Abstract
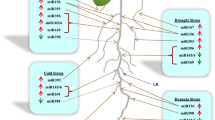
Sweet potato (Ipomoea batatas) is one of the major food and vegetable crops around the world. Sweet potato produces two types of roots: storage roots and fibrous roots. The tuberous root is a modified lateral root that functions as a storage organ, and is the portion of the plant consumed by humans. However, the molecular mechanisms controlling transformation of lateral roots to storage roots are still unclear. In this study, we systematically investigated the potential role of 16 conserved microRNAs (miRNAs) in floral root development. miRNAs are an extensive class of small regulatory RNAs controlling almost all biological and metabolic process in plants. Our results show that some miRNAs are expressed in an organ-dependent manner. For example, the expression level of miR156 and miR162 was significantly lower in storage roots than in leaves and fibrous roots. This suggests that miRNAs may play a role during storage root initiation and development. The expression of miR167 is higher in stamens than that in other tissues tested, indicating that miR167 is crucial to stamen development. miR398—a stress-responsive miRNA—may be involved in fibrous root and storage root development. Additionally, three miRNAs, miR160, miR164 and miR166, also appear to be important in fibrous root and storage root development.





Similar content being viewed by others
Abbreviations
- miRNA:
-
MicroRNA
- qRT-PCR:
-
Quantitative real time PCR
- RISC:
-
RNA-induced silencing complex
- SPB:
-
SQUAMOSA promoter binding transcription factor
- AP2:
-
APETALA2
References
Achard P, Herr A, Baulcombe DC, Harberd NP (2004) Modulation of floral development by a gibberellin-regulated microRNA. Development 131(14):3357–3365. doi:10.1242/dev.01206
Allen RS, Li J, Stahle MI, Dubroue A, Gubler F, Millar AA (2007) Genetic analysis reveals functional redundancy and the major target genes of the Arabidopsis miR159 family. Proc Natl Acad Sci USA 104(41):16371–16376. doi:10.1073/pnas.0707653104
Ambros V, Bartel B, Bartel DP, Burge CB, Carrington JC, Chen X, Dreyfuss G, Eddy SR, Griffiths-Jones S, Marshall M, Matzke M, Ruvkun G, Tuschl T (2003) A uniform system for microRNA annotation. RNA 9(3):277–279
Aukerman MJ, Sakai H (2003) Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. Plant Cell 15(11):2730–2741. doi:10.1105/tpc.016238
Axtell MJ, Westholm JO, Lai EC (2011) Vive la difference: biogenesis and evolution of microRNAs in plants and animals. Genome Biol 12(4):221. doi:10.1186/gb-2011-12-4-221
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116(2):281–297
Blazquez MA, Green R, Nilsson O, Sussman MR, Weigel D (1998) Gibberellins promote flowering of arabidopsis by activating the LEAFY promoter. Plant cell 10(5):791–800
Carlsbecker A, Lee JY, Roberts CJ, Dettmer J, Lehesranta S, Zhou J, Lindgren O, Moreno-Risueno MA, Vaten A, Thitamadee S, Campilho A, Sebastian J, Bowman JL, Helariutta Y, Benfey PN (2010) Cell signalling by microRNA165/6 directs gene dose-dependent root cell fate. Nature 465(7296):316–321. doi:10.1038/nature08977
Carrington JC, Ambros V (2003) Role of microRNAs in plant and animal development. Science 301(5631):336–338. doi:10.1126/science.1085242
Chapman EJ, Estelle M (2009) Mechanism of auxin-regulated gene expression in plants. Annu Rev Genet 43:265–285. doi:10.1146/annurev-genet-102108-134148
Chen X (2004) A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303(5666):2022–2025. doi:10.1126/science.1088060
Chen X (2009) Small RNAs and their roles in plant development. Annu Rev Cell Dev Biol 25:21–44. doi:10.1146/annurev.cellbio.042308.113417
Chen R, Hu Z, Zhang H (2009) Identification of microRNAs in wild soybean (Glycine soja). J Integr Plant Biol 51(12):1071–1079. doi:10.1111/j.1744-7909.2009.00887.x
Dehury B, Panda D, Sahu J, Sahu M, Sarma K, Barooah M, Sen P, Modi M (2013) In silico identification and characterization of conserved miRNAs and their target genes in sweet potato (Ipomoea batatas L.) expressed sequence tags (ESTs). Plant Signal Behav 8:e26543. doi:10.4161/psb.26543
Ding Y, Tao Y, Zhu C (2013) Emerging roles of microRNAs in the mediation of drought stress response in plants. J Exp Bot 64(11):3077–3086. doi:10.1093/jxb/ert164
Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ (2006) miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 34(Database issue):D140–144. doi:10.1093/nar/gkj112
Guo HS, Xie Q, Fei JF, Chua NH (2005) MicroRNA directs mRNA cleavage of the transcription factor NAC1 to downregulate auxin signals for arabidopsis lateral root development. Plant Cell 17(5):1376–1386. doi:10.1105/tpc.105.030841
Han Y, Luan F, Zhu H, Shao Y, Chen A, Lu C, Luo Y, Zhu B (2009) Computational identification of microRNAs and their targets in wheat (Triticum aestivum L.). Science in China Series C, Life sciences / Chinese Academy of Sciences 52(11):1091–1100. doi:10.1007/s11427-009-0144-y
Khan Barozai MY, Irfan M, Yousaf R, Ali I, Qaisar U, Maqbool A, Zahoor M, Rashid B, Hussnain T, Riazuddin S (2008) Identification of micro-RNAs in cotton. Plant physiol biochem PPB / Societe francaise de physiologie vegetale 46(8–9):739–751. doi:10.1016/j.plaphy.2008.05.009
Kurihara Y, Watanabe Y (2004) Arabidopsis micro-RNA biogenesis through Dicer-like 1 protein functions. Proc Natl Acad Sci USA 101(34):12753–12758. doi:10.1073/pnas.0403115101
Liang M, Haroldsen V, Cai X, Wu Y (2006) Expression of a putative laccase gene, ZmLAC1, in maize primary roots under stress. Plant Cell Environ 29(5):746–753. doi:10.1111/j.1365-3040.2005.01435.x
Liu Q, Chen YQ (2009) Insights into the mechanism of plant development: interactions of miRNAs pathway with phytohormone response. Biochem Biophys Res Commun 384(1):1–5. doi:10.1016/j.bbrc.2009.04.028
Low JW, Arimond M, Osman N, Cunguara B, Zano F, Tschirley D (2007) A food-based approach introducing orange-fleshed sweet potatoes increased vitamin A intake and serum retinol concentrations in young children in rural Mozambique. J Nutr 137(5):1320–1327
Mallory AC, Bartel DP, Bartel B (2005) MicroRNA-directed regulation of Arabidopsis AUXIN RESPONSE FACTOR17 is essential for proper development and modulates expression of early auxin response genes. Plant Cell 17(5):1360–1375. doi:10.1105/tpc.105.031716
Miyashima S, Koi S, Hashimoto T, Nakajima K (2011) Non-cell-autonomous microRNA165 acts in a dose-dependent manner to regulate multiple differentiation status in the Arabidopsis root. Development 138(11):2303–2313. doi:10.1242/dev.060491
Nagpal P, Ellis CM, Weber H, Ploense SE, Barkawi LS, Guilfoyle TJ, Hagen G, Alonso JM, Cohen JD, Farmer EE, Ecker JR, Reed JW (2005) Auxin response factors ARF6 and ARF8 promote jasmonic acid production and flower maturation. Development 132(18):4107–4118. doi:10.1242/dev.01955
Qiu CX, Xie FL, Zhu YY, Guo K, Huang SQ, Nie L, Yang ZM (2007) Computational identification of microRNAs and their targets in Gossypium hirsutum expressed sequence tags. Gene 395(1–2):49–61. doi:10.1016/j.gene.2007.01.034
Saeed AI, Bhagabati NK, Braisted JC, Liang W, Sharov V, Howe EA, Li J, Thiagarajan M, White JA, Quackenbush J (2006) TM4 microarray software suite. Methods Enzymol 411:134–193. doi:10.1016/S0076-6879(06)11009-5
Schmittgen TD, Lee EJ, Jiang J (2008) High-throughput real-time PCR. Methods Mol Biol 429:89–98. doi:10.1007/978-1-60327-040-3_7
Schwab R, Palatnik JF, Riester M, Schommer C, Schmid M, Weigel D (2005) Specific effects of microRNAs on the plant transcriptome. Dev Cell 8(4):517–527. doi:10.1016/j.devcel.2005.01.018
Taki FA, Pan X, Zhang B (2014) Chronic nicotine exposure systemically alters microRNA expression profiles during post-embryonic stages in Caenorhabditis elegans. J Cell Physiol 229(1):79–89. doi:10.1002/jcp.24419
Tang G, Reinhart BJ, Bartel DP, Zamore PD (2003) A biochemical framework for RNA silencing in plants. Genes Dev 17(1):49–63. doi:10.1101/gad.1048103
Uberti-Manassero NG, Lucero LE, Viola IL, Vegetti AC, Gonzalez DH (2012) The class I protein AtTCP15 modulates plant development through a pathway that overlaps with the one affected by CIN-like TCP proteins. J Exp Bot 63(2):809–823. doi:10.1093/jxb/err305
Voinnet O (2009) Origin, biogenesis, and activity of plant microRNAs. Cell 136(4):669–687. doi:10.1016/j.cell.2009.01.046
Wang JW, Wang LJ, Mao YB, Cai WJ, Xue HW, Chen XY (2005) Control of root cap formation by MicroRNA-targeted auxin response factors in Arabidopsis. Plant cell 17(8):2204–2216. doi:10.1105/tpc.105.033076
Wang JW, Czech B, Weigel D (2009) miR156-regulated SPL transcription factors define an endogenous flowering pathway in Arabidopsis thaliana. Cell 138(4):738–749. doi:10.1016/j.cell.2009.06.014
Wang J, Hou X, Yang X (2011) Identification of conserved microRNAs and their targets in Chinese cabbage (Brassica rapa subsp. pekinensis). Genome / National Research Council Canada = Genome / Conseil national de recherches Canada 54(12):1029–1040. doi:10.1139/g11-069
Wu MF, Tian Q, Reed JW (2006) Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development 133(21):4211–4218. doi:10.1242/dev.02602
Xie F, Frazier TP, Zhang B (2010) Identification and characterization of microRNAs and their targets in the bioenergy plant switchgrass (Panicum virgatum). Planta 232(2):417–434. doi:10.1007/s00425-010-1182-1
Xie K, Shen J, Hou X, Yao J, Li X, Xiao J, Xiong L (2012) Gradual increase of miR156 regulates temporal expression changes of numerous genes during leaf development in rice. Plant Physiol 158(3):1382–1394. doi:10.1104/pp. 111.190488
Xie F, Jones DC, Wang Q, Sun R, Zhang B (2015) Small RNA sequencing identifier miRNA roles iun ovule and fibre development. Plant Biotechnol J 13. doi:10.1111/pbi.12296
Yang JH, Han SJ, Yoon EK, Lee WS (2006) Evidence of an auxin signal pathway, microRNA167-ARF8-GH3, and its response to exogenous auxin in cultured rice cells. Nucleic Acids Res 34(6):1892–1899. doi:10.1093/nar/gkl118
Yin ZJ, Shen FF (2010) Identification and characterization of conserved microRNAs and their target genes in wheat (Triticum aestivum). Genet Mole Res GMR 9(2):1186–1196. doi:10.4238/vol9-2gmr805
Yoon EK, Yang JH, Lim J, Kim SH, Kim SK, Lee WS (2010) Auxin regulation of the microRNA390-dependent transacting small interfering RNA pathway in Arabidopsis lateral root development. Nucleic Acids Res 38(4):1382–1391. doi:10.1093/nar/gkp1128
Zhang BH (2015) MicroRNA: a new target for improving plant tolerance to abiotic stress. J Exp Bot. doi:10.1093/jxb/erv013
Zhang B, Pan X (2009) Expression of microRNAs in cotton. Mol Biotechnol 42(3):269–274. doi:10.1007/s12033-009-9163-y
Zhang B, Pan X, Anderson TA (2006a) Identification of 188 conserved maize microRNAs and their targets. FEBS Lett 580(15):3753–3762. doi:10.1016/j.febslet.2006.05.063
Zhang B, Pan X, Cannon CH, Cobb GP, Anderson TA (2006b) Conservation and divergence of plant microRNA genes. Plant J Cell Mol Biol 46(2):243–259. doi:10.1111/j.1365-313X.2006.02697.x
Zhang B, Pan X, Cobb GP, Anderson TA (2006c) Plant microRNA: a small regulatory molecule with big impact. Dev Biol 289(1):3–16. doi:10.1016/j.ydbio.2005.10.036
Zhang B, Wang Q (2015) MicroRNA-based biotechnology for plant improvement. J Cell Physiol 230(1):1–15. doi:10.1002/jcp.24685
Zhang B, Wang Q, Wang K, Pan X, Liu F, Guo T, Cobb GP, Anderson TA (2007) Identification of cotton microRNAs and their targets. Gene 397(1–2):26–37. doi:10.1016/j.gene.2007.03.020
Zhang B, Pan X, Stellwag EJ (2008) Identification of soybean microRNAs and their targets. Planta 229(1):161–182. doi:10.1007/s00425-008-0818-x
Zhang W, Luo Y, Gong X, Zeng W, Li S (2009) Computational identification of 48 potato microRNAs and their targets. Comput Biol Chem 33(1):84–93. doi:10.1016/j.compbiolchem.2008.07.006
Compliance with Ethical Standards
No potential conflict of interest is declared. This manuscript does not concern human and animal matters.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Sun, R., Guo, T., Cobb, J. et al. Role of microRNAs During Flower and Storage Root Development in Sweet Potato. Plant Mol Biol Rep 33, 1731–1739 (2015). https://doi.org/10.1007/s11105-015-0869-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-015-0869-7