Abstract
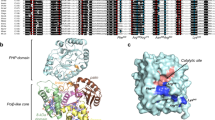
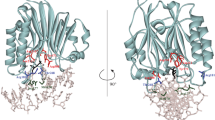
The active sites of the membrane-bound nucleoside triphosphate diphosphohydrolases (NTPDases) regulate and are regulated by coordinated and spatially distant movements of their transmembrane helices, modulating enzyme activity, and substrate specificity. Using site-directed mutagenesis, the roles of the conserved proline residues (N-terminal: P52 and P53; C-terminal: P472, P476, P481, P484, and P485) of human NTPDase3, located in the “linker regions” that connect the N- and C-terminal transmembrane helices with the extracellular active site, were examined. Single cysteine substitutions were strategically placed in the transmembrane domain (N-terminal helix: V42C; C-terminal helix: G489C) to serve as cross-linking “sensors” of helical interactions. These “sensor” background mutant proteins (V42C and G489C NTPDase3) are enzymatically active and are cross-linked by copper phenanthroline less efficiently in the presence of adenosine triphosphate (ATP). Proline to alanine substitutions at P53, P481, P484, and P485 in the V42C background, as well as P53, P481, and P484 in the G489C background, exhibited decreased nucleotidase activities. More importantly, alanine substitutions at P53 and P481 in the V42C background and P481 in the G489C background no longer exhibited the ATP-induced decrease in transmembrane cross-linking efficiency. Interestingly, the P485A mutation abolished oxidative cross-linking at G489C both in the presence and absence of ATP. Taken together, these results suggest a role for proline residues 53 and 481 in the linker regions of human NTPDase3 for coupling nucleotide binding at the enzyme active site to movements and/or rearrangements of the transmembrane helices necessary for optimal nucleotide hydrolysis.






Similar content being viewed by others
Abbreviations
- NTPDase3:
-
Nucleoside triphosphate diphosphohydrolase 3
- TM:
-
Transmembrane
- TMD:
-
Transmembrane domain
- CuPhen:
-
Copper phenanthroline
- NEM:
-
N-ethylmaleimide
- MOPS:
-
3-(N-Morpholino)-propanesulfonic acid
References
Zimmermann H (1999) Two novel families of ectonucleotidases: molecular structures, catalytic properties and a search for function. Trends Pharmacol Sci 20:231–236
Ivanenkov VV, Meller J, Kirley TL (2005) Characterization of disulfide bonds in human nucleoside triphosphate diphosphohydrolase 3 (NTPDase3): implications for NTPDase structural modeling. Biochemistry 44:8998–9012
Murphy-Piedmonte DM, Crawford PA, Kirley TL (2005) Bacterial expression, folding, purification and characterization of soluble NTPDase5 (CD39L4) ecto-nucleotidase. Biochim Biophys Acta 1747:251–259
Ivanenkov VV, Murphy-Piedmonte DM, Kirley TL (2003) Bacterial expression, characterization, and disulfide bond determination of soluble human NTPDase6 (CD39L2) nucleotidase: implications for structure and function. Biochemistry 42:11726–11735
Stout JG, Strobel RS, Kirley TL (1995) Identification and immunolocalization of ecto-ATPDase in chicken stomach. Biochem Mol Biol Int 36:529–535
Kansas GS, Wood GS, Tedder TF (1991) Expression, distribution, and biochemistry of human CD39: role in activation-associated homotypic adhesion of lymphocytes. J Immunol 146:2235–2244
Knowles AF (1988) Differential expression of ectoMg2 + -ATPase and ectoCa2 + -ATPase activities in human hepatoma cells. Arch Biochem Biophys 263:264–271
Karasaki S, Simard A, de Lamirande G (1977) Surface morphology and nucleoside phosphatase activity of rat liver epithelial cells during oncogenic transformation in vitro. Cancer Res 37:3516–3525
Plesner L (1995) Ecto-ATPases: identities and functions. Int Rev Cytol 158:141–214
Kirley TL, Crawford PA, Smith TM (2006) The structure of the nucleoside triphosphate diphosphohydrolases (NTPDases) as revealed by mutagenic and computational modeling analysis. Purinergic Signal 2:379–389
Handa M, Guidotti G (1996) Purification and cloning of a soluble ATP-diphosphohydrolase (apyrase) from potato tubers (Solanum tuberosum). Biochem Biophys Res Commun 218:916–923
Bork P, Sander C, Valencia A (1992) An ATPase domain common to prokaryotic cell cycle proteins, sugar kinases, actin, and hsp70 heat shock proteins. Proc Natl Acad Sci USA 89:7290–7294
Zebisch M, Sträter N (2008) Structural insight into signal conversion and inactivation by NTPDase2 in purinergic signaling. Proc Natl Acad Sci USA 105:6882–6887
Hurley JH, Mason DA, Matthews BW (1992) Flexible-geometry conformational energy maps for the amino acid residue preceding a proline. Biopolymers 32:1443–1446
MacArthur MW, Thornton JM (1991) Influence of proline residues on protein conformation. J Mol Biol 218:397–412
Nicholson H, Tronrud DE, Becktel WJ, Matthews BW (1992) Analysis of the effectiveness of proline substitutions and glycine replacements in increasing the stability of phage T4 lysozyme. Biopolymers 32:1431–1441
Balasubramanian R, Lakshminarayanan AV, Sabesan MN, Tegoni G, Venkatesan K, Ramachandran GN (1971) Studies on the conformation of amino acids. VI. Conformation of the proline ring as observed in crystal structures of amino acids and peptides. Int J Protein Res 3:25–33
Morris AL, MacArthur MW, Hutchinson EG, Thornton JM (1992) Stereochemical quality of protein structure coordinates. Proteins 12:345–364
Wang TF, Ou Y, Guidotti G (1998) The transmembrane domains of ectoapyrase (CD39) affect its enzymatic activity and quaternary structure. J Biol Chem 273:24814–24821
Wang CJ, Vlajkovic SM, Housley GD, Braun N, Zimmermann H, Robson SC, Sevigny J, Soeller C, Thorne PR (2005) C-terminal splicing of NTPDase2 provides distinctive catalytic properties, cellular distribution and enzyme regulation. Biochem J 385:729–736
Chiang WC, Knowles AF (2008) Transmembrane domain interactions affect the stability of the extracellular domain of the human NTPDase 2. Arch Biochem Biophys 472:89–99
Knowles AF, Li C (2006) Molecular cloning and characterization of expressed human ecto-nucleoside triphosphate diphosphohydrolase 8 (E-NTPDase 8) and its soluble extracellular domain. Biochemistry 45:7323–7333
Grinthal A, Guidotti G (2004) Dynamic motions of CD39 transmembrane domains regulate and are regulated by the enzymatic active site. Biochemistry 43:13849–13858
Smith TM, Kirley TL (1999) Site-directed mutagenesis of a human brain ecto-apyrase: evidence that the E-type ATPases are related to the actin/heat shock 70/sugar kinase superfamily. Biochemistry 38:321–328
Murphy DM, Ivanenkov VV, Kirley TL (2002) Identification of cysteine residues responsible for oxidative cross-linking and chemical inhibition of human nucleoside triphosphate diphosphohydrolase 3. J Biol Chem 277:6162–6169
Smith TM, Kirley TL (1998) Cloning, sequencing, and expression of a human brain ecto-apyrase related to both the ecto-ATPases and CD39 ecto-apyrases. Biochim Biophys Acta 1386:65–78
Stoscheck CM (1987) Protein assay sensitive at nanogram levels. Anal Biochem 160:301–305
Fiske CH, Subbarow Y (1925) The colorometric determination of phosphorous. J Biol Chem 66:375–400
Smith TM, Kirley TL (1999) Glycosylation is essential for functional expression of a human brain ecto-apyrase. Biochemistry 38:1509–1516
Careaga CL, Falke JJ (1992) Thermal motions of surface alpha-helices in the D-galactose chemosensory receptor. Detection by disulfide trapping. J Mol Biol 226:1219–1235
Gaddie KJ, Kirley TL (2009) Conserved polar residues stabilize transmembrane domains and promote oligomerization in human nucleoside triphosphate diphosphohydrolase 3. Biochemistry 48:9437–9447
Gerstein M, Lesk AM, Chothia C (1994) Structural mechanisms for domain movements in proteins. Biochemistry 33:6739–6749
Grinthal A, Guidotti G (2002) Transmembrane domains confer different substrate specificities and adenosine diphosphate hydrolysis mechanisms on CD39, CD39L1, and chimeras. Biochemistry 41:1947–1956
Drosopoulos JH, Broekman MJ, Islam N, Maliszewski CR, Gayle RB 3rd, Marcus AJ (2000) Site-directed mutagenesis of human endothelial cell ecto-ADPase/soluble CD39: requirement of glutamate 174 and serine 218 for enzyme activity and inhibition of platelet recruitment. Biochemistry 39:6936–6943
Murphy DM, Kirley TL (2003) Asparagine 81, an invariant glycosylation site near apyrase conserved region 1, is essential for full enzymatic activity of ecto nucleoside triphosphate diphosphohydrolase 3. Arch Biochem Biophys 413:107–115
Mateo J, Kreda S, Henry CE, Harden TK, Boyer JL (2003) Requirement of Cys399 for processing of the human ecto-ATPase (NTPDase2) and its implications for determination of the activities of splice variants of the enzyme. J Biol Chem 278:39960–39968
Basu S, Kirley TL (2005) Identification of a tyrosine residue responsible for N-acetylimidazole-induced increase of activity of ecto-nucleoside triphosphate diphosphohydrolase 3. Purinergic Signalling 1:271–280
Papanikolaou A, Papafotika A, Murphy C, Papamarcaki T, Tsolas O, Drab M, Kurzchalia TV, Kasper M, Christoforidis S (2005) Cholesterol-dependent lipid assemblies regulate the activity of the ecto-nucleotidase CD39. J Biol Chem 280:26406–26414
Mukasa T, Lee Y, Knowles AF (2005) Either the carboxyl- or the amino-terminal region of the human ecto-ATPase (E-NTPDase 2) confers detergent and temperature sensitivity to the chicken ecto-ATP-diphosphohydrolase (E-NTPDase 8). Biochemistry 44:11160–11170
Chen W, Guidotti G (2001) Soluble apyrases release ADP during ATP hydrolysis. Biochem Biophys Res Commun 282:90–95
Zebisch M, Sträter N (2007) Characterization of rat NTPDase1, -2, and -3 ectodomains refolded from bacterial inclusion bodies. Biochemistry 46:11945–11956
Chiang WC, Knowles AF (2008) Inhibition of human NTPDase 2 by modification of an intramembrane cysteine by p-chloromercuriphenylsulfonate and oxidative cross-linking of the transmembrane domains. Biochemistry 47:8775–8785
Stapley BJ, Creamer TP (1999) A survey of left-handed polyproline II helices. Protein Sci 8:587–595
Creamer TP (1998) Left-handed polyproline II helix formation is (very) locally driven. Proteins 33:218–226
Ma K, Kan L, Wang K (2001) Polyproline II helix is a key structural motif of the elastic PEVK segment of titin. Biochemistry 40:3427–3438
Acknowledgments
This work was supported by the National Institutes of Health R01 grant HL72382/HL72382-S1 to T.L.K.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gaddie, K.J., Kirley, T.L. Proline residues link the active site to transmembrane domain movements in human nucleoside triphosphate diphosphohydrolase 3 (NTPDase3). Purinergic Signalling 6, 327–337 (2010). https://doi.org/10.1007/s11302-010-9180-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11302-010-9180-0