundance of clinically relevant antimicrobial resistance genes in environmental samples after the Brumadinho dam disaster, Brazil. Science of The Total Environment 726:138100. https://doi.org/10.1016/j.scitotenv.2020.138100
Gomes RP, Pena CB, Rezende J, Coutrim MX, Afonso RJCF (2017) Validation of a new high-throughput method to determine urinary S-phenylmercapturic acid using low-temperature partitioning extraction and ultra high performance liquid chromatography–mass spectrometry. Journal of Separation Science 40:550–557. https://doi.org/10.1002/jssc.201600540
Gros M, Rodríguez-Mozaz S, Barceló D (2013) Rapid analysis of multiclass antibiotic residues and some of their metabolites in hospital, urban wastewater and river water by ultra-high-performance liquid chromatography coupled to quadrupole-linear ion trap tandem mass spectrometry. Journal of Chromatography A 1292:173–188. https://doi.org/10.1016/j.chroma.2012.12.072
Gulkowska A, Leung HW, So MK, Taniyasu S, Yamashita N, Yeung LW, Richardson BJ, Lei A, Giesy JP, Lam PK (2008) Removal of antibiotics from wastewater by sewage treatment facilities in Hong Kong and Shenzhen, China. Water Research 42:395–403. https://doi.org/10.1016/j.watres.2007.07.031
Hu Y, Jiang L, Zhang T, Jin L, Han Q, Zhang D, Lin K, Cui C (2018) Occurrence and removal of sulfonamide antibiotics and antibiotic resistance genes in conventional and advanced drinking water treatment processes. Journal of hazardous materials 360:364–372. https://doi.org/10.1016/j.jhazmat.2018.08.012
Jaén-Gil A, Hom-Diaz A, Llorca M, Vicent T, Blánquez P, Barceló D, Rodríguez-Mozaz S (2018) An automated on-line turbulent flow liquid-chromatography technology coupled to a high resolution mass spectrometer LTQ-Orbitrap for suspect screening of antibiotic transformation products during microalgae wastewater treatment. Journal of Chromatography A 1568:57–68. https://doi.org/10.1016/j.chroma.2018.06.027
Jiang L, Hu X, Xu T, Zhang H, Sheng D, Yin D (2013) Prevalence of antibiotic resistance genes and their relationship with antibiotics in the Huangpu River and the drinking water sources, Shanghai, China. Science of the Total Environment 458:267–272. https://doi.org/10.1016/j.scitotenv.2013.04.038
Kiki C, Rashid A, Wang Y, Li Y, Zeng Q, Yu C-P, Sun Q (2020) Dissipation of antibiotics by microalgae: kinetics, identification of transformation products and pathways. Journal of Hazardous Materials 387:121985. https://doi.org/10.1016/j.jhazmat.2019.121985
Le-Minh N, Khan S, Drewes J, Stuetz R (2010) Fate of antibiotics during municipal water recycling treatment processes. Water research 44:4295–4323. https://doi.org/10.1016/j.watres.2010.06.020
Li B, Zhang T, Xu Z, Fang HHP (2009) Rapid analysis of 21 antibiotics of multiple classes in municipal wastewater using ultra performance liquid chromatography-tandem mass spectrometry. Analytica Chimica Acta 645:64–72. https://doi.org/10.1016/j.aca.2009.04.042
Liang X, Guan F, Chen B, Luo P, Guo C, Wu G, Ye Y, Zhou Q, Fang H (2020) Spatial and seasonal variations of antibiotic resistance genes and antibiotics in the surface waters of Poyang Lake in China. Ecotoxicology and Environmental Safety 196:110543. https://doi.org/10.1016/j.ecoenv.2020.110543
Luo Y, Guo W, Ngo HH, Nghiem LD, Hai FI, Zhang J, Liang S, Wang XC (2014) A review on the occurrence of micropollutants in the aquatic environment and their fate and removal during wastewater treatment. Science of the Total Environment 473:619–641. https://doi.org/10.1016/j.scitotenv.2013.12.065
Manaia CM, Graham D, Topp E, Martinez JL, Collignon P, Gaze WH (2020): Antibiotic resistance in the environment: Expert Perspectives.
Mao D, Yu S, Rysz M, Luo Y, Yang F, Li F, Hou J, Mu Q, Alvarez P (2015) Prevalence and proliferation of antibiotic resistance genes in two municipal wastewater treatment plants. Water research 85:458–466. https://doi.org/10.1016/j.watres.2015.09.010
Meng L, Wang J, Li X (2020) Insight into effect of high-level cephalexin on fate and driver mechanism of antibiotics resistance genes in antibiotic wastewater treatment system. Ecotoxicology and Environmental Safety 201:110739. https://doi.org/10.1016/j.ecoenv.2020.110739
Neves e Castro PB, da Silva Rodrigues DA, Roeser HMP, da Fonseca Santiago A, de Cássia Franco Afonso RJ (2020): Antibiotic consumption in developing countries defies global commitments: an overview on Brazilian growth in consumption. Environmental Science and Pollution Research https://doi.org/10.1007/s11356-020-08574-x
Nush (1981): Nederlands Norm-NEN 6520. Water: spectrophotometric determination of chlorophyll a content
Pomati F, Netting AG, Calamari D, Neilan BA (2004) Effects of erythromycin, tetracycline and ibuprofen on the growth of Synechocystis sp. and Lemna minor. Aquatic Toxicology 67:387–396. https://doi.org/10.1016/j.aquatox.2004.02.001
Pugajeva I, Rusko J, Perkons I, Lundanes E, Bartkevics V (2017) Determination of pharmaceutical residues in wastewater using high performance liquid chromatography coupled to quadrupole-Orbitrap mass spectrometry. Journal of Pharmaceutical and Biomedical Analysis 133:64–74. https://doi.org/10.1016/j.jpba.2016.11.008
Qi C, Liu X, Lin C, Zhang X, Ma J, Tan H, Ye W (2014) Degradation of sulfamethoxazole by microwave-activated persulfate: kinetics, mechanism and acute toxicity. Chemical Engineering Journal 249:6–14. https://doi.org/10.1016/j.cej.2014.03.086
Ramsundar P, Guldhe A, Singh P, Bux F (2017) Assessment of municipal wastewaters at various stages of treatment process as potential growth media for Chlorella sorokiniana under different modes of cultivation. Bioresource technology 227:82–92. https://doi.org/10.1016/j.biortech.2016.12.037
Rodriguez-Mozaz S, Chamorro S, Marti E, Huerta B, Gros M, Sànchez-Melsió A, Borrego CM, Barceló D, Balcázar JL (2015) Occurrence of antibiotics and antibiotic resistance genes in hospital and urban wastewaters and their impact on the receiving river. Water research 69:234–242. https://doi.org/10.1016/j.watres.2014.11.021
Rossi S, Sforza E, Pastore M, Bellucci M, Casagli F, Marazzi F, Ficara E (2020) Photo-respirometry to shed light on microalgae-bacteria consortia—a review. Reviews in Environmental Science and Bio/Technology 19:43–72. https://doi.org/10.1007/s11157-020-09524-2
Saravanane R, Sundararaman S (2009) Effect of loading rate and HRT on the removal of cephalosporin and their intermediates during the operation of a membrane bioreactor treating pharmaceutical wastewater. Environmental technology 30:1017–1022. https://doi.org/10.1080/09593330903032865
Szekeres E, Baricz A, Chiriac CM, Farkas A, Opris O, Soran M-L, Andrei A-S, Rudi K, Balcázar JL, Dragos N (2017) Abundance of antibiotics, antibiotic resistance genes and bacterial community composition in wastewater effluents from different Romanian hospitals. Environmental Pollution 225:304–315. https://doi.org/10.1016/j.envpol.2017.01.054
Tao C-W, Hsu B-M, Ji W-T, Hsu T-K, Kao P-M, Hsu C-P, Shen S-M, Shen T-Y, Wan T-J, Huang Y-L (2014) Evaluation of five antibiotic resistance genes in wastewater treatment systems of swine farms by real-time PCR. Science of the Total Environment 496:116–121. https://doi.org/10.1016/j.scitotenv.2014.07.024
Wang Y, Ho S-H, Cheng C-L, Guo W-Q, Nagarajan D, Ren N-Q, Lee D-J, Chang J-S (2016) Perspectives on the feasibility of using microalgae for industrial wastewater treatment. Bioresource Technology 222:485–497. https://doi.org/10.1016/j.biortech.2016.09.106
WHO (2014): World Health Organization - Antimicrobial resistance: Global Report on Surveillance. World Health Organization, http://apps.who.int/bookorders/anglais/detart1.jsp?codlan=1&codcol=15&codcch=870
Wu D-L, Zhang M, He L-X, Zou H-Y, Liu Y-S, Li B-B, Yang Y-Y, Liu C, He L-Y, Ying G-G (2020) Contamination profile of antibiotic resistance genes in ground water in comparison with surface water. Science of The Total Environment 715:136975. https://doi.org/10.1016/j.scitotenv.2020.136975
Xiong J-Q, Kurade MB, Jeon B-H (2016) Biodegradation of levofloxacin by an acclimated freshwater microalga, Chlorella vulgaris. Chemical Engineering Journal 313:1251–1257. https://doi.org/10.1016/j.cej.2016.11.017
Xu L, Ouyang W, Qian Y, Su C, Su J, Chen H (2016a) High-throughput profiling of antibiotic resistance genes in drinking water treatment plants and distribution systems. Environmental Pollution 213:119–126. https://doi.org/10.1016/j.envpol.2016.02.013
Xu Y, Guo C, Luo Y, Lv J, Zhang Y, Lin H, Wang L, Xu J (2016b) Occurrence and distribution of antibiotics, antibiotic resistance genes in the urban rivers in Beijing, China. Environmental pollution 213:833–840. https://doi.org/10.1016/j.envpol.2016.03.054
Yao S, Lyu S, An Y, Lu J, Gjermansen C, Schramm A (2019) Microalgae–bacteria symbiosis in microalgal growth and biofuel production: a review. Journal of applied microbiology 126:359–368. https://doi.org/10.1111/jam.14095
Zhu S, Huo S, Feng P (2019): Developing designer microalgal consortia: a suitable approach to sustainable wastewater treatment, microalgae biotechnology for development of biofuel and wastewater treatment. Springer, pp. 569-598
Acknowledgements
The authors would like to thank the Federal University of Ouro Preto (UFOP), Multicenter Postgraduation Program in Chemistry - Minas Gerais (PPGMQ-MG), and the Autonomous Water and Sewage Service (SAAE) in Itabirito District, Minas Gerais State, Brazil.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Funding
This study received financial support from the Coordination for Improvement of Higher-Level Personnel (CAPES), the Foundation for Research Support of the Minas Gerais State (FAPEMIG), the National Research Council of Brazil (CNPq), the Research and Projects Financier (FINEP), the Brazilian National Health Foundation (FUNASA), and the Federal University of Ouro Preto (UFOP).
Author information
Authors and Affiliations
Contributions
DASR: Conceptualization, methodology, investigation, and writing—original draft
CCRFC: Methodology and investigation
DRES: Methodology and investigation
ALCB: Methodology, investigation, and formal analysis
ARP: Methodology and investigation
SQS: Resources and writing—review and editing
AFS: Conceptualization, resources, visualization, and writing—review and editing
RJCFA: Conceptualization, writing—review and editing—project administration, funding acquisition, and supervision
All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable for that specific section.
Consent for publication
Not applicable for that specific section.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
da Silva Rodrigues, .A., da Cunha, C.C.R.F., do Espirito Santo, D.R. et al. Removal of cephalexin and erythromycin antibiotics, and their resistance genes, by microalgae-bacteria consortium from wastewater treatment plant secondary effluents. Environ Sci Pollut Res 28, 67822–67832 (2021). https://doi.org/10.1007/s11356-021-15351-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-021-15351-x