Abstract
With advantages of low substrates cost, high optical purity of end products and environmentally friendly fermentation process, microbial production of valuable chemicals grow rapidly. Compared with static microbial strain engineering strategies, such as gene deletion, overexpression and mutation, dynamic pathway regulation is a new approach that balances cellular growth and chemical production. Quorum sensing is a natural microbial communication system responsible for cell-density-related cell behaviors. Accordingly, quorum sensing systems can be employed to achieve dynamic regulation in microorganisms without the need for manual intervention or the use of chemical inducers. In this review, natural quorum sensing systems are firstly summarized. Then, recent progress in using quorum sensing circuits in the field of metabolic engineering is highlighted. The current application challenges of quorum sensing systems and future perspectives in microbial synthesis of chemicals are also discussed.
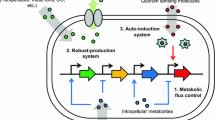
Graphical abstract


Similar content being viewed by others
References
Ansaldi M, Marolt D, Stebe T, Mandic-Mulec I, Dubnau D (2002) Specific activation of the Bacillus quorum-sensing systems by isoprenylated pheromone variants. Mol Microbiol 44(6):1561–1573. https://doi.org/10.1046/j.1365-2958.2002.02977.x
Balagadde FK, Song H, Ozaki J, Collins CH, Barnet M, Arnold FH, Quake SR, You L (2008) A synthetic Escherichia coli predator-prey ecosystem. Mol Syst Biol 4:187. https://doi.org/10.1038/msb.2008.24
Chen GQ, Jiang XR (2018) Engineering microorganisms for improving polyhydroxyalkanoate biosynthesis. Curr Opin Biotechnol 53:20–25. https://doi.org/10.1016/j.copbio.2017.10.008
Core L, Perego M (2003) TPR-mediated interaction of RapC with ComA inhibits response regulator-DNA binding for competence development in Bacillus subtilis. Mol Microbiol 49(6):1509–1522. https://doi.org/10.1046/j.1365-2958.2003.03659.x
Correa GG, Lins M, Silva BF, de Paiva GB, Zocca VFB, Ribeiro NV, Picheli FP, Mack M, Pedrolli DB (2020) A modular autoinduction device for control of gene expression in Bacillus subtilis. Metab Eng 61:326–334. https://doi.org/10.1016/j.ymben.2020.03.012
Dahl RH, Zhang F, Alonso-Gutierrez J, Baidoo E, Batth TS, Redding-Johanson AM, Petzold CJ, Mukhopadhyay A, Lee TS, Adams PD, Keasling JD (2013) Engineering dynamic pathway regulation using stress-response promoters. Nat Biotechnol 31(11):1039–1046. https://doi.org/10.1038/nbt.2689
Danino T, Mondragon-Palomino O, Tsimring L, Hasty J (2010) A synchronized quorum of genetic clocks. Nature 463(7279):326–330. https://doi.org/10.1038/nature08753
Das S, Das S, Ghangrekar MM (2019) Quorum-sensing mediated signals: A promising multi-functional modulators for separately enhancing algal yield and power generation in microbial fuel cell. Bioresour Technol 294:122138. https://doi.org/10.1016/j.biortech.2019.122138
Das S, Das S, Ghangrekar MM (2022) Bacterial signalling mechanism: An innovative microbial intervention with multifaceted applications in microbial electrochemical technologies: a review. Bioresour Technol 344(Pt B):126218. https://doi.org/10.1016/j.biortech.2021.126218
DeLisa MP, Bentley WE (2002) Bacterial autoinduction: looking outside the cell for new metabolic engineering targets. Microb Cell Fact 1(1):5. https://doi.org/10.1186/1475-2859-1-5
Deng J, Chen C, Gu Y, Lv X, Liu Y, Li J, Ledesma-Amaro R, Du G, Liu L (2019) Creating an in vivo bifunctional gene expression circuit through an aptamer-based regulatory mechanism for dynamic metabolic engineering in Bacillus subtilis. Metab Eng 55:179–190. https://doi.org/10.1016/j.ymben.2019.07.008
Din MO, Danino T, Prindle A, Skalak M, Selimkhanov J, Allen K, Julio E, Atolia E, Tsimring LS, Bhatia SN, Hasty J (2016) Synchronized cycles of bacterial lysis for in vivo delivery. Nature 536(7614):81–85. https://doi.org/10.1038/nature18930
Dinh CV, Prather KLJ (2019) Development of an autonomous and bifunctional quorum-sensing circuit for metabolic flux control in engineered Escherichia coli. Proc Natl Acad Sci USA 116(51):25562–25568. https://doi.org/10.1073/pnas.1911144116
Dusane DH, Zinjarde SS, Venugopalan VP, McLean RJ, Weber MM, Rahman PK (2010) Quorum sensing: implications on rhamnolipid biosurfactant production. Biotechnol Genet Eng Rev 27:159–184. https://doi.org/10.1080/02648725.2010.10648149
Farmer WR, Liao JC (2000) Improving lycopene production in Escherichia coli by engineering metabolic control. Nat Biotechnol 18(5):533–537. https://doi.org/10.1038/75398
Fuqua C, Greenberg EP (1998) Self perception in bacteria: quorum sensing with acylated homoserine lactones. Curr Opin Microbiol 1(2):183–189. https://doi.org/10.1016/s1369-5274(98)80009-x
Ge C, Yu Z, Sheng H, Shen X, Sun X, Zhang Y, Yan Y, Wang J, Yuan Q (2022) Redesigning regulatory components of quorum-sensing system for diverse metabolic control. Nat Commun 13(1):2182. https://doi.org/10.1038/s41467-022-29933-x
Goo E, An JH, Kang Y, Hwang I (2015) Control of bacterial metabolism by quorum sensing. Trends Microbiol 23(9):567–576. https://doi.org/10.1016/j.tim.2015.05.007
Gu F, Jiang W, Mu Y, Huang H, Su T, Luo Y, Liang Q, Qi Q (2020) Quorum sensing-based dual-function switch and its application in solving two key metabolic engineering problems. ACS Synth Biol 9(2):209–217. https://doi.org/10.1021/acssynbio.9b00290
Gupta A, Reizman IM, Reisch CR, Prather KL (2017) Dynamic regulation of metabolic flux in engineered bacteria using a pathway-independent quorum-sensing circuit. Nat Biotechnol 35(3):273–279. https://doi.org/10.1038/nbt.3796
He X, Chen Y, Liang Q, Qi Q (2017) Autoinduced AND gate controls metabolic pathway dynamically in response to microbial communities and cell physiological state. ACS Synth Biol 6(3):463–470. https://doi.org/10.1021/acssynbio.6b00177
Janssens Y, Wynendaele E, Verbeke F, Debunne N, Gevaert B, Audenaert K, Van DeWiele C, De Spiegeleer B (2018) Screening of quorum sensing peptides for biological effects in neuronal cells. Peptides 101:150–156. https://doi.org/10.1016/j.peptides.2018.01.013
Jiang W, He X, Luo Y, Mu Y, Gu F, Liang Q, Qi Q (2020) Two completely orthogonal quorum sensing systems with self-produced autoinducers enable automatic delayed cascade control. ACS Synth Biol 9(9):2588–2599. https://doi.org/10.1021/acssynbio.0c00370
Kim EM, Woo HM, Tian T, Yilmaz S, Javidpour P, Keasling JD, Lee TS (2017) Autonomous control of metabolic state by a quorum sensing (QS)-mediated regulator for bisabolene production in engineered E. coli. Metab Eng 44:325–336. https://doi.org/10.1016/j.ymben.2017.11.004
Li Y, Li M, Zhang X, Yang P, Liang Q, Qi Q (2013) A novel whole-phase succinate fermentation strategy with high volumetric productivity in engineered Escherichia coli. Bioresour Technol 149:333–340. https://doi.org/10.1016/j.biortech.2013.09.077
Liu Y, Liu L, Li J, Du G, Chen J (2019) Synthetic biology toolbox and chassis development in Bacillus subtilis. Trends Biotechnol 37(5):548–562. https://doi.org/10.1016/j.tibtech.2018.10.005
Lv Y, Qian S, Du G, Chen J, Zhou J, Xu P (2019) Coupling feedback genetic circuits with growth phenotype for dynamic population control and intelligent bioproduction. Metab Eng 54:109–116. https://doi.org/10.1016/j.ymben.2019.03.009
Ma C, Zhang K, Zhang X, Liu G, Zhu T, Che Q, Li D, Zhang G (2021) Heterologous expression and metabolic engineering tools for improving terpenoids production. Curr Opin Biotechnol 69:281–289. https://doi.org/10.1016/j.copbio.2021.02.008
Mahto KU, Kumari S, Das S (2022) Unraveling the complex regulatory networks in biofilm formation in bacteria and relevance of biofilms in environmental remediation. Crit Rev Biochem Mol Biol 57(3):305–332. https://doi.org/10.1080/10409238.2021.2015747
Mao Y, Li G, Chang Z, Tao R, Cui Z, Wang Z, Tang YJ, Chen T, Zhao X (2018) Metabolic engineering of Corynebacterium glutamicum for efficient production of succinate from lignocellulosic hydrolysate. Biotechnol Biofuels 11:95. https://doi.org/10.1186/s13068-018-1094-z
March JC, Bentley WE (2004) Quorum sensing and bacterial cross-talk in biotechnology. Curr Opin Biotechnol 15(5):495–502. https://doi.org/10.1016/j.copbio.2004.08.013
Martin VJ, Pitera DJ, Withers ST, Newman JD, Keasling JD (2003) Engineering a mevalonate pathway in Escherichia coli for production of terpenoids. Nat Biotechnol 21(7):796–802. https://doi.org/10.1038/nbt833
Mayville P, Ji GY, Beavis R, Yang HM, Goger M, Novick Muir TW (1999) Structure-activity analysis of synthetic autoinducing thiolactone peptides from Staphylococcus aureus responsible for virulence. Proc Natl Acad Sci U S A 96:1218–1223. https://doi.org/10.1073/pnas.96.4.1218
McGivney E, Jones KE, Weber B, Valentine AM, VanBriesen JM, Gregory KB (2018) Quorum sensing signals form complexes with Ag+ and Cu2+ cations. ACS Chem Biol 13(4):894–899. https://doi.org/10.1021/acschembio.7b01000
Medina G, Juarez K, Valderrama B, Soberon-Chavez G (2003) Mechanism of Pseudomonas aeruginosa RhlR transcriptional regulation of the rhlAB promoter. J Bacteriol 185(20):5976–5983. https://doi.org/10.1128/JB.185.20.5976-5983.2003
Miyashiro T, Ruby EG (2012) Shedding light on bioluminescence regulation in Vibrio fischeri. Mol Microbiol 84(5):795–806. https://doi.org/10.1111/j.1365-2958.2012.08065.x
Nakagami G, Minematsu T, Asada M, Nagase T, Akase T, Huang L, Morohoshi T, Ikeda T, Ohta Y, Sanada H (2011) The Pseudomonas aeruginosa quorum-sensing signal N-(3-oxododecanoyl) homoserine lactone can accelerate cutaneous wound healing through myofibroblast differentiation in rats. FEMS Immunol Med Microbiol 62(2):157–163. https://doi.org/10.1111/j.1574-695X.2011.00796.x
Ng WL, Bassler BL (2009) Bacterial quorum-sensing network architectures. Annu Rev Genet 43:197–222. https://doi.org/10.1146/annurev-genet-102108-134304
Pai A, Tanouchi Y, You L (2012) Optimality and robustness in quorum sensing (QS)-mediated regulation of a costly public good enzyme. Proc Natl Acad Sci USA 109(48):19810–19815. https://doi.org/10.1073/pnas.1211072109
Patel K, Rodriguez C, Stabb EV, Hagen SJ (2020) Spatially propagating activation of quorum sensing in Vibrio fischeri and the transition to low population density. Phys Rev E 101(6–1):062421. https://doi.org/10.1103/PhysRevE.101.062421
Potumarthi R, Subhakar C, Vanajakshi J, Jetty A (2008) Effect of aeration and agitation regimes on lipase production by newly isolated Rhodotorula mucilaginosa-MTCC 8737 in stirred tank reactor using molasses as sole production medium. Appl Biochem Biotechnol 151(2–3):700–710. https://doi.org/10.1007/s12010-008-8293-1
Rodrigues AL, Trachtmann N, Becker J, Lohanatha AF, Blotenberg J, Bolten CJ, Korneli C, de Souza Lima AO, Porto LM, Sprenger GA, Wittmann C (2013) Systems metabolic engineering of Escherichia coli for production of the antitumor drugs violacein and deoxyviolacein. Metab Eng 20:29–41. https://doi.org/10.1016/j.ymben.2013.08.004
Ryan RP, Dow JM (2008) Diffusible signals and interspecies communication in bacteria. Microbiology 154(Pt 7):1845–1858. https://doi.org/10.1099/mic.0.2008/017871-0
Scott SR, Din MO, Bittihn P, Xiong L, Tsimring LS, Hasty J (2017) A stabilized microbial ecosystem of self-limiting bacteria using synthetic quorum-regulated lysis. Nat Microbiol 2:17083. https://doi.org/10.1038/nmicrobiol.2017.83
Shen YP, Fong LS, Yan ZB, Liu JZ (2019) Combining directed evolution of pathway enzymes and dynamic pathway regulation using a quorum-sensing circuit to improve the production of 4-hydroxyphenylacetic acid in Escherichia coli. Biotechnol Biofuels 12:94. https://doi.org/10.1186/s13068-019-1438-3
Singh BN, Singh HB, Singh A, Singh BR, Mishra A, Nautiyal CS (2012) Lagerstroemia speciosa fruit extract modulates quorum sensing-controlled virulence factor production and biofilm formation in Pseudomonas aeruginosa. Microbiology 158(Pt 2):529–538. https://doi.org/10.1099/mic.0.052985-0
Soma Y, Hanai T (2015) Self-induced metabolic state switching by a tunable cell density sensor for microbial isopropanol production. Metab Eng 30:7–15. https://doi.org/10.1016/j.ymben.2015.04.005
Soma Y, Takahashi M, Fujiwara Y, Shinohara T, Izumi Y, Hanai T, Bamba T (2021) Design of synthetic quorum sensing achieving induction timing-independent signal stabilization for dynamic metabolic engineering of E. coli. ACS Synth Biol 10(6):1384–1393. https://doi.org/10.1021/acssynbio.1c00008
Song H, Ding MZ, Jia XQ, Ma Q, Yuan YJ (2014) Synthetic microbial consortia: from systematic analysis to construction and applications. Chem Soc Rev 43(20):6954–6981. https://doi.org/10.1039/c4cs00114a
Stevens AM, Dolan KM, Greenberg EP (1994) Synergistic binding of the Vibrio fischeri LuxR transcriptional activator domain and RNA polymerase to the lux promoter region. Proc Natl Acad Sci USA 91(26):12619–19623. https://doi.org/10.1073/pnas.91.26.12619
Tateda K, Ishii Y, Horikawa M, Matsumoto T, Miyairi S, Pechere JC, Standiford TJ, Ishiguro M, Yamaguchi K (2003) The Pseudomonas aeruginosa autoinducer N-3-oxododecanoyl homoserine lactone accelerates apoptosis in macrophages and neutrophils. Infect Immun 71(10):5785–5793. https://doi.org/10.1128/IAI.71.10.5785-5793.2003
Wang B, Kitney RI, Joly N, Buck M (2011) Engineering modular and orthogonal genetic logic gates for robust digital-like synthetic biology. Nat Commun 2:508. https://doi.org/10.1038/ncomms1516
Williams TC, Averesch NJH, Winter G, Plan MR, Vickers CE, Nielsen LK, Kromer JO (2015) Quorum-sensing linked RNA interference for dynamic metabolic pathway control in Saccharomyces cerevisiae. Metab Eng 29:124–134. https://doi.org/10.1016/j.ymben.2015.03.008
Wu J, Bao M, Duan X, Zhou P, Chen C, Gao J, Cheng S, Zhuang Q, Zhao Z (2020) Developing a pathway-independent and full-autonomous global resource allocation strategy to dynamically switching phenotypic states. Nat Commun 11(1):5521. https://doi.org/10.1038/s41467-020-19432-2
Wu J, Zhou L, Duan X, Peng H, Liu S, Zhuang Q, Pablo CM, Fan X, Ding S, Dong M, Zhou J (2021a) Applied evolution: dual dynamic regulations-based approaches in engineering intracellular malonyl-CoA availability. Metab Eng 67:403–416. https://doi.org/10.1016/j.ymben.2021.08.004
Wu S, Xue Y, Yang S, Xu C, Liu C, Liu X, Liu J, Zhu H, Zhao GR, Yang A, Qiao J (2021b) Combinational quorum sensing devices for dynamic control in cross-feeding cocultivation. Metab Eng 67:186–197. https://doi.org/10.1016/j.ymben.2021.07.002
Yang Y, Lin Y, Wang J, Wu Y, Zhang R, Cheng M, Shen X, Wang J, Chen Z, Li C, Yuan Q, Yan Y (2018) Sensor-regulator and RNAi based bifunctional dynamic control network for engineered microbial synthesis. Nat Commun 9(1):3043. https://doi.org/10.1038/s41467-018-05466-0
Zhang H, Wang X (2016) Modular co-culture engineering, a new approach for metabolic engineering. Metab Eng 37:114–121. https://doi.org/10.1016/j.ymben.2016.05.007
Zhang F, Carothers JM, Keasling JD (2012) Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat Biotechnol 30(4):354–359. https://doi.org/10.1038/nbt.2149
Funding
This work was financially supported by the National Natural Science Foundation of China (31600066, 32070111, 31870105), Rizhao Science and Technology Innovation Project (2020CXZX1206), the Science and Technology Program of the University of Jinan (XKY2028), and the Higher Educational Science and Technology Program of Jinan City (2021GXRC088).
Author information
Authors and Affiliations
Contributions
All the authors were involved in the conception and design of this review. PG, QM, SZ, CL, HZ, and SJ wrote the article; PG, JG, and QL read and corrected the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gu, P., Ma, Q., Zhao, S. et al. Application of quorum sensing system in microbial synthesis of valuable chemicals: a mini-review. World J Microbiol Biotechnol 38, 192 (2022). https://doi.org/10.1007/s11274-022-03382-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-022-03382-6