Abstract
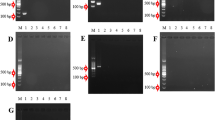
Citrus taxonomy is very complex mainly due to specific aspects of its reproductive biology. A number of studies have been performed using various molecular markers in order to evaluate the level of genetic variability in Citrus. SNP markers have been used for genetic diversity assessment using a variety of different methods. Recently, the availability of EST database and whole genome sequences has made it possible to develop more markers such as SNPs. In the present study, the high-resolution melting curve analysis (HRM) was used to detect SNPs or INDELs in Citrus genus for the first time. We aimed to develop a panel of SNPs to differentiate Citrus genotypes which can also be applied to Citrus biodiversity studies. The results showed that 21 SNP containing markers produced distinct polymorphic melting curves among the Citrus spp. investigated through HRM analysis. It was proved that HRM is an efficient, cost-effective, and accurate method for discriminating citrus SNPs as well as a method to analyze more polymorphisms in a single PCR amplicon, representing a useful tool for genetic, biodiversity, and breeding studies. SNPs developed based on Citrus sinensis EST database showed a good transferability within the Citrus genus. Moreover, HRM analysis allowed the discrimination of citrus genotypes at specific level and the resulting genetic distance analysis clustered these genotypes into three main branches. The results suggested that the panel of SNP markers could be used in a variety of applications in citrus biodiversity assessment and breeding programs using HRM analysis.



Similar content being viewed by others
References
Amar MH, Biswas MK, Zhang ZW, Guo WW (2011) Exploitation of SSR, SRAP and CAPS–SNP markers for genetic diversity of Citrus germplasm collection. Sci Hortic 128:220–227
Barkley NA, Roose ML, Krueger RR, Federici CT (2006) Assessing genetic diversity and population structure in a citrus germplasm collection utilizing simple sequence repeat markers (SSRs). Theor Appl Genet 112:1519–1531
Batley J, Edwards D (2007) SNP applications in plants. In: Oraguzie N, Rikkerink E, Gardiner S, De Silva NH (eds) Association mapping in plants. Springer, New York, pp 95–102
Biswas MK, Chai LJ, Qiang X, Deng XX (2012) Generation, functional analysis and utility of Citrus grandis EST from a flower-derived cDNA library. Mol Biol Rep 39:7221–7235
Cameron JW, Frost HB (1968) Genetics, breeding and nucellar embryony. In: Reuthen W, Webber HJ, Batchelor LD (eds) The citrus industry. University of California Press, Berkeley, pp 325–370
Chagne D, Gasic K, Crowhurst RN, Han Y, Bassett HC, Bowatte DR, Lawrence TJ, Rikkerink EH, Gardiner SE, Korban SS (2008) Development of a set of SNP markers present in expressed genes of the apple. Genomics 92:353–358
Chaisan T, Van K, Kim MY, Kim KD, Choi BS, Lee SH (2012) In silico single nucleotide polymorphism discovery and application to marker-assisted selection in soybean. Mol Breeding 29:221–233
Chen C, Zhou P, Choi YA, Huang S, Gmitter FG (2006) Mining and characterizing microsatellites from citrus ESTs. Theor Appl Genet 112:1248–1257
Corbett Research (2006) High resolution melt assay design and analysis CorProtocolTM. Corbett Research, Sydney
Crifò T, Petrone G, Lo Cicero L, Lo Piero AR (2012) Short cold storage enhances the anthocyanin contents and level of transcripts related to their biosynthesis in blood oranges. J Agr Food Chem 60:476–481
Deng ZN, Gentile A, Nicolosi E, Domina F, Vardi A, Tribulato E (1995) Identification of in vivo and in vitro lemon mutants by RAPD markers. J Hortic Sci 70:117–125
Distefano G, Caruso M, La Malfa S, Gentile A, Wu S-B (2012) High resolution melting analysis is a more sensitive and effective alternative to gel-based platforms in analysis of SSR—an example in Citrus. Plos One 7(8):e44202. doi:10.1371/journal.pone.0044202
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Fang DQ, Roose ML (1997) Identification of closely related Citrus cultivars with inter-simple sequence repeat markers. Theor Appl Genet 95:408–417
Federici CT, Fang DQ, Scora RW, Roose ML (1998) Phylogenetic relationships within the genus Citrus (Rutaceae) and related genera as revealed by RFLP and RAPD analysis. Theor Appl Genet 94:812–822
Flot JF (2007) Champuru 1.0: a computer software for unraveling mixtures of two DNA sequences of unequal lengths. Mol Ecol Notes 7:974–977
Fujii H, Shimada T, Nonaka K, Kita M, Kuniga T, Endo T, Ikoma Y, Omura M (2012) High-throughput genotyping in citrus accessions using an SNP genotyping array. Tree Genet Genomes: 1–9
Ganal MW, Altmann T, Roder MS (2009) SNP identification in crop plants. Curr Opin Plant Biol 12:211–217
Garcia-Lor A, Luro F, Navarro L, Ollitrault P (2012) Comparative use of InDel and SSR markers in deciphering the interspecific structure of cultivated citrus genetic diversity; a perspective for genetic association studies. Mol Genet Genomics 287:77–94
Garritano S, Gemignani F, Voegele C, Nguyen-Dumont T, Le Calvez-Kelm F et al (2009) Determining the effectiveness of high resolution melting analysis for SNP genotyping and mutation scanning at the TP53 locus. BMC Genet 10:5
Garvin MR, Saitoh K, Gharret AJ (2010) Application of single nucleotide polymorphisms to non-model species: a technical review. Mol Ecol Res 10:915–934
Gmitter FG, Chen C, Machado MA, de Souza AA, Ollitrault P, Froehlicher Y, Shimizu T (2012) Citrus genomics. Tree Genet Genomes 8:611–626
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:78–86
Gupta PK, Roy JK, Prasad M (2001) Single nucleotide polymorphisms: a new paradigm for molecular marker technology and DNA polymorphism detection with emphasis on their use in plants. Curr Science 80:524–535
Han Y, Khu DM, Monteros MJ (2012) High-resolution melting analysis for SNP genotyping and mapping in tetraploid alfalfa (Medicago sativa L.). Mol Breeding 29:489–501
Hayashi K, Hashimoto N, Daigen M, Ashikawa I (2004) Development of PCR-based SNP markers for rice blast resistance genes at the Piz locus. Theor Appl Genet 108:1212–1220
Herrero R, Asins MJ, Carbonell EA, Navarro L (1996) Genetic diversity in the orange subfamily Aurantioideae. I. Intraspecies and intragenus genetic variability. Theor Appl Genet 92:599–609
Herrmann MG, Durtschi JD, Wittwer CT, Voelkerding KV (2007) Expanded instrument comparison of amplicon DNA melting analysis for mutation scanning and genotyping. Clin Chem 53:1544–1548
Jeong HJ, Jo YD, Kang BC (2010) Identification of Capsicum species using SNP markers based on high resolution melting analysis. Genome 53:1029–1040
Jiang D, Ye QL, Wang FS, Cao L (2010) The mining of citrus EST–SNP and its application in cultivar discrimination. Agricultural Sciences in China 9:179–190
Lehmensiek A, Sutherland M, McNamara R (2008) The use of high resolution melting (HRM) to map single nucleotide polymorphism markers linked to a covered smut resistance gene in barley. Theor Appl Genet 117:721–728
Liew M, Pryor R, Palais R, Meadows C, Erali M, Lyon E, Wittwer C (2004) Genotyping of single-nucleotide polymorphisms by high-resolution melting of small amplicons. Clin Chem 50:1156–1164
Liu K, Muse SV (2005) Powermarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Luro FL, Laigret F, Bove JM, Ollitrault P (1995) DNA amplified fingerprinting, a useful tool for determination of genetic origin and diversity analysis in Citrus. Hort Sci 30:1063–1067
Luro FL, Costantino G, Terol J, Argout X, Allario T, Wincker P, Talon M, Ollitrault P, Morillon R (2008) Transferability of the EST–SSRs developed on Nules clementine (Citrus clementina Hort ex Tan) to other Citrus species and their effectiveness for genetic mapping. BMC Genom 9:287–299
Mackay JF, Wright CD, Bonfiglioli RG (2008) A new approach to varietal identification in plants by microsatellite high resolution melting analysis: application to the verification of grapevine and olive cultivars. Plant Methods 4:8
Markham NR, Zuker M (2005) DINAMelt web server for nucleic acid melting prediction. Nucleic Acids Res 33:W577–W581
Moore GA (2001) Oranges and lemons: clues to the taxonomy of Citrus from molecular markers. Trends in Genet 17:536–540
Muleo R, Colao MC, Miano D, Cirilli M, Intrieri MC, Baldoni L, Rugini E (2009) Mutation scanning and genotyping by high-resolution DNA melting analysis in olive germplasm. Genome 52:252–260
Nakano M, Shimizu T, Fujii H, Shimada T, Endo T, Nesumi H, Kuniga T, Omura M (2008) Marker enrichment and construction of haplotype-specific BAC contigs for the polyembryony genomic region in Citrus. Breeding Sci 58:375–383
Nicolosi E, Deng ZN, Gentile A, La Malfa S, Continella G, Tribulato E (2000) Citrus phylogeny and genetic origin of important species as investigated by molecular markers. Theor Appl Genet 100:1155–1166
Novelli VM, Takita MA, Machado MA (2004) Identification and analysis of single nucleotide polymorphisms (SNPs) in citrus. Euphytica 138:227–237
Novelli VM, Cristofani M, Souza AA, Machado MA (2006) Development and characterization of polymorphic microsatellite markers for the sweet orange (Citrus sinensis L. Osbeck). Genet Mol Biol 29:90–96
Ohnishi Y, Tanaka T, Ozaki K, Yamada R, Suzuki H, Nakamura Y (2001) A high-throughput SNP typing system for genome-wide association studies. J Hum Genet 46:471–477
Oliver RE, Lazo GR, Lutz JD, Rubenfield MJ, Tinker NA, Anderson JM, Wisniewski Morehead NH, Adhikary D, Jellen EN, Maughan PJ, Brown Guedira GL, Chao S, Beattie AD, Carson ML, Rines HW, Obert DE, Bonman JM, Jackson EW (2011) Model SNP development for complex genomes based on hexaploid oat using high-throughput 454 sequencing technology. BMC Genom 12:77
Ollitrault F, Terol J, Pina JA, Navarro L, Talon M, Ollitrault P (2010) Development of SSR markers from Citrus clementina (Rutaceae) BAC end sequences and interspecific transferability in Citrus. Am J Bot 97:e124–e129
Ollitrault P, Terol J, Garcia-Lor A, Berard A, Chauveau A, Froelicher Y, Belzile C, Morillon R, Navarro L, Brunel D, Talon M (2012) SNP mining in C. clementina BAC end sequences; transferability in the Citrus genus (Rutaceae), phylogenetic inferences and perspectives for genetic mapping. BMC Genom 13:13
Raghuvanshi SS (1962) Cytological studies in the genus Citrus. IV. Evolution in the genus Citrus. Cytologia 27:172–188
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, pp 365–386
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Seipp MT, Durtschi JD, Voelkerding KV, Wittwer CT (2009) Multiplex amplicon genotyping by high-resolution melting. J Biomol Tech 20:160–164
Terol J, Naranjo MA, Ollitrault P, Talon M (2008) Development of genomic resources for Citrus clementina: characterization of three deep-coverage BAC libraries and analysis of 46 000 BAC end sequences. BMC Genom 9:423
van Tienderen PH, de Haan AA, van der Linden CG, Vosman B (2002) Biodiversity assessment using markers for ecologically important traits. Trends Ecol Evol 17:577–582
Wang HT, Kim MK, Kim YJ, Lee HN, Jin HZ, Chen J, Yang DC (2012) Molecular authentication of the Oriental medicines Pericarpium Citri Reticulatae and Citri Unshius Pericarpium using SNP markers. Gene 494:92–95
White HE, Hall VJ, Cross NCP (2007) Methylation-sensitive high resolution melting-curve analysis of the SNRPN gene as a diagnostic screen for Prader–Willi and Angelman syndromes. Clin Chem 53:1960–1962
Wu S-B, Wirthensohn MG, Hunt P, Gibson JP, Sedgley M (2008) High resolution melting analysis of almond SNPs derived from ESTs. Theor Appl Genet 118:1–14
Wu S-B, Tavassolian I, Rabiei G, Hunt P, Wirthensohn M, Gibson J, Ford CM, Sedgley M (2009) Mapping SNP anchored genes using high resolution melting analysis in almond. Mol Genet Genomics 282:273–281
Wu S-B, Franks T, Hunt P, Wirthensohn M, Gibson J, Sedgley M (2010) Discrimination of SNP genotypes associated with complex haplotypes by high resolution melting analysis in almond: implications for improved marker efficiencies. Mol Breed 25:351–357
Wu B, Yang R-T, Zhu S-P, Zhong Y, Jiang B, Zeng J-W, Zhong G (2012) Genotyping single nucleotide polymorphisms in mandarin cultivars using high resolution melting analysis. Acta Hort Sinica 39:777–782
Acknowledgments
This research was supported by Regione Siciliana, Project “Innovazioni bioagronomiche e fitopatologiche per il contenimento di CTV e per il rinnovamento dell’agrumicoltura siciliana”.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by W.-W. Guo
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online Resource 1
Identification and reference number of the ESTs (HarvEST: Citrus 1.32 Assembly: #C54CS) forming the contigs used for SNPs identification (PDF 192 kb)
Online Resource 2
SNP haplotypes in 18 citrus genotypes as revealed by HRM analysis performed with 21 selected EST-SNPs markers (PDF 20 kb)
Rights and permissions
About this article
Cite this article
Distefano, G., La Malfa, S., Gentile, A. et al. EST-SNP genotyping of citrus species using high-resolution melting curve analysis. Tree Genetics & Genomes 9, 1271–1281 (2013). https://doi.org/10.1007/s11295-013-0636-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-013-0636-6