Abstract
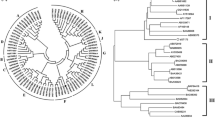
Flavonoids are a large class of phenylpropanoid-derived secondary metabolites, which are usually glycosylated by UDP-glycosyltransferases with one or more sugar groups. Here, we report the cloning and biochemical characterization of a flavonoid glycosyltransferase gene from Withania somnifera (WsGT), which is an important medicinal plant used in Ayurvedic formulations. Using PCR primers, designed for a highly conserved region of previously reported glycosyltransferases, we were able to isolate the corresponding fragment of the WsGT gene. Rapid amplification of cDNA ends (RACE) was then employed to isolate full-length cDNA, which had an open reading frame of 1,371 bp that encode for 456 amino acids. Phylogenetic analysis indicated that WsGT was similar to that of family 1 GT-B glycosyltransferase. Biochemical analysis revealed that WsGT interacts with UDP-glucose and was capable of regiospecifically glycosylating flavonoid-7-ols, such as apigenin, naringenin, luteolin, diadzein and genistein. Expression profiling studies showed that WsGT was highly expressed in young and mature leaves of W. somnifera. Furthermore, exposure to salicylic acid enhanced the expression of WsGT in the leaves and heat shock treatment resulted in decreased expression of WsGT after an initial increase. This may suggest the role of WsGT in response to abiotic/biotic stresses.



Similar content being viewed by others
References
Arend J, Warzecha H, Hefner T, Stockigt J (2001) Utilizing genetically engineered bacteria to produce plant-specific glucosides. Biotechnol Bioeng 76:126–131. doi:10.1002/bit.1152
Arun Kumar MKK, Bhan MK, Khanna PK, Suri KA (2007) Morphological and chemical variation in 25 collections of the Indian medicinal plant, Withania somnifera (L.) Dunal (Solanaceae). Genet Resour Crop Ev 54:655–660
Bowles D, Isayenkova J, Lim EK, Poppenberger B (2005) Glycosyltransferases: Managers of small molecules. Curr Opin Plant Biol 8:254–263
Bowles D, Lim EK, Poppenberger B, Vaistij FE (2006) Glycosyltransferases of lipophilic small molecules. Annu Rev Plant Biol 57:567–597. doi:10.1146/annurev.arplant.57.032905.105429
Caputi LMM, Goremykin V, Nikiforova S, Martens S (2012) A genome-wide phylogenetic reconstruction of family 1UDP-glycosyltransferases revealed the expansion of the family during the adaptation of plants to life on land. The Plant J 69:1030–1042
Chaturvedi P, Misra P, Tuli R (2011) Sterol glycosyltransferases—the enzymes that modify sterols. Appl Biochem Biotechnol 165:47–68. doi:10.1007/s12010-011-9232-0
Chevallet M, Luche S, Rabilloud T (2006) Silver staining of proteins in polyacrylamide gels. Nature Protoc 1:1852–1858
Ward ER et al (1991) Coordinate gene activity in response to agents that induce systemic acquired resistance. Plant cell 3:1085–1094
Gachon CM, Langlois-Meurinne M, Saindrenan P (2005) Plant secondary metabolism glycosyltransferases: The emerging functional analysis. Trends Plant Sci 10:542–549. doi:10.1016/j.tplants.2005.09.007
Gupta NSP, Santosh Kumar RJ, Vishwakarma RK, Khan BM (2012) Functional characterization and differential expression studies of squalene synthase from Withania somnifera. Mol Biol Rep 39:8803–8812
Hong BS, Kim JH, Kim NY, Kim BG, Chong Y, Ahn JH (2007) Characterization of uridine-diphosphate dependent flavonoid glucosyltransferase from Oryza sativa. J Biochem Mol Biol 40:870–874
Kim JH et al (2006) Characterization of flavonoid 7-O-glucosyltransferase from Arabidopsis thaliana. Biosci Biotechnol Biochem 70:1471–1477
Ko JH, Kim BG, Hur HG, Lim Y, Ahn JH (2006) Molecular cloning, expression and characterization of a glycosyltransferase from rice. Plant Cell Rep 25:741–746. doi:10.1007/s00299-006-0119-4
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Larkin MA et al (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948. doi:10.1093/bioinformatics/btm404
Li Y, Baldauf S, Lim EK, Bowles DJ (2001) Phylogenetic analysis of the UDP-glycosyltransferase multigene family of Arabidopsis thaliana. J Biol Chem 276:4338–4343. doi:10.1074/jbc.M007447200M007447200
Liuji Wu XZ, Xintao W, Anguo S, Jun Z, Shunxi W, Yanhui C (2012) Comparative proteomic analysis of the effects of salicylic acid and abscisic acid on maize (Zea mays L.) Leaves. Plant Mol Biol Rep. doi:10.1007/s11105-012-0522-7
Mackenzie PI et al (2005) Nomenclature update for the mammalian UDP glycosyltransferase (UGT) gene superfamily. Pharmacogenet Genomics 15:677–685
Noguchi A, Saito A, Homma Y, Nakao M, Sasaki N, Nischino T, Takahashi S, Nakayama T (2007) A UDP-glucose:Isoflavone 7-O-glucosyltransferase from the roots of soybean (Glycine max) seedlings. Purification, gene cloning, phylogenetics, and an implication for an alternative strategy of enzyme catalysis. J Biol Chem 282:23581–23590
Noguchi A, Sasaki N, Nakao M, Fukami H, Takahashi S, Nishino T, Nakayama T (2008) cDNA cloning of glycosyltransferases from Chinese wolfberry (Lycium barbarum L.) fruits and enzymatic synthesis of a catechin glucoside using a recombinant enzyme (UGT73A10). J Mol Catal B-Enzym 55:84–92. doi:10.1016/j.molcatb.2008.02.001
Ono E, Homma Y, Horikawa M, Kunikane-Doi S, Imai H, Takahashi S, Kawai Y, Ishiguro M, Fukui Y, Nakayama T (2010) Functional differentiation of the glycosyltransferases that contribute to the chemical diversity of bioactive flavonol glycosides in grapevines (Vitis vinifera). The Plant Cell 22:2856–2871
Paquette SMB, Bak S (2003) On the origin of family 1 plant glycosyltransferases. Phytochemistry 62:399–413
Vishwakarma RK, Singh R, Sonawane PD, Srivastava S, Kumari U, Santosh Kumar RJ, Khan BM (2012) Molecular cloning, biochemical characterization, and differential expression of an acetyl-CoA C-acetyltransferase gene (AACT) of Brahmi (Bacopa monniera). Plant Mol Biol Rep. doi:10.1007/s11105-012-0523-6
Saritha KVNC (2007) vitro flowering of Withania somnifera Dunal.—an important antitumor medicinal plant. Plant Sci 172:847–851
Shao H, He X, Achnine L, Blount JW, Dixon RA, Wang X (2005) Crystal structures of a multifunctional triterpene/flavonoid glycosyltransferase from Medicago truncatula. Plant Cell 17:3141–3154. doi:10.1105/tpc.105.035055
Sharma LK, Madina BR, Chaturvedi P, Sangwan RS, Tuli R (2007) Molecular cloning and characterization of one member of 3beta-hydroxy sterol glucosyltransferase gene family in Withania somnifera. Arch Biochem Biophys 460:48–55. doi:10.1016/j.abb.2007.01.024
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599. doi:10.1093/molbev/msm092
Udayakumar R, Kasthurirengan S, Mariashibu TS, Rajesh M, Anbazhagan VR, Kim SC, Ganapathi A, Choi CW (2009) Hypoglycaemic and hypolipidaemic effects of Withania somnifera root and leaf extracts on alloxan-induced diabetic rats. Int J Mol Sci 10:2367–2382. doi:10.3390/ijms10052367
Vogt T, Jones P (2000) Glycosyltransferases in plant natural product synthesis: Characterization of a supergene family. Trends Plant Sci 5:380–386. doi:10.1016/s1360-1385(00)01720-9
Wang J, Hou B (2009) Glycosyltransferases: Key players involved in the modification of plant secondary metabolites. Frontiers Biol China 4:39–46. doi:10.1007/s11515-008-0111-1
Acknowledgments
RJSK is thankful to Council of Scientific and Industrial Research (CSIR) for senior research fellowship. We are thankful to Dr. Bhushan Dholakia and Dr. D. Shanmugam for comments on manuscript. We are grateful to Dr. Shrinivas Hotha for providing LC-MS facility and his student Ashif Shaikh for help in LC-MS experiment and analysis. This work was supported by grants from network project CSIR (NWP 0008), India.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Fig. S1
Nucleotide and deduced amino acid sequences of WsGT from Withania somnifera with 5′ (18bp) and 3′ (158bp) UTRs followed by a poly-A tail. Start codon (ATG) and stop codon (TAA) highlighted in bold and poly-A signal site (AATAAA) are underlined (JPEG 1661 kb)
Fig. S2
Amino acid sequence alignment of WsGT protein with reported glycosyltransfearses. Asterisk indicates identical or conserved residues; colon indicates a conserved substitution; dot indicates a semi-conserved substitution. The conserved PSPG box found in glycosyltransferases is highlighted with an underline (JPEG 2618 kb)
Fig. S3
SDS-PAGE analysis of WsGT expressed in E. coli. Lane M- protein molecular size marker; lane 1 & 2- Ni-NTA purified WsGT protein; lane 3, 4 & 5- anion exchange purified WsGT. The position of purified protein is about 52kDa (JPEG 341 kb)
Fig. S4
The LC-MS spectrum of the substrate and product in the reaction of WsGT enzyme with (a) diadzein and (b) genistein (JPEG 617 kb)
Rights and permissions
About this article
Cite this article
Kumar, R.J.S., Ruby, Singh, S. et al. Functional Characterization of a Glucosyltransferase Specific to Flavonoid 7-O-Glucosides from Withania somnifera . Plant Mol Biol Rep 31, 1100–1108 (2013). https://doi.org/10.1007/s11105-013-0573-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-013-0573-4