Abstract
Key message
A dehydration-inducible Arabidopsis CIN-like TCP gene, TCP13, acts as a key regulator of plant growth in leaves and roots under dehydration stress conditions.
Abstract
Plants modulate their shape and growth in response to environmental stress. However, regulatory mechanisms underlying the changes in shape and growth under environmental stress remain elusive. The CINCINNATA (CIN)-like TEOSINTE BRANCHED1/CYCLOIDEA/PCF (TCP) family of transcription factors (TFs) are key regulators for limiting the growth of leaves through negative effect of auxin response. Here, we report that stress-inducible CIN-like TCP13 plays a key role in inducing morphological changes in leaves and growth regulation in leaves and roots that confer dehydration stress tolerance in Arabidopsis thaliana. Transgenic Arabidopsis plants overexpressing TCP13 (35Spro::TCP13OX) exhibited leaf rolling, and reduced leaf growth under osmotic stress. The 35Spro::TCP13OX transgenic leaves showed decreased water loss from leaves, and enhanced dehydration tolerance compared with their control counterparts. Plants overexpressing a chimeric repressor domain SRDX-fused TCP13 (TCP13pro::TCP13SRDX) showed severely serrated leaves and enhanced root growth. Transcriptome analysis of TCP13pro::TCP13SRDX transgenic plants revealed that TCP13 affects the expression of dehydration- and abscisic acid (ABA)-regulated genes. TCP13 is also required for the expression of dehydration-inducible auxin-regulated genes, INDOLE-3-ACETIC ACID5 (IAA5) and LATERAL ORGAN BOUNDARIES (LOB) DOMAIN 1 (LBD1). Furthermore, tcp13 knockout mutant plants showed ABA-insensitive root growth and reduced dehydration-inducible gene expression. Our findings provide new insight into the molecular mechanism of CIN-like TCP that is involved in both auxin and ABA response under dehydration stress.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants are exposed to various environmental stresses, such as drought, high salt, and low temperature. To withstand these abiotic stresses, plants have evolved numerous mechanisms of adaptation. Inhibition of shoot growth under water deficit conditions is often observed, which improves water balance and stress tolerance, thus ensuring plant survival (Claeys and Inze 2013). Environmental cues such as water and nutrient availability, salt, temperature, and light conditions have profound effects on leaf and root architecture (Ding and De Smet 2013). Morphological changes in leaves and roots allow plants to acclimate to and tolerate environmental stress conditions. When the effect of environmental stress is temporary, continued restriction of growth can lead to a competitive disadvantage and unnecessary yield losses. Therefore, plants have evolved several tightly regulated mechanisms to balance growth and survival in response to severe abiotic stress conditions. Although the regulation of shoot and root growth under water deficit conditions is important for plant survival, the molecular mechanisms regulating morphological changes in these organs are not well understood.
Plants utilize transcriptional modulation to regulate the balance between growth and survival under abiotic stress conditions. Previous studies have shown that genes performing different functions are either upregulated or downregulated under stress conditions (Dubois et al. 2013; Kreps et al. 2002; Maruyama et al. 2004, 2009; Seki et al. 2002a, 2002b; Urano et al. 2017, 2009). During evolution, the number of transcription factors (TFs) involved in the adaptation of plants to complex environmental stresses has increased. Several different TF families are known to be involved in abiotic stress responses, including dehydration-responsive element-binding (DREB) protein, basic leucine zipper (bZIP) domain, ethylene-responsive element-binding factor (ERF), zinc-finger, WRKY, MYB, and basic helix-loop-helix (bHLH) families. These TFs mainly function as transcriptional activators of downstream genes involved in stress responses and tolerance (Chen et al. 2002; Kodaira et al. 2011; Schommer et al. 2008; Yamaguchi-Shinozaki and Shinozaki 2006). Recently, the results of high-throughput techniques, such as chromatin immunoprecipitation sequencing (ChIP-seq) and DNA affinity purification sequencing (DAP-seq), showed that differential binding of multiple TFs to downstream gene promoters ensures robust responsiveness of downstream genes to the environmental stimulus (O'Malley et al. 2016; Song et al. 2016; Sullivan et al. 2014).
The CINCINNATA-like (CIN-like) TEOSINTE BRANCHED1/CYCLOIDEA/PCF (TCP) TF family plays essential roles in the determination of leaf size and shape (Nath et al. 2003; Palatnik et al. 2003). TCPs harbor a conserved noncanonical bHLH domain, which mediates their binding to DNA or interaction with other proteins (Cubas et al. 1999; Kosugi and Ohashi 2002). The TCP proteins are grouped into two subclasses, class I and class II, based on sequence similarity (Martin-Trillo and Cubas 2010). A total of 13 class I and 11 class II TCPs have been identified in Arabidopsis thaliana. Among the 11 class II TCP genes, TCP2, TCP3, TCP4, TCP5, TCP10, TCP13, TCP17, and TCP24 belong to the CIN-like TCP family. Simultaneous disruption of multiple CIN-like TCP genes greatly affects leaf development (Koyama et al. 2010; Schommer et al. 2008). TCP3 directly activates the expression of microRNA164 (miR164), ASYMMETRIC LEAVES1 (AS1), INDOLE-3-ACETIC ACID3/SHORT HYPOCOTYL2 (IAA3/SHY2), and several auxin-inducible genes including SMALL AUXIN UP RNA (SAUR) proteins, PIN FORMED (PIN) family of auxin efflux carriers, and LATERAL ORGAN BOUNDARIES (LOB) DOMAIN (LBD) TFs (Koyama et al. 2010). These target genes of TCP3 act as negative regulators of CUP-SHAPED COTYLEDON (CUC) genes for regulating leaf differentiation (Koyama et al. 2010). TCP4 directly activates the expression of HAT2, a HD-ZIP II TF to induce the maturation of leaf pavement cells via both auxin-dependent and independent pathways (Challa et al. 2019). TCP5 controls leaf margin development by regulating KNAT3, a KNOTTED1-like homeobox (KNOX) gene and SAW1, a BEL-like transcription factors (Yu et al. 2020). CIN-like TCP genes also regulate jasmonic acid (JA) and flavonol biosynthesis. JA biosynthesis is mediated by Lipoxygenase2 (LOX2), which is upregulated by TCP3 (Koyama et al. 2007) and TCP4 (Schommer et al. 2008), and downregulated in jaw-d mutants (Schommer et al. 2008). TCP3 also interacts with the R2R3-MYB protein MYB12, which promotes flavonoid biosynthesis and represses auxin signaling (Li and Zachgo 2013). In addition, CIN-like TCP genes are involved in the control of axillary bud outgrowth. The jaw‐D, tcp5 and tcp5/13/17 mutant plants show a significant reduction in the number of secondary branches (van Es et al. 2019).
The activity of five CIN-like TCPs, including TCP2, TCP3, TCP4, TCP10, and TCP24, during leaf development is tightly regulated by miR319 at the post-transcriptional level. Expression of TCPs carrying synonymous mutations, responsible for the resistance to miR319-mediated cleavage, causes severe defects in leaf morphology or seedling death in Arabidopsis, and the transition from compound leaves to simple leaves in tomato (Solanum lycopersicum) (Ori et al. 2007; Palatnik et al. 2003, 2007). The activity of CIN-like TCPs is also modulated at the protein level through protein–protein interactions. ARMADILLO BTB ARABIDOPSIS PROTEIN1 (ABAP1) interacts with TCP24 to modify its activity for the regulation of leaf cell proliferation (Masuda et al. 2008). The SWI/SNF chromatin remodeling ATPase, BRAHMA (BRM), modulates the activity of CIN-like TCPs to reduce the cytokinin sensitivity of leaves by increasing the expression of the negative cytokinin regulator, ARABIDOPSIS RESPONSE REGULATOR16 (ARR16) (Efroni et al. 2013). The TCP INTERACTOR-CONTAINING EAR MOTIF PROTEIN1 (TIE1), a transcriptional repressor, as well as TOPLESS (TPL) and TIE1-ASSOCIATED RING-TYPE E3 LIGASE1 (TEAR1) regulate leaf development by physically interacting with CIN-like TCPs (Tao et al. 2013; Zhang et al. 2017). However, the role of CIN-like TCP in the regulation of plant growth under abiotic stress conditions is not well characterized.
In this study, we show that an Arabidopsis CIN-like TCP gene, TCP13, is significantly induced by dehydration. We show that TCP13 plays important roles in regulating the growth of leaves and roots under abiotic stress. Arabidopsis plants overexpressing TCP13 (35Spro::TCP13OX) exhibited leaf rolling and leaf growth inhibition, abscisic acid (ABA)-sensitive root growth, and elevated dehydration stress tolerance. Additionally, tcp13 knockout mutant plants showed ABA-insensitive root growth and reduced dehydration-inducible gene expression, which supports the idea that TCP13 acts downstream of ABA signaling pathway under dehydration stress. TCP13 also positively regulates the dehydration-inducible auxin-regulated genes, IAA5 and LBD1. Overall, our results suggest that TCP13 is an important component of a regulatory module that controls plant growth under dehydration stress.
Materials and methods
Plant materials and stress treatments
Arabidopsis thaliana ecotype Columbia (Col-0; WT) was used for the generation of transgenic lines used in this study. Unless otherwise stated, plants were grown on MS medium (Murashige and Skoog 1962), supplemented with 3% sucrose and 0.8% agar (MS-agar), or in soil (Dio Professional for grafting, Innovex Co., Ltd., Tokyo) at 22 °C under long-day conditions (a 16 h light/8 h dark photoperiod and 60 ± 10 μmol photons m−2 s−1 light intensity, as described previously (Urano et al. 2009; Urano et al. 2004)). The T-DNA insertion tcp5 (SM_3_29639), tcp13 (GK_182B12), and tcp17 (SALK_148580) mutant lines were obtained from the Arabidopsis Biological Resource Center (ABRC), OH, USA, and the site of T-DNA insertion site in each of these mutants has been described previously (Koyama et al. 2007, 2010). The tcp13tcp15tcp17 triple mutant was generated by crossing the single mutants described previously (Koyama et al. 2017). To perform ABA or salt (NaCl) stress treatments, 2-week-old whole WT plants were transferred from the agar medium to water (control) or to water containing 10 μM ABA or175 mM NaCl for 3–6 h. To induce dehydration stress, whole plants were transferred on to a parafilm (Parafilm M PM999, Bemis Company, Neenah, WI) for 3–6 h (WT) or 3–8 h (tcp13 and WT).
Plasmid construction and plant transformation
To generate 35Spro::TCP13OX, 35Spro::TCP5OX, and 35Spro::TCP17OX constructs, coding sequences (CDSs) of TCP13, TCP5, and TCP17, respectively, were amplified by PCR using sequence-specific primers and cloned into the EcoRV site of pGKX, as described previously (Maruyama et al. 2009). To generate the 35Spro::TCP13SRDX and 35Spro::TCP13-sGFP constructs, the CDS of TCP13 minus the stop codon was cloned into the SmaI site of pGKX-SRDX and pGKX-sGFP, respectively, as described previously (Fujita et al. 2005; Maruyama et al. 2009; Qin et al. 2008). To construct the TCP13pro::TCP13SRDX plasmid, the TCP13 promoter (∼ 1000 bp upstream of the transcription start site) was PCR amplified and cloned into 35Spro::TCP13SRDX between KpnI and XbaI restriction sites. To construct the TCP13pro::GUS plasmid, first the pGKX-GUS construct was generated by cloning the GUS reporter gene amplified from the pBI101 vector (Fujita et al. 2005; Maruyama et al. 2009; Qin et al. 2008; Urano et al. 2004) into the pGKX-sGFP plasmid between BamHI and EcoRV restriction sites. Then, the TCP13 promoter region was cloned into the pGKX-GUS plasmid between KpnI and XbaI restriction sites. These constructs were introduced into Agrobacterium tumefaciens strain C58, which was then used to transform Arabidopsis plants via vacuum infiltration (Bechtold and Pelletier 1998). Seeds of T2 or T3 plants were used for subsequent experiments.
Histochemical GUS staining and GFP detection in protoplasts
Histochemical GUS staining was performed as described previously (Urano et al. 2004), and GUS was observed using an M205C stereomicroscope equipped with a DFC490 digital color camera (Leica Microsystems). To determine the subcellular localization of TCP13, plasmid DNA (5 μg) isolated from 35Spro::TCP13-sGFP plants was transfected into Arabidopsis mesophyll protoplasts, and GFP was detected under a confocal laser scanning microscope (LSM510; Zeiss). Arabidopsis protoplast isolation and PEG-calcium transfection were performed as described previously (Yoo et al. 2007).
ABA sensitivity of transgenic and mutant plants
The ABA sensitivity test was performed as described previously (Yoshida et al. 2010) with minor modification. 35Spro::TCP13OX and control plants were grown in plates containing 1/2 MS medium supplemented with 1% sucrose, 30 mg/L kanamycin, and 0.8% agar or tcp13 mutant and WT (Col-0) plants were grown in plates containing half-strength MS (1/2 MS) medium supplemented with 1% sucrose and 0.8% agar at 22 °C under long-day conditions. To perform the ABA sensitivity test, 5-day-old plants were transferred to plates containing 1/2 MS medium supplemented with 1% sucrose, 1.2% agar, and ABA (0, 50, or 100 μM for 35Spro::TCP13OX and control plants; 0, 5, or 10 μM for tcp13 and WT plants). After 7 days of incubation, the root elongation length of 7 plants per experiment was measured.
Osmotic stress sensitivity test of transgenic and mutant plants
The osmotic stress sensitivity test was performed using the 35Spro::TCP13OX and control plants grown in plates containing 1/2 MS medium supplemented with 1% sucrose, 30 mg/L kanamycin, and 0.8% agar or tcp13 mutant, tcp13tcp15tcp17 triple mutant and WT (Col-0) plants grown on1/2 MS medium supplemented with 1% sucrose and 0.8% agar at 22 °C under long-day conditions. Plates were overlaid with a nylon mesh (Prosep; pore size = 20 μm) to prevent roots from growing into the medium in a plate (diameter = 150 mm). At 9 days after sowing (DAS), when the third leaf was fully expanded, the nylon mesh was gently lifted using forceps, and seedlings were transferred to plates containing 1/2 MS medium (control) or 1/2 MS medium supplemented with 25 mM mannitol (Sigma-Aldrich). After 13 days of incubation, the leaf area of five leaves per line was measured using ImageJ.
Dehydration tolerance test
The dehydration tolerance test was performed using the 35Spro::TCP13OX and control plants grown in soil at 22 °C under long-day conditions. One pot contained 5 plants of each line. 3 pots of 35Spro::TCP13OXa and b, and 6 pots of control plants were transferred to empty tray and exposed dehydration stress by withholding watering for 14 days in 45–60 relative humidity. After dehydration treatment, plants were grown for 7 days with well-water conditions and evaluated the survival rate of plants. We performed dehydration tolerance test three times. Total of 90 plants of control, 45 plants of 35Spro::TCP13OXa and b were tested.
Transcriptome analysis using an oligo DNA array
The Agilent Arabidopsis 4 Oligo Microarray (Agilent Technologies) containing 21,500 probes was used to identify genes downstream of TCP13. Total RNA was isolated from 2-week-old TCP13pro::TCP13SRDX whole plants using extraction buffer (0.2 M Tris-HCl [pH 9.0], 0.4 M LiCl, 25 mM EDTA, and 1% SDS). Total RNA of 10 plants of each transgenic line was pooled. This procedure was repeated to produce two biologically independent RNA pools per transgenic line. The Cy3- and Cy5-labeled cRNA of each transgenic line and control sample was hybridized to the microarray. Additionally, the color swapping experiment was performed as a technical replicate. After hybridization, the microarray slides were scanned (scanner model G2505C; scan control software version A.8.5.1; Agilent Tech, Inc., Santa Clara, CA, USA), and data were analyzed using the Feature Extraction software (version 10.10.1.1; Agilent). Raw data were analyzed using the GeneSpring GX software (version 12.0; Agilent). Expression log ratios and Benjamini–Hochberg FDR p-values were calculated using GeneSpring GX. The microarray design and data were deposited at ArrayExpress (Accession Number E-MTAB-9336).
Gene expression analysis by qRT-PCR
Gene expression was analyzed by qRT-PCR as described previously (Urano et al. 2017). The AGI code- and gene-specific primers were designed based on the sequence of a single exon using Primer Express 2.0 (Applied Biosystems, Foster City, CA, USA) (Methods S1). All qRT-PCR reactions were performed in three technical replicates, and mRNA levels of genes were normalized relative to the constitutive control, At2g32170.
Transactivation analysis of the AHG3, Gols2, IAA5, and LBD1 promoter
Protoplasts were isolated from peeled rosette leaves of 3–4-week-old WT and tcp13 mutant Arabidopsis plants using the Tape Arabidopsis Sandwich method (Wu et al. 2009), as described previously (Sakamoto et al. 2016; Yoshida et al. 2013). The reporter construct containing the firefly (Photinus pyralis) LUC gene driven by the AHG3, Gols2, IAA5, or LBD1 promoter was co-transfected into the WT and tcp13 mutant protoplasts using the PEG transfection method, as described previously (Sakamoto et al. 2016; Yoshida et al. 2013), along with a construct containing the modified Renilla reniformis luciferase gene driven by the CaMV 35S promoter (phRLHSP; internal control). The transfected protoplasts were incubated at 22 °C in the dark in medium supplemented with or without 5 μM ABA or 1 μM IAA for 16–18 h. The dual-LUC assay was carried out using the Pikka Gene Dual Assay Kit (Toyo Ink, Inc., Tokyo, Japan). The reporter activity was normalized with the activity of the Renilla luciferase gene.
Results
TCP13 shows dehydration-inducible expression
Water deficit stress induces morphological changes in plants to ensure survival. To identify key factors that control these morphological changes under dehydration stress, we analyzed the expression patterns of Arabidopsis TCP genes in leaves under dehydration stress using our previously published microarray data (Urano et al. 2009). Of the 24 Arabidopsis TCP genes on the Agilent oligo array, 11 were expressed (Fig. S1). This microarray analysis showed that only TCP13 was upregulated under dehydration stress, whereas other TCPs were downregulated. The up-regulation of TCP13 under dehydration stress is also supported by the Arabidopsis RNA-seq Database (http://ipf.sustech.edu.cn/pub/athrna/; Zhang et al. 2020) (Table S1).
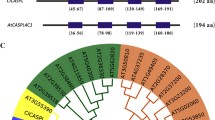
Amino acid sequence alignment of Arabidopsis class II TCPs suggests that TCP13 is relatively similar to TCP5 and TCP17 (Fig. 1a). TCP5, TCP13, and TCP17 do not contain the miR319A/JAW target sequence (Palatnik et al. 2003), and all three TCPs have been previously shown to be involved in leaf differentiation, which is a common function of CIN-like TCPs (Koyama et al. 2007, 2010). Figure 1b shows the results of quantitative real-time PCR (qRT-PCR) analysis of TCP5, TCP13, and TCP17 in Arabidopsis plants exposed to ABA, high salt, or dehydration stress for 3 or 6 h. The expression of TCP13 was highly upregulated during the early phase of dehydration stress, and slightly upregulated by exogenous ABA and high salinity. In contrast, the expression of TCP5 decreased during the early phase of dehydration. Expression of TCP17 was not affected by any of the three abiotic stresses (Fig. 1B).
Expression pattern of TCP13 in wild-type (WT; Col-0) Arabidopsis plants under different abiotic stresses. a Phylogenetic analysis of Arabidopsis class II TCP proteins and representative members of Antirrhinum majus CINCINNATA (CIN) and CYCLOIDEA (CYC) (Luo et al. 1996) families. The phylogenetic tree was constructed using the neighbor-joining method with the MEGA X software (Kumar et al. 2018). Rice (Oryza sativa) class I TCPs, PROLIFERATING CELL FACTOR1 (PCF1), and PCF2 (Kosugi and Ohashi 2002) were used as an outgroup. The underlined names represent proteins containing the miR319A/JAW target sequence (Palatnik et al. 2003). Black dots indicate proteins involved in leaf differentiation (Koyama et al. 2007, 2010). b Quantitative real-time PCR (qRT-PCR) analysis of the relative transcript levels of TCP13, TCP5, and TCP17. Plants were treated with water (control), dehydration stress, 10 μM abscisic acid (ABA), or salt stress (175 mM NaCl), and gene expression was examined at 3 and 6 h post-treatment. Data represent mean ± standard deviation (SD; n = 3). Asterisks indicate significant differences (*P < 0.05, **P < 0.01; one-way ANOVA with Welch’s t test). c Schematic showing the structure of the 1000 bp promoter of TCP13, TCP5, and TCP17. Red and blue circles indicate the ABA-responsive element (ABRE)-related sequences (ACGTGG/T) (Busk and Pages 1998; Yamaguchi-Shinozaki and Shinozaki 2006) and GMSAUR (CATATG) motif found in the promoter of the soybean SAUR15A gene (Xu et al. 1997), respectively
To investigate why the response of TCP13 to abiotic stress was different from that of TCP5 and TCP17, we analyzed the promoter sequences of all three genes. The TCP13 promoter contained two typical ABRE motifs (ACGTGG) (Fig. 1c). By contrast, TCP5 and TCP17 promoters lacked the ABRE motif but contained an auxin-responsive element, named GMSAUR (CATATG), which was found in the promoter of an auxin-responsive gene, SAUR15A, of soybean (Glycine max) (Xu et al. 1997). These results suggest that TCP13 might perform a unique function under dehydration stress via ABA signaling. In addition, amino acid sequence alignment of class II TCPs of selected vascular land plants such as Arabidopsis, Brassica napus, soybean, alfalfa (Medicago truncatula), rice (Oryza sativa), maize (Zea mays), and a bryophyte (Physcomitrella patens) was shown in Fig. S2.
Tissue-specific expression of TCP13, and subcellular localization of TCP13 protein
To analyze the tissue-specific expression profile of TCP13, we performed histochemical staining of transgenic Arabidopsis plants expressing the β-glucuronidase (GUS) gene under the control of the TCP13 promoter (TCP13pro::GUS) (Fig. 2a–c), and analyzed the expression of TCP13 in leaves and roots of wild-type (WT) plants by qRT-PCR (Fig. 2d). GUS activity was detected in the leaves, but not in the roots, of 5- and 10-day-old plants (Fig. 2a and b). GUS staining was detected in cotyledon and true leaves (Fig. 2a and b). GUS staining was specifically detected in expanding cells (upper region) but not in dividing cells (lower region) in young leaves (Fig. 2c). Additionally, qRT-PCR analysis showed that the expression level of TCP13 was high in leaves and low in roots (Fig. 2d). Tissue-specific TCP13 expression data obtained from the Arabidopsis EFP browser (http://bar.utoronto.ca/efp/cgi-bin/efpWeb.cgi) (Winter et al. 2007) showed that TCP13 is expressed in mesophyll cells, but not in guard cells, of leaves (Fig. S3a), and in the endodermis and cortex of roots under salt stress (Fig. S3b).
Expression pattern of TCP13 in Arabidopsis plants at different developmental stages, and subcellular localization of the TCP13 protein. a–c Histochemical staining showing β-glucuronidase (GUS) reporter activity (blue) under the control of the TCP13 promoter in seedlings (a), leaves (b), and in an enlarged view of the true leaves (c). Scale bars = 1 mm (a and c); 5 mm (b). d Analysis of the relative transcript level of TCP13 in leaves and roots by qRT-PCR. Transcript levels of genes were normalized relative to the constitutive control, At2g32170 (Czechowski et al. 2005). Data represent mean ± SD (n = 3). Asterisks indicate significant differences (**P < 0.001; one-way ANOVA with Welch’s t test). e Subcellular localization of TCP13-sGFP fusion protein in Arabidopsis mesophyll protoplasts. Scale bars = 5 μm
To determine the subcellular localization of TCP13, we fused the full-length TCP13 gene to a synthetic green fluorescent protein (sGFP) gene (TCP13-sGFP), and introduced the construct into Arabidopsis leaf protoplasts via polyethylene glycol (PEG)-mediated transfection. The TCP13-sGFP fusion protein was transiently expressed in the nuclei (Fig. 2e). Together, these data suggest that TCP13 exhibits strong expression in leaves, and the encoded protein localizes to the nuclei.
Characterization of TCP13 overexpression (TCP13OX) lines and tcp5tcp13tcp17 triple mutant plants under normal growth conditions
First, we analyzed transgenic plants overexpressing TCP13 cDNA under the control of the constitutive Cauliflower mosaic virus (CaMV) 35S promoter (35Spro::TCP13OX). The 35Spro::TCP13OX seedlings showed longer hypocotyls and shorter roots than plants transformed with the empty vector (control) (Fig. 3a–c). True leaves of 35Spro::TCP13OX plants were narrower than those of control plants and showed downward rolling (Fig. 3d, e, i, and j). We also examined the effect of TCP13 repression on plant morphology. The Chimeric REpressor Silencing Technology (CRES-T) is a gene silencing system, in which a TF fused to the EAR-motif repression domain (Leu-Asp-Leu-Asp-Leu-Glu-Leu-Arg-Leu-Gly-Phe-Ala;SRDX) dominantly represses the transcription of its target genes, regardless of the absence or presence of endogenous and functionally redundant TFs (Hiratsu et al. 2003). We generated transgenic plants expressing SRDX-fused TCP13 cDNA (TCP13SRDX). Transgenic Arabidopsis plants expressing the chimeric repressor under the control of the constitutive CaMV 35S promoter (35Spro::TCP13SRDX) produced ectopic shoots and failed to grow when transferred to soil (Fig. 3f). Transgenic seedlings expressing the chimeric repressor under the control of the TCP13 promoter (TCP13pro::TCP13SRDX) showed greater root growth (Fig. 3a and c) and wavy and serrated rosette leaves (Fig. 3g, m, and n) compared with control plants (Fig. 3d and j). Rosette leaves of tcp13 mutant plants showed no severe phenotypic abnormality (Fig. 3k) compared with those of WT plants (Fig. 3l). By contrast, tcp5tcp13tcp17 triple knockout mutant plants (tcp5/13/17) showed shorter petioles and smaller leaves (Fig. 3h and o), with slightly wavy and serrated margins (Fig. 3q), compared with the respective control (Fig. 3d) or WT plants (Fig. 3p and r), suggesting that TCP13, together with TCP5 and TCP17, regulates leaf morphology.
Characterization of TCP13 overexpression (TCP13OX) lines and tcp13 loss-of-function mutant plants at different developmental stages. a Root growth of 7-day-old 35Spro::TCP13OX and TCP13pro::TCP13SRDX seedlings. Scale bars = 5 mm. b and c Hypocotyl (b) and root (c) length of 7-day-old 35Spro::TCP13OX and TCP13pro::TCP13SRDX seedlings. Data represent mean ± SD (n = 6). An asterisk shows that the indicated mean is significantly different from the mean value of the control plant (**P < 0.01, ***P < 0.001; one-way ANOVA with Welch’s t test). d–h Images of 14-day-old empty vector control (d), 35Spro::TCP13OX (e), 35Spro::TCP13SRDX (f), TCP13pro::TCP13SRDX (g), and tcp5/13/17 triple mutant (h) plants grown in agar. Scale bar = 5 mm (d, e, g, and h); 1 mm (f). i–r Images of 21-day-old 35Spro::TCP13OX (i) and corresponding empty vector control (j) plants; 28-day-old tcp13 mutant (k) and corresponding WT (l) plants; 28-day-old TCP13pro::TCP13SRDX (m and n) plants; 21-day-old tcp5/13/17 (o) and corresponding WT (p) plants; and 28-day-old tcp5/13/17 mutant (q) and corresponding WT (r) plants. Scale bars = 10 mm
We also generated transgenic Arabidopsis plants overexpressing TCP5 and TCP17 under the control of the constitutive CaMV 35S promoter (35Spro::TCP5OX and 35Spro::TCP17OX, respectively). Transgenic 35Spro::TCP5OX and 35Spro::TCP17OX seedlings showed longer hypocotyls than empty vector-transformed control plants (Fig. S4a). This phenotype of 35Spro::TCP5OX and 35Spro::TCP17OX seedlings was similar to that of transgenic seedlings overexpressing a mutant form of TCP3 (35Spro::mTCP3OX), in which the target site of miR319/JAW was replaced with a non-target sequence (Koyama et al. 2007), and to that of previously generated transgenic seedlings overexpressing TCP5 (35Spro::TCP5OX) (Han et al. 2019) and TCP17 (35Spro::TCP17OX) (Han et al. 2019; Zhou et al. 2019). Furthermore, TCP5, TCP13, and TCP17 were shown to play basic roles in promoting thermo-responsive hypocotyl growth by positively regulating PIF4 activity (Han et al. 2019; Zhou et al. 2019). In contrast to 35Spro::TCP13OX plants, the 35Spro::TCP5OX and 35Spro::TCP17OX plants showed no change in root growth in comparison with control plants (Fig. S4b). These results suggest that inhibition of root growth is a unique function of TCP13 among CIN-like TCP genes.
TCP13 inhibits leaf and root growth in response to ABA and osmotic stress, and is involved in tolerance to dehydration stress
The TCP13 gene is significantly induced by dehydration stress and slightly induced by high salinity and ABA treatments (Fig. 1b). To analyze the function of TCP13 in ABA response, we examined the response of the 35Spro::TCP13OX transgenic seedling and tcp13 knockout mutant seedlings to the ABA treatment (Fig. 4). To carry out the ABA sensitivity test, the 35Spro::TCP13OX and tcp13 plants grown on agar plates for 5 days were transferred to plates containing agar supplemented with or without ABA. After 7 days, the root length of these seedlings was measured (Fig. 4c), and the root growth in the presence of ABA was compared with that in the absence of ABA (Fig. 4d). The 35Spro::TCP13OX plants showed reduced root growth compared with control plants in the absence and presence of ABA treatment (Fig. 4a and c). The enhanced root growth retardation relative to the control plants was detected in the 35Spro::TCP13OX plants under ABA treatment (Fig. 4d). In contrast, the WT and tcp13 mutant plants showed similar growth in the absence of ABA, whereas the tcp13 mutant plants showed greater root elongation than WT plants in response to ABA (Fig. 4b, c and d). These results suggest that TCP13 negatively regulates root growth in response to the ABA treatment.
Characterization of TCP13OX and tcp13 mutant plants in the ABA treatment. Five-day-old seedlings were transferred to control and ABA-containing medium and grown for 7 days and root length was measured. Root elongation under the ABA treatment was compared with that under the control treatment, and the rate of root elongation was calculated. a and b Images of the root growth of 35Spro::TCP13OX transgenic and control plants (a) and tcp13 mutant and WT plants (b) in the ABA treatment. Scale bars = 10 mm. c The root length of 35Spro::TCP13OX transgenic and control plants (left panel) and tcp13 mutant and WT plants (right panel) in the ABA treatment. d Relative level of root elongation of 35Spro::TCP13OX transgenic and control plants (left panel) and tcp13 mutant and WT plants (right panel) in the ABA treatment. In each transgenic or mutant plants, average of root length in the control condition was set to 100. Data represent mean ± SD (n = 7). An asterisk shows that the indicated mean is significantly different from the mean value of the wild-type plant under the corresponding condition (***P < 0.001, one-way ANOVA with Welch’s t test)
To analyze the effect of TCP13 on plant response to osmotic stress, we examined changes in the leaf growth of the 35Spro::TCP13OX, tcp13 and tcp5/13/17 knockout mutant plants on agar plates under mannitol stress (Fig. 5). Nine-day-old plants were transferred to plates containing half-strength Murashige and Skoog (1/2 MS) medium or 1/2 MS medium supplemented with 25 mM mannitol, which has been shown to restrict leaf growth by approximately 50% (Skirycz et al. 2011). After 13 days, each rosette leaf was separated (Fig. 5a and d), and size of third (L3), fourth (L4) and fifth (L5) leaf in the mannitol treatment was compared with that in the control treatment (Fig. 5b, c, e, and f). The 35Spro::TCP13OX plants showed growth retardation with smaller leaves compared with the control plants in control and mannitol conditions (Fig. 5 a and b). Especially, leaves of 35Spro::TCP13OXa plants in the mannitol treatment showed severe rolling from the mid-point to the tip (Fig. 5a). The enhanced leaf growth retardation relative to the control plants was detected in L3-L5 of the 35Spro::TCP13OXa and L4 of the 35Spro::TCP13OXb plants under mannitol treatment (Fig. 5c). These results indicate that 35Spro::TCP13OX leaves are hypersensitive to osmotic stress. On the other hand, the tcp13 mutant plants treated with mannitol showed no significant changes in leaf growth compared with control plants (Fig. 5d–f). tcp5/13/17 triple mutant plants showed growth retardation with smaller leaves compared with the control plants in control (L3-5) and mannitol (L4 and L5) conditions (Fig. 5e). The reduced leaf growth retardation relative to the WT plants was detected in L4 of the tcp5/13/17 mutant plants under mannitol treatment (Fig. 5f), indicating that tcp5/13/17 showed insensitive response to osmotic stress. TCP13 probably exhibit functional redundancies between TCP5 and 17 for leaf growth regulation under osmotic stress.
Characterization of 35Spro::TCP13OX, tcp13 and tcp5/13/17 mutant plants under osmotic stress conditions. Nine-day-old 35Spro::TCP13OX and control plants, and tcp13 mutant, tcp5/13/17 mutant and WT plants were transferred to 1/2 MS medium containing 25 mM mannitol for 13 days. The leaf area of third (L3), fourth (L4), and fifth (L5) leaves was measured. Leaf growth under the mannitol treatment was compared with that under the control treatment, and the rate of leaf growth was calculated. a and d Images of cotyledon (C) and true leaves (L) of 35Spro::TCP13OX and control plants (a) and tcp13 mutant, tcp5/13/17 mutant and WT plants (d) exposed to control and mannitol treatments. Scale bars = 5 mm. b and e The leaf area in 35Spro::TCP13OX transgenic and control plants (b) and tcp13 mutant, tcp5/13/17 mutant and WT plants (e) treated with control and mannitol medium. c and f Relative level of leaf growth in 35Spro::TCP13OX transgenic and control plants (c) and tcp13 mutant, tcp5/13/17 mutant and WT plants (f) treated with mannitol. In each transgenic or mutant plants, average of leaf area in the control condition was set to 100. Data represent mean ± SD (n = 5). An asterisk shows that the indicated mean is significantly different from the mean value of the wild-type plant under the corresponding condition (*P < 0.05, **P < 0.01, ***P < 0.001; one-way ANOVA with Welch’s t test)
We then examined the response of 35Spro::TCP13OX plants to severe dehydration stress. Watering of 35Spro::TCP13OX plants was withheld for 14 days, and the survival rate of plants was calculated at 7 days after rewatering. The 35Spro::TCP13OX plants showed greater dehydration stress tolerance, as evident from the reduced downward leaf rolling phenotype and higher survival rate, than control plants (Fig. 6a and b). This result was consistent with a lower reduction in the water content of 35Spro::TCP13OX plants, based on the measurement of the weight of detached leaves, than in the water content of control plants (Fig. 6c). On the other hand, detached leaves of nced3-2 mutant plants (negative control), carrying a knockout mutation in a key gene responsible for dehydration-inducible ABA accumulation, showed a greater reduction in the water content than control plants (Fig. 6c). These results suggest that leaves of 35Spro::TCP13OX plants show less water loss than control plants under dehydration stress.
Characterization of 35Spro::TCP13OX plants under severe dehydration stress. 35Spro::TCP13OX and control plants were grown in soil for 14 days with well-water conditions. One pot contained 5 plants of each line. 3 pots of 35Spro::TCP13OX 35Sa and b, and 6 pots of control plants were transferred to empty tray and exposed dehydration stress by withholding watering for 14 days. After dehydration treatment, plants were grown for 7 days with well-water conditions and evaluated the survival rate of plants. The dehydration tolerance test was performed three times. a Images of plants before dehydration stress (Untreated) and after dehydration (Dehydration). Scale bars = 10 mm. b The survival rate of plants after dehydration. The error bars indicate the SD from three replicates. n = 90 (WT), n = 45 (TCP13OXa) and n = 45 (TCP13OXb), total. c Water loss assay of detached leaves of 5-week-old 35Spro::TCP13OX 35S and control plants. The nced3-2 plants represent a negative control. Data represent mean ± SD (n = 4). An asterisk shows that the indicated mean is significantly different from the mean value of the control plant under the corresponding condition (*P < 0.05, **P < 0.01, ***P < 0.001, one-way ANOVA with Welch’s t test)
Transcriptome analysis of TCP13pro::TCP13SRDX plants
To investigate the transcriptional regulation of TCP13, we performed microarray analysis of whole plants expressing the TCP13pro::TCP13SRDX and focused on genes downregulated in TCP13pro::TCP13SRDX plants (Fig. 7). The results showed that 555 genes were downregulated in TCP13pro::TCP13SRDX plants (fold-change [FC] < 0.5; p < 0.05; false discovery rate [FDR] < 0.0277) (Table S2) compared with control plants.
Transcriptome analysis of genes downregulated in TCP13pro::TCP13SRDX plants. a GO enrichment analyses of downregulated in TCP13pro::TCP13SRDX plants. b Venn diagram showing the overlap between genes downregulated in TCP13pro::TCP13SRDX plants and those upregulated by dehydration stress (Urano et al. 2017). c–e Relative transcript levels of downregulated genes including ABA- and dehydration-inducible genes (c), CIN-like TCP common downstream genes involved in auxin signaling (d), and flavonoid biosynthesis genes (e) in TCP13pro::TCP13SRDX plants. Data represent mean ± SD (n = 4). f Enrichment of hexamer motifs in the promoter regions (~ 1000 bp) of downstream candidate genes of TCP13. Hexamer motifs in the promoter regions of the top 100 downregulated genes in TCP13pro::TCP13SRDX plants were analyzed as described previously (Maruyama et al. 2012). g Comparison of TCP binding and ABRE motifs identified in the promoter regions of downregulated genes in TCP13pro::TCP13SRDX plants. h Comparison of the 1000 bp promoter regions of TCP13SRDX-downregulated genes with those of 23,739 Arabidopsis genes used previously for motif analyses (Maruyama et al. 2012) by χ2 tests. ACGTGT and ACGTGG were used as the ABRE sequences, and GGACCA and TGGTCC were used as the TCP binding motifs. Significant differences between the observed and expected values of the frequency of overlap between the ABRE and TCP binding motifs (**P < 0.01, ***P < 0.001)
Next, we performed Gene Ontology (GO) enrichment analysis (PANTHER Classification System, https://www.arabidopsis.org/tools/go_term_enrichment.jsp) of these 555 genes to determine their potential functions. A total of 48 GO terms were significantly enriched (p < 0.05) (Table S3), and the top 10 categories are shown in Fig. 7a. Among these GO terms, “response to stimulus” was the most highly enriched GO term, followed by “response to abiotic stimulus” and “response to chemical”. Then, we compared the 555 genes downregulated in TCP13pro::TCP13SRDX plants with dehydration-inducible genes reported previously (Urano et al. 2017). The results showed that 230 out of 555 genes were upregulated under dehydration stress in the study of Urano et al. (2017) (FC < 2; p < 0.05) (Fig. 7b). These results suggest that numerous downstream candidates of TCP13 are involved in dehydration stress response.
We further analyzed the expression of dehydration stress-responsive and ABA-regulated genes downregulated in TCP13pro::TCP13SRDX plants in our microarray results (Fig. 7c). These genes included Highly ABA-induced Protein Phosphatase 2C 1 (HAI1), HAI2, ABA-hypersensitive Germination3 (AHG3), Response to Desiccation 20 (RD20), RD26, and RD29B, and key genes involved in raffinose, proline, and ABA metabolism such as Galactinol Synthase2 (GolS2), delta-1-pyrroline-5-carboxylate synthase1 (P5CS1), and 9-cis-epoxycarotenoid dioxygenase3 (NCED3) (Fig. 7c). These genes were not identified as downstream candidates of CIN-like TCPs in previous studies (Koyama et al. 2007, 2010, 2017; Schommer et al. 2008) but were newly identified as downstream candidates of only the dehydration-inducible TCP13. In addition, we also found genes related to the common targets of CIN-like TCP (Koyama et al. 2007, 2010, 2017; Schommer et al. 2008) as being potential downstream candidates of TCP13 in our microarray results (Fig. 7d, e). These included related to JA metabolic gene and auxin-regulated genes such as LOX2, IAA1, IAA5, IAA14, IAA17, IAA34, SAUR17, SAUR34, PIN4, and LBD1 (Fig. 7d), and related to flavonoid metabolic genes such as MYB12, MYB111, Flavonol Synthase1 (FLS1), FLS2, FLS4, Chalcone Isomerase (CHI/TT5), CHI-like (CHIL), flavone 3-hydroxylase (F3H/TT6), UDP-glucosyl transferase 73B2 (UGT73B2), and UGT78D1 (Fig. 7e). We also analyzed the expression of ABA-regulated genes such as AHG3, RD20, Gols2, and NCED3, and auxin-regulated genes, such as IAA5 and LBD1, in 35Spro::TCP13-OX plants. Among these genes, Gols2, NCED3, IAA5, and LBD1 were upregulated in 35Spro::TCP13-OX plants compared with WT plants (Figure S8). These results revealed that expression of TCP13SRDX triggers changes in the expression patterns of downstream genes regulated specifically by TCP13 as well as those regulated by various CIN-like TCPs.
Furthermore, to identify common motifs present in the promoters of the top 100 most downregulated genes in TCP13pro::TCP13SRDX plants, we searched the frequency of hexamer motifs in 1000 bp promoter regions (Table S4), as described previously (Maruyama et al. 2012). Among the top 10 motifs identified in gene promoters, three TCP binding motifs-related sequences (GGACCA) (Schommer et al. 2008) and four ABRE-related sequences (ACGTGG/T) (Busk and Pages 1998; Yamaguchi-Shinozaki and Shinozaki 2006) were found (Fig. 7f). Among all 555 downregulated genes, 218 genes contained the TCP binding motifs in their promoter regions, and 123 genes contained both TCP binding and ABRE motifs (Fig. 7g). The repressive activity of TCP13 induced by the SRDX motif accurately downregulated TCP-regulated downstream genes, most of which were also responsive to ABA signaling or dehydration stress. Although many downstream candidate genes of TCP13 contained the ABRE motifs in their promoters, TCP13 probably does not directly bind to these motifs (Fig. S6). To statistically clarify the overlap between ABRE sequences (ACGTGG and ACGTGT) and TCP binding motifs (GGACCA and TGGTCC) in the promoter regions of genes downregulated by TCP13SRDX, we performed χ2 tests to compare the 1000 bp promoter region of TCP13SRDX-downregulated genes with that of 23,739 Arabidopsis genes previously used for motif analyses (Maruyama et al. 2012) (Fig. 7h). Significant differences were detected between the observed and expected values of the frequency of overlap between the ABRE and TCP binding motifs in the 1000 bp promoter regions of 218 TCP13SRDX-down-regulated genes. Combinations of typical ABRE sequences (ACGTGG and ACGTGT) with the TCP binding motifs (GGACCA or TGGTCC) were frequently found in the promoter regions of TCP13SRDX-downregulated genes. These results show that TCP13-target genes harbor both TCP binding and ABRE motifs in their promoter regions.
Genes downregulated in tcp13 mutant plants under dehydration stress
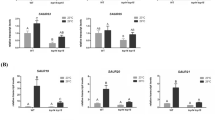
To investigate the transcriptional regulation by TCP13 under dehydration stress conditions, the expression of genes downregulated in TCP13pro::TCP13SRDX plants (determined by microarray) was examined in tcp13 mutant plants under dehydration stress by qRT-PCR (Fig. 8). First, we analyzed the expression of common downstream targets of CIN-like TCPs by qRT-PCR. IAA5 and closely related genes, IAA19, LBD1, and LOX2, showing reduced expression in tcp13 mutant plants (Fig. 8a). Compared with the WT, the expression of IAA5 and LOX2 was slightly reduced at 3 h after dehydration stress, while that of IAA19 induction was reduced at 8 h in tcp13 mutant plants. The expression of LBD1 was reduced at both 3 and 8 h after dehydration stress in tcp13 mutant plants compared with the WT. By contrast, PIN4 and FLS1 expression levels were reduced in tcp13 mutant plants compared with the WT under normal conditions.
Expression analysis of TCP13 downstream genes in tcp13 mutant plants under dehydration stress. a and b Expression of common downstream target genes of CIN-like TCPs (a), and expression of ABA-inducible genes (b) analyzed by qRT-PCR. Transcript levels of genes were normalized relative to the constitutive control, At2g32170 (Czechowski et al. 2005). In each experiment, the maximum gene transcript level was set to 100. Data represent mean ± SD (n = 3 technical replicates). An asterisk shows that the indicated mean is significantly different from the mean value of the wild-type plant under the corresponding condition (*P < 0.05, one-way ANOVA with Welch’s t test)
We found that dehydration-inducible expression of selected ABA-regulated genes was also reduced in tcp13 mutant plants compared with WT plants (Fig. 8b). The expression of RD20, NCED3, and GolS2 genes was slightly reduced at 3 h after the dehydration treatment, while that of AHG3 was reduced at 8 h after dehydration stress in tcp13 mutant plants. We also compared the expression of AHG3, RD20, Gols2, and NCED3 between ABA-treated WT and ABA-treated tcp13 mutant plants (Figure S7). These genes were upregulated in WT plants, but those upregulations were reduced in tcp13 mutant plants after the ABA treatment. Knockout of TCP13 slightly reduced the response of ABA- and dehydration-inducible genes under dehydration stress. These results suggest that TCP13 is involved in the regulation of not only common target genes of CIN-like TCPs but also ABA- and dehydration-inducible genes.
Transcriptional regulation of ABA- and auxin-regulated genes by TCP13
The results of qRT-PCR analysis showed that selected ABA-regulated genes (such as AHG3 and Gols2) and common target genes of CIN-like TCPs (such as IAA5 and LBD1) were downregulated in tcp13 mutant plants after dehydration stress (Fig. 8). The IAA/AUX proteins are auxin-sensitive repressors (Weijers and Wagner 2016). To clarify the effect of ABA- and auxin-regulated genes by TCP13, we selected AHG3 and Gols2 as the TCP-regulated ABA signaling genes and IAA5 and LBD1 as the TCP-regulated auxin signaling genes for reporter assay (Fig. 9). We investigated the activity of AHG3, Gols2, IAA5 and LBD1 gene promoter fused to the luciferase (LUC) reporter in mesophyll protoplasts of WT and tcp13 in response to the ABA and IAA treatments (Fig. 9). Figure 9a showed the existence of three DNA binding motifs of ABRE, auxin-response elements (AuxREs) (Ulmasov et al. 1995) and TCP binding motif in the promoter regions of AHG3, Gols2, IAA5 and LBD1. AHG3 and Gols2 harbor both ABRE and TCP binding motifs, whereas those of IAA5 and LBD1 contain three types of motifs including the ABRE, AuxRE, and TCP binding motif or TCP binding motif like. The AHG3 and Gols2 promoters-LUC activities increased in WT plants and those activities were reduced in the tcp13 mutant plants after the ABA treatment (Fig. 9b). However, the activities of these promoters-LUC were not affected by the IAA treatment, indicating that TCP13 is required for the activation of AHG3 and Gols2 genes during ABA signaling. IAA5 and LBD1 promoter-LUC activities increased in WT plants and those activity were reduced in tcp13 mutant plants after the IAA treatment (Fig. 9c). However, the activities of these promoters-LUC were not affected by the ABA treatment, indicating that TCP13 is required to activates IAA5 and LBD1 during auxin signaling.
Protoplast transient assays of ABA- and auxin-regulated genes in tcp13 mutant plants. a Existence of the ABREs (ACGTGT, ACGTGG, ACACGT, and CCACGT), AuxREs (TGTCTC and GAGACA), and TCP binding motifs (GGACCA, GGTCCT, TGGTCC, and AGGACC) in AHG3, Gols2, IAA5, and LBD1 promoters. The LBD1 promoter also contained the TCP binding motif like (TGGTCA). b and c Protoplast transient assays for the activation analysis of ABA-regulated genes (b) and auxin-regulated genes (c) in WT and tcp13 mutant protoplasts in response to 5 μM ABA or 1 μM IAA. In each experiment, relative luciferase (LUC) activity in the control condition was set to 100. Data represent mean ± SD (n = 3 technical replicates). An asterisk shows that the indicated mean is significantly different from the mean value of the wild-type plant under the corresponding condition (*P < 0.05, **P < 0.01, one-way ANOVA with Welch’s t test)
Discussion
A possible role of TCP13 in plant growth regulation in response to dehydration stress
Leaves often change their architecture and growth to cope with environmental fluctuations, such as limited water availability (Hsiao et al. 1984; Zhang et al. 2018). Leaf rolling is a spontaneous response of plants to dehydration stress. Water deficit causes downward rolling of leaves that prevents water loss and modulates the speed of plant growth in Arabidopsis (Fujita et al. 2018). However, factors regulating these morphological changes under water limiting conditions have not yet been elucidated. Identification of the genetic components that modulate leaf architecture and growth may lead to useful strategies to enhance crop yield under stress conditions.
In this study, we demonstrated that the CIN-like TCP TF, TCP13, plays a pivotal role in the regulation of plant growth under dehydration stress conditions. Among the CIN-like TCP gene family members, TCP13 is significantly upregulated by dehydration stress (Fig. 1). The TCP13 promoter contains two ABRE motifs. The TCP13OX transgenic Arabidopsis plants produced narrow leaves, with the downward rolling phenotype, compared with control plants, and these leaf phenotypes were drastically enhanced in response to the mannitol treatment and dehydration stress (Figs. 3, 5 and 6). Additionally, TCP13OX transgenic plants showed higher tolerance and reduced water loss under dehydration stress (Fig. 6). These results led us to speculate that TCP13 modulates the leaf growth to cope with dehydration stress conditions. The leaf growth retardation of the tcp5/13/17 triple mutants were slightly reduced in response to the mannitol treatment, whereas the tcp13 mutant plants showed no specific leaf phenotype under osmotic stress conditions (Fig. 5). These observations indicate that TCP13 probably contributes to the leaf growth together with TCP5 and TCP17 under stress conditions.
Genome-wide expression and qRT-PCR analyses revealed that IAA1, 5, 14, 17, and 34 were repressed in TCP13pro::TCP13SRDX plants (Fig. 7), and dehydration-inducible expression of IAA5 and 19 was reduced in tcp13 mutant plants under dehydration stress (Fig. 8). The IAA/AUX proteins are auxin-sensitive repressors that mediate diverse physiological and developmental processes in plants (Weijers and Wagner 2016). The incurvata6 (icu6) semi-dominant allele of the AUXIN RESISTANT3 (AXR3)/IAA17 gene increases auxin response and triggers adaxial leaf rolling because of the reduced size of adaxial pavement cells, and an abnormal expansion of palisade mesophyll cells (Perez-Perez et al. 2010). The downstream target genes of the well-characterized TCP3 include 6 of the 29 members of the IAA/AUX gene family as well as IAA3/SHY2 (Koyama et al. 2010). TCP5 and TCP17 interact with PIF4 to up-regulate the expression of IAA19 and consequently regulate thermomorphogenesis, and TCP13 also contributes to this response (Han et al. 2019; Zhou et al. 2019). Among the 29 IAA/AUX genes in Arabidopsis (Bargmann et al. 2013; Weijers and Wagner 2016), IAA5, IAA10, IAA19, and IAA31 are upregulated, whereas IAA29 is downregulated by dehydration stress (Shani et al. 2017). The IAA5, IAA6, and IAA19 genes are positively regulated by the DREB2A TF to promote stress-induced growth inhibition required for dehydration tolerance (Shani et al. 2017). Thus, we speculate that TCP13 might induces the growth inhibition through the dehydration-inducible auxin repressor, IAA5 and 19 under dehydration stress.
Our results showed that TCP13 is required for the expression of LBD1 under dehydration stress (Fig. 8) and in response to IAA treatment (Fig. 9). LBD1 is member of the ASYMMETRIC LEAVES2 (AS2)/LOB family in Arabidopsis (Iwakawa et al. 2002). AS2 forms a repressor complex with ASYMMETRIC LEAVES1 (AS1) that acts directly on class I KNOX genes (Guo et al. 2008), which promote stem cell activity and must be repressed to form determinate lateral organs (Jackson et al. 1994; Long et al. 1996). AS2 also forms a protein complex with CIN-like TCPs, including TCP2, 3, 4, 10, and 24, to repress the KNOX genes (Li et al. 2012). TCP3 transcriptionally activates AS1 expression to repress CUC genes that causes the repression of KNOX genes (Aida et al. 1999; Hibara et al. 2003; Koyama et al. 2010). Additionally, AS2 forms a trimetric complex with AS1 and another LBD protein, JAGGED LATERAL ORGANS (JLO), to coordinate auxin distribution and meristem function through the regulation of PIN and KNOX expression in shoots and roots (Rast and Simon 2012). Overexpression of the poplar (Populus tremula × Populus alba) LBD1 ortholog (PtaLBD1) enhances secondary phloem production in populus. Overexpression of PtaLBD1 downregulates the expression of KNOX genes, ARK1 and ARK2, involved in vascular cambium maintenance (Yordanov et al. 2010). The negative regulation of KNOX genes by CIN-like TCPs in a CUC-dependent and -independent manner is the core process underlying the promotion of differentiation of leaves (Koyama et al. 2010). Based on these reports, our results support the idea that TCP13 is required for the dehydration-inducible LBD1 expression, which potentially affects the leaf development in response to dehydration stress.
Although an increase in ABA level and signaling has long been recognized as an inhibitor of primary root growth (Antoni et al. 2013; Fujii et al. 2007; Gonzalez-Guzman et al. 2012; Park et al. 2009; Yoshida et al. 2010), knowledge about the underlying mechanisms is incomplete. Interestingly, tcp13 mutant plants were insensitive to the ABA treatment and showed greater root growth than WT plants (Fig. 4). Additionally, 35Spro::TCP13OX transgenic seedlings showed reduced root growth compared with control plants (Fig. 3), and this reduction was dramatically enhanced upon ABA treatment (Fig. 4). Moreover, TCP13pro::TCP13SRDX transgenic seedlings showed greater root growth than control plants under normal growth conditions (Fig. 3). Our transcriptome analysis showed that genes containing both ABRE and TCP binding motifs in their promoter regions were significantly downregulated in TCP13pro::TCP13SRDX plants (Fig. 7). The expression of ABA-regulated genes such as AHG3, RD20, NCED3, and GolS2 was reduced in tcp13 mutant plants under dehydration stress (Fig. 8) and in response to ABA treatment (Fig. 9). These results suggest that TCP13 might acts downstream of ABA signaling and participates in the inhibition of root growth under stress conditions. Several factors are involved in root growth inhibition via the ABA signaling pathway, with auxins being of prime importance. Auxin accumulation, distribution, transport, and signal transduction significantly affect primary root development (Overvoorde et al. 2010; Petricka et al. 2012; Sun et al. 2018). In Arabidopsis, ABA reduces the auxin level in roots, resulting in root growth arrest (Sun et al. 2018). ABA, dehydration, and PEG treatments prominently attenuate the response of roots to auxin in transgenic plants expressing auxin sensor constructs such as DR5::GUS, IAA2pro::GUS, and IAA19pro::Venus (Shani et al. 2017; Wang et al. 2011; Yang et al. 2014). High concentrations of ABA decrease the expression of genes encoding auxin transporters, such as AUX1 and PINs, in roots (Promchuea et al. 2017; Yang et al. 2014). ABI5 also suppresses PIN1 expression, and the abi5 mutant exhibits enhanced auxin transport in roots (Yuan et al. 2014). These results suggest that ABA inhibits root development by impacting auxin transport and signaling. Our transcriptome analysis showed that expression levels of IAA5, IAA19, and PIN4 were reduced in TCP13pro::TCP13SRDX (Fig. 7) and tcp13 mutant plants under normal growth or dehydration stress conditions (Fig. 8). Based on these reports, our results support the idea that TCP13 probably contributes the root growth inhibition under dehydration stress through negative regulation of auxin signaling.
In conclusion, the present study proposes that TCP13 might act as a positive regulator of dehydration stress tolerance through regulation of ABA and auxin signaling genes. Our results suggest that the dehydration-inducible TCP13 performs unique functions in regulating plant growth to cope with dehydration stress, and facilitates the ABA signaling. Hypothetical model of molecular framework of TCP13 under dehydration stress is presented in Figure S8. Thus, the results of this study enhance our understanding of the molecular mechanisms regulating plant growth and stress tolerance in response to dehydration stress. However, additional experiments are necessary to elucidate the detailed molecular mechanisms, especially direct regulation of ABA signaling and auxin signaling genes by TCP13. Future investigations should determine how TCP13 affect the ABA and auxin signaling genes in leaves and roots and what factors regulate the plant growth and stress tolerance through TCP13 regulation.
Data availability
Microarray design and data were deposited at ArrayExpress (https://www.ebi.ac.uk/arrayexpress/) (Accession No. E-MTAB-9336).
Change history
24 February 2022
A Correction to this paper has been published: https://doi.org/10.1007/s11103-022-01251-2
References
Aida M, Ishida T, Tasaka M (1999) Shoot apical meristem and cotyledon formation during Arabidopsis embryogenesis: interaction among the cup-shaped cotyledon and shoot meristemless genes. Development 126:1563–1570
Antoni R, Gonzalez-Guzman M, Rodriguez L, Peirats-Llobet M, Pizzio GA, Fernandez MA, De Winne N, De Jaeger G, Dietrich D, Bennett MJ, Rodriguez PL (2013) Pyrabactin Resistance1-Like8 plays an important role for the regulation of abscisic acid signaling in root. Plant Physiol 161:931–941. https://doi.org/10.1104/pp.112.208678
Bargmann BO, Vanneste S, Krouk G, Nawy T, Efroni I, Shani E, Choe G, Friml J, Bergmann DC, Estelle M, Birnbaum KD (2013) A map of cell type-specific auxin responses. Mol Syst Biol 9:688. https://doi.org/10.1038/msb.2013.40
Bechtold N, Pelletier G (1998) In planta Agrobacterium-mediated transformation of adult Arabidopsis thaliana plants by vacuum infiltration. Methods Mol Biol 82:259–266. https://doi.org/10.1385/0-89603-391-0:259
Busk PK, Pages M (1998) Regulation of abscisic acid-induced transcription. Plant Mol Biol 37:425–435
Challa KR, Rath M, Nath U (2019) The CIN-TCP transcription factors promote commitment to differentiation in Arabidopsis leaf pavement cells via both auxin-dependent and independent pathways. PLoS Genet 15:e1007988. https://doi.org/10.1371/journal.pgen.1007988
Chen W, Provart NJ, Glazebrook J, Katagiri F, Chang HS, Eulgem T, Mauch F, Luan S, Zou G, Whitham SA, Budworth PR, Tao Y, Xie Z, Chen X, Lam S, Kreps JA, Harper JF, Si-Ammour A, Mauch-Mani B, Heinlein M, Kobayashi K, Hohn T, Dangl JL, Wang X, Zhu T (2002) Expression profile matrix of Arabidopsis transcription factor genes suggests their putative functions in response to environmental stresses. Plant Cell 14:559–574
Claeys H, Inze D (2013) The agony of choice: how plants balance growth and survival under water-limiting conditions. Plant Physiol 162:1768–1779. https://doi.org/10.1104/pp.113.220921
Cubas P, Lauter N, Doebley J, Coen E (1999) The TCP domain: a motif found in proteins regulating plant growth and development. Plant J 18:215–222
Czechowski T, Stitt M, Altmann T, Udvardi MK, Scheible WR (2005) Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol 139:5–17. https://doi.org/10.1104/pp.105.063743
Ding Z, De Smet I (2013) Localised ABA signalling mediates root growth plasticity. Trends Plant Sci 18:533–535. https://doi.org/10.1016/j.tplants.2013.08.009
Dubois M, Skirycz A, Claeys H, Maleux K, Dhondt S, De Bodt S, Vanden Bossche R, De Milde L, Yoshizumi T, Matsui M, Inze D (2013) Ethylene response factor6 acts as a central regulator of leaf growth under water-limiting conditions in Arabidopsis. Plant Physiol 162:319–332. https://doi.org/10.1104/pp.113.216341
Efroni I, Han SK, Kim HJ, Wu MF, Steiner E, Birnbaum KD, Hong JC, Eshed Y, Wagner D (2013) Regulation of leaf maturation by chromatin-mediated modulation of cytokinin responses. Dev Cell 24:438–445. https://doi.org/10.1016/j.devcel.2013.01.019
Fujii H, Verslues PE, Zhu JK (2007) Identification of two protein kinases required for abscisic acid regulation of seed germination, root growth, and gene expression in Arabidopsis. Plant Cell 19:485–494. https://doi.org/10.1105/tpc.106.048538
Fujita Y, Fujita M, Satoh R, Maruyama K, Parvez MM, Seki M, Hiratsu K, Ohme-Takagi M, Shinozaki K, Yamaguchi-Shinozaki K (2005) AREB1 is a transcription activator of novel ABRE-dependent ABA signaling that enhances drought stress tolerance in Arabidopsis. Plant Cell 17:3470–3488. https://doi.org/10.1105/tpc.105.035659
Fujita M, Tanabata T, Urano K, Kikuchi S, Shinozaki K (2018) RIPPS: a plant phenotyping system for quantitative evaluation of growth under controlled environmental stress conditions. Plant Cell Physiol 59:2030–2038. https://doi.org/10.1093/pcp/pcy122
Gonzalez-Guzman M, Pizzio GA, Antoni R, Vera-Sirera F, Merilo E, Bassel GW, Fernandez MA, Holdsworth MJ, Perez-Amador MA, Kollist H, Rodriguez PL (2012) Arabidopsis PYR/PYL/RCAR receptors play a major role in quantitative regulation of stomatal aperture and transcriptional response to abscisic acid. Plant Cell 24:2483–2496. https://doi.org/10.1105/tpc.112.098574
Guo M, Thomas J, Collins G, Timmermans MC (2008) Direct repression of KNOX loci by the ASYMMETRIC LEAVES1 complex of Arabidopsis. Plant Cell 20:48–58. https://doi.org/10.1105/tpc.107.056127
Han X, Yu H, Yuan R, Yang Y, An F, Qin G (2019) Arabidopsis transcription factor TCP5 controls plant thermomorphogenesis by positively regulating PIF4 activity. iScience 15:611–622. https://doi.org/10.1016/j.isci.2019.04.005
Hibara K, Takada S, Tasaka M (2003) CUC1 gene activates the expression of SAM-related genes to induce adventitious shoot formation. Plant J 36:687–696. https://doi.org/10.1046/j.1365-313x.2003.01911.x
Hiratsu K, Matsui K, Koyama T, Ohme-Takagi M (2003) Dominant repression of target genes by chimeric repressors that include the EAR motif, a repression domain, in Arabidopsis. Plant J 34:733–739
Hsiao TC, O’Toole JC, Yambao EB, Turner NC (1984) Influence of osmotic adjustment on leaf rolling and tissue death in rice (Oryza sativa L.). Plant Physiol 75:338–341. https://doi.org/10.1104/pp.75.2.338
Iwakawa H, Ueno Y, Semiarti E, Onouchi H, Kojima S, Tsukaya H, Hasebe M, Soma T, Ikezaki M, Machida C, Machida Y (2002) The ASYMMETRIC LEAVES2 gene of Arabidopsis thaliana, required for formation of a symmetric flat leaf lamina, encodes a member of a novel family of proteins characterized by cysteine repeats and a leucine zipper. Plant Cell Physiol 43:467–478. https://doi.org/10.1093/pcp/pcf077
Jackson D, Veit B, Hake S (1994) Expression of maize KNOTTED1 related homeobox genes in the shoot apical meristem predicts patterns of morphogenesis in the vegetative shoot. Development 120:405–413. https://doi.org/10.1242/dev.120.2.405
Kodaira KS, Qin F, Tran LS, Maruyama K, Kidokoro S, Fujita Y, Shinozaki K, Yamaguchi-Shinozaki K (2011) Arabidopsis Cys2/His2 zinc-finger proteins AZF1 and AZF2 negatively regulate abscisic acid-repressive and auxin-inducible genes under abiotic stress conditions. Plant Physiol 157:742–756. https://doi.org/10.1104/pp.111.182683
Kosugi S, Ohashi Y (2002) DNA binding and dimerization specificity and potential targets for the TCP protein family. Plant J 30:337–348
Koyama T, Furutani M, Tasaka M, Ohme-Takagi M (2007) TCP transcription factors control the morphology of shoot lateral organs via negative regulation of the expression of boundary-specific genes in Arabidopsis. Plant Cell 19:473–484. https://doi.org/10.1105/tpc.106.044792
Koyama T, Mitsuda N, Seki M, Shinozaki K, Ohme-Takagi M (2010) TCP transcription factors regulate the activities of ASYMMETRIC LEAVES1 and miR164, as well as the auxin response, during differentiation of leaves in Arabidopsis. Plant Cell 22:3574–3588. https://doi.org/10.1105/tpc.110.075598
Koyama T, Sato F, Ohme-Takagi M (2017) Roles of miR319 and TCP transcription factors in leaf development. Plant Physiol 175:874–885. https://doi.org/10.1104/pp.17.00732
Kreps JA, Wu Y, Chang HS, Zhu T, Wang X, Harper JF (2002) Transcriptome changes for Arabidopsis in response to salt, osmotic, and cold stress. Plant Physiol 130:2129–2141. https://doi.org/10.1104/pp.008532
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Li S, Zachgo S (2013) TCP3 interacts with R2R3-MYB proteins, promotes flavonoid biosynthesis and negatively regulates the auxin response in Arabidopsis thaliana. Plant J 76:901–913. https://doi.org/10.1111/tpj.12348
Li Z, Li B, Shen WH, Huang H, Dong A (2012) TCP transcription factors interact with AS2 in the repression of class-I KNOX genes in Arabidopsis thaliana. Plant J 71:99–107. https://doi.org/10.1111/j.1365-313X.2012.04973.x
Long JA, Moan EI, Medford JI, Barton MK (1996) A member of the KNOTTED class of homeodomain proteins encoded by the STM gene of Arabidopsis. Nature 379:66–69. https://doi.org/10.1038/379066a0
Luo D, Carpenter R, Vincent C, Copsey L, Coen E (1996) Origin of floral asymmetry in Antirrhinum. Nature 38:794–799. https://doi.org/10.1038/383794a0
Martin-Trillo M, Cubas P (2010) TCP genes: a family snapshot ten years later. Trends Plant Sci 15:31–39. https://doi.org/10.1016/j.tplants.2009.11.003
Maruyama K, Sakuma Y, Kasuga M, Ito Y, Seki M, Goda H, Shimada Y, Yoshida S, Shinozaki K, Yamaguchi-Shinozaki K (2004) Identification of cold-inducible downstream genes of the Arabidopsis DREB1A/CBF3 transcriptional factor using two microarray systems. Plant J 38:982–993. https://doi.org/10.1111/j.1365-313X.2004.02100.x
Maruyama K, Takeda M, Kidokoro S, Yamada K, Sakuma Y, Urano K, Fujita M, Yoshiwara K, Matsukura S, Morishita Y, Sasaki R, Suzuki H, Saito K, Shibata D, Shinozaki K, Yamaguchi-Shinozaki K (2009) Metabolic pathways involved in cold acclimation identified by integrated analysis of metabolites and transcripts regulated by DREB1A and DREB2A. Plant Physiol 150:1972–1980. https://doi.org/10.1104/pp.109.135327
Maruyama K, Todaka D, Mizoi J, Yoshida T, Kidokoro S, Matsukura S, Takasaki H, Sakurai T, Yamamoto YY, Yoshiwara K, Kojima M, Sakakibara H, Shinozaki K, Yamaguchi-Shinozaki K (2012) Identification of cis-acting promoter elements in cold- and dehydration-induced transcriptional pathways in Arabidopsis, rice, and soybean. DNA Res 19:37–49. https://doi.org/10.1093/dnares/dsr040
Masuda HP, Cabral LM, De Veylder L, Tanurdzic M, de Almeida EJ, Geelen D, Inze D, Martienssen RA, Ferreira PC, Hemerly AS (2008) ABAP1 is a novel plant Armadillo BTB protein involved in DNA replication and transcription. EMBO J 27:2746–2756. https://doi.org/10.1038/emboj.2008.191
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Plant Physiol 15:473–497. https://doi.org/10.1111/j.1399-3054.1962.tb08052.x
Nath U, Crawford BC, Carpenter R, Coen E (2003) Genetic control of surface curvature. Science 299:1404–1407. https://doi.org/10.1126/science.1079354
O’Malley RC, Huang SC, Song L, Lewsey MG, Bartlett A, Nery JR, Galli M, Gallavotti A, Ecker JR (2016) Cistrome and epicistrome features shape the regulatory DNA landscape. Cell 165:1280–1292. https://doi.org/10.1016/j.cell.2016.04.038
Ori N, Cohen AR, Etzioni A, Brand A, Yanai O, Shleizer S, Menda N, Amsellem Z, Efroni I, Pekker I, Alvarez JP, Blum E, Zamir D, Eshed Y (2007) Regulation of LANCEOLATE by miR319 is required for compound-leaf development in tomato. Nat Genet 39:787–791
Overvoorde P, Fukaki H, Beeckman T (2010) Auxin control of root development. Cold Spring Harb Perspect Biol 2:a001537. https://doi.org/10.1101/cshperspect.a001537
Palatnik JF, Allen E, Wu X, Schommer C, Schwab R, Carrington JC, Weigel D (2003) Control of leaf morphogenesis by microRNAs. Nature 425:257–263. https://doi.org/10.1038/nature01958
Palatnik JF, Wollmann H, Schommer C, Schwab R, Boisbouvier J, Rodriguez R, Warthmann N, Allen E, Dezulian T, Huson D, Carrington JC, Weigel D (2007) Sequence and expression differences underlie functional specialization of Arabidopsis microRNAs miR159 and miR319. Dev Cell 13:115–125. https://doi.org/10.1016/j.devcel.2007.04.012
Park SY, Fung P, Nishimura N, Jensen DR, Fujii H, Zhao Y, Lumba S, Santiago J, Rodrigues A, Chow TF, Alfred SE, Bonetta D, Finkelstein R, Provart NJ, Desveaux D, Rodriguez PL, McCourt P, Zhu JK, Schroeder JI, Volkman BF, Cutler SR (2009) Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 324:1068–1071. https://doi.org/10.1126/science.1173041
Perez-Perez JM, Candela H, Robles P, Lopez-Torrejon G, del Pozo JC, Micol JL (2010) A role for AUXIN RESISTANT3 in the coordination of leaf growth. Plant Cell Physiol 51:1661–1673. https://doi.org/10.1093/pcp/pcq123
Petricka JJ, Winter CM, Benfey PN (2012) Control of Arabidopsis root development. Annu Rev Plant Biol 63:563–590. https://doi.org/10.1146/annurev-arplant-042811-105501
Promchuea S, Zhu Y, Chen Z, Zhang J, Gong Z (2017) ARF2 coordinates with PLETHORAs and PINs to orchestrate ABA-mediated root meristem activity in Arabidopsis. J Integr Plant Biol 59:30–43. https://doi.org/10.1111/jipb.12506
Qin F, Sakuma Y, Tran LS, Maruyama K, Kidokoro S, Fujita Y, Fujita M, Umezawa T, Sawano Y, Miyazono K, Tanokura M, Shinozaki K, Yamaguchi-Shinozaki K (2008) Arabidopsis DREB2A-interacting proteins function as RING E3 ligases and negatively regulate plant drought stress-responsive gene expression. Plant Cell 20:1693–1707. https://doi.org/10.1105/tpc.107.057380
Rast MI, Simon R (2012) Arabidopsis JAGGED LATERAL ORGANS acts with ASYMMETRIC LEAVES2 to coordinate KNOX and PIN expression in shoot and root meristems. Plant Cell 24:2917–2933. https://doi.org/10.1105/tpc.112.099978
Sakamoto S, Takata N, Oshima Y, Yoshida K, Taniguchi T, Mitsuda N (2016) Wood reinforcement of poplar by rice NAC transcription factor. Sci Rep 6:19925. https://doi.org/10.1038/srep19925
Schommer C, Palatnik JF, Aggarwal P, Chetelat A, Cubas P, Farmer EE, Nath U, Weigel D (2008) Control of jasmonate biosynthesis and senescence by miR319 targets. PLoS Biol 6:e230. https://doi.org/10.1371/journal.pbio.0060230
Seki M, Ishida J, Narusaka M, Fujita M, Nanjo T, Umezawa T, Kamiya A, Nakajima M, Enju A, Sakurai T, Satou M, Akiyama K, Yamaguchi-Shinozaki K, Carninci P, Kawai J, Hayashizaki Y, Shinozaki K (2002a) Monitoring the expression pattern of around 7,000 Arabidopsis genes under ABA treatments using a full-length cDNA microarray. Funct Integr Genomics 2:282–291. https://doi.org/10.1007/s10142-002-0070-6
Seki M, Narusaka M, Ishida J, Nanjo T, Fujita M, Oono Y, Kamiya A, Nakajima M, Enju A, Sakurai T, Satou M, Akiyama K, Taji T, Yamaguchi-Shinozaki K, Carninci P, Kawai J, Hayashizaki Y, Shinozaki K (2002b) Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J 31:279–292
Shani E, Salehin M, Zhang Y, Sanchez SE, Doherty C, Wang R, Mangado CC, Song L, Tal I, Pisanty O, Ecker JR, Kay SA, Pruneda-Paz J, Estelle M (2017) Plant stress tolerance requires auxin-sensitive Aux/IAA transcriptional repressors. Curr Biol 27:437–444. https://doi.org/10.1016/j.cub.2016.12.016
Skirycz A, Claeys H, De Bodt S, Oikawa A, Shinoda S, Andriankaja M, Maleux K, Eloy NB, Coppens F, Yoo SD, Saito K, Inze D (2011) Pause-and-stop: the effects of osmotic stress on cell proliferation during early leaf development in Arabidopsis and a role for ethylene signaling in cell cycle arrest. Plant Cell 23:1876–1888. https://doi.org/10.1105/tpc.111.084160
Song L, Huang SC, Wise A, Castanon R, Nery JR, Chen H, Watanabe M, Thomas J, Bar-Joseph Z, Ecker JR (2016) A transcription factor hierarchy defines an environmental stress response network. Science. https://doi.org/10.1126/science.aag1550
Sullivan AM, Arsovski AA, Lempe J, Bubb KL, Weirauch MT, Sabo PJ, Sandstrom R, Thurman RE, Neph S, Reynolds AP, Stergachis AB, Vernot B, Johnson AK, Haugen E, Sullivan ST, Thompson A, Neri FV 3rd, Weaver M, Diegel M, Mnaimneh S, Yang A, Hughes TR, Nemhauser JL, Queitsch C, Stamatoyannopoulos JA (2014) Mapping and dynamics of regulatory DNA and transcription factor networks in A. thaliana. Cell Rep 8:2015–2030. https://doi.org/10.1016/j.celrep.2014.08.019
Sun LR, Wang YB, He SB, Hao FS (2018) Mechanisms for abscisic acid inhibition of primary root growth. Plant Signal Behav 13:e1500069. https://doi.org/10.1080/15592324.2018.1500069
Tao Q, Guo D, Wei B, Zhang F, Pang C, Jiang H, Zhang J, Wei T, Gu H, Qu LJ, Qin G (2013) The TIE1 transcriptional repressor links TCP transcription factors with TOPLESS/TOPLESS-RELATED corepressors and modulates leaf development in Arabidopsis. Plant Cell 25:421–437. https://doi.org/10.1105/tpc.113.109223
Ulmasov T, Liu ZB, Hagen G, Guilfoyle TJ (1995) Composite structure of auxin response elements. Plant Cell 7:1611–1623. https://doi.org/10.1105/tpc.7.10.1611
Urano K, Yoshiba Y, Nanjo T, Ito T, Yamaguchi-Shinozaki K, Shinozaki K (2004) Arabidopsis stress-inducible gene for arginine decarboxylase AtADC2 is required for accumulation of putrescine in salt tolerance. Biochem Biophys Res Commun 313:369–375. https://doi.org/10.1016/j.bbrc.2003.11.119
Urano K, Maruyama K, Ogata Y, Morishita Y, Takeda M, Sakurai N, Suzuki H, Saito K, Shibata D, Kobayashi M, Yamaguchi-Shinozaki K, Shinozaki K (2009) Characterization of the ABA-regulated global responses to dehydration in Arabidopsis by metabolomics. Plant J 57:1065–1078. https://doi.org/10.1111/j.1365-313X.2008.03748.x
Urano K, Maruyama K, Jikumaru Y, Kamiya Y, Yamaguchi-Shinozaki K, Shinozaki K (2017) Analysis of plant hormone profiles in response to moderate dehydration stress. Plant J 90:17–36. https://doi.org/10.1111/tpj.13460
van Es SW, van der Auweraert EB, Silveira SR, Angenent GC, van Dijk ADJ, Immink RGH (2019) Comprehensive phenotyping reveals interactions and functions of Arabidopsis thaliana TCP genes in yield determination. Plant J 99:316–328. https://doi.org/10.1111/tpj.14326
Wang L, Hua D, He J, Duan Y, Chen Z, Hong X, Gong Z (2011) Auxin response factor2 (ARF2) and its regulated homeodomain gene HB33 mediate abscisic acid response in Arabidopsis. PLoS Genet 7:e1002172. https://doi.org/10.1371/journal.pgen.1002172
Weijers D, Wagner D (2016) Transcriptional responses to the auxin hormone. Annu Rev Plant Biol 67:539–574. https://doi.org/10.1146/annurev-arplant-043015-112122
Winter D, Vinegar B, Nahal H, Ammar R, Wilson GV, Provart NJ (2007) An “Electronic Fluorescent Pictograph” browser for exploring and analyzing large-scale biological data sets. PLoS ONE 2:e718. https://doi.org/10.1371/journal.pone.0000718
Wu FH, Shen SC, Lee LY, Lee SH, Chan MT, Lin CS (2009) Tape-Arabidopsis Sandwich—a simpler Arabidopsis protoplast isolation method. Plant Methods 5:16. https://doi.org/10.1186/1746-4811-5-16
Xu NF, Hagen G, Guilfoyle T (1997) Multiple auxin response modules in the soybean SAUR 15A promoter. Plant Sci 126:193–201. https://doi.org/10.1016/S0168-9452(97)00110-6
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803. https://doi.org/10.1146/annurev.arplant.57.032905.105444
Yang L, Zhang J, He J, Qin Y, Hua D, Duan Y, Chen Z, Gong Z (2014) ABA-mediated ROS in mitochondria regulate root meristem activity by controlling PLETHORA expression in Arabidopsis. PLoS Genet 10:e1004791. https://doi.org/10.1371/journal.pgen.1004791
Yoo SD, Cho YH, Sheen J (2007) Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat Protoc 2:1565–1572. https://doi.org/10.1038/nprot.2007.199
Yordanov YS, Regan S, Busov V (2010) Members of the lateral organ boundaries domain transcription factor family are involved in the regulation of secondary growth in Populus. Plant Cell 22:3662–3677. https://doi.org/10.1105/tpc.110.078634
Yoshida T, Fujita Y, Sayama H, Kidokoro S, Maruyama K, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2010) AREB1, AREB2, and ABF3 are master transcription factors that cooperatively regulate ABRE-dependent ABA signaling involved in drought stress tolerance and require ABA for full activation. Plant J 61:672–685. https://doi.org/10.1111/j.1365-313X.2009.04092.x
Yoshida K, Sakamoto S, Kawai T, Kobayashi Y, Sato K, Ichinose Y, Yaoi K, Akiyoshi-Endo M, Sato H, Takamizo T, Ohme-Takagi M, Mitsuda N (2013) Engineering the Oryza sativa cell wall with rice NAC transcription factors regulating secondary wall formation. Front Plant Sci 4:383. https://doi.org/10.3389/fpls.2013.00383
Yu H, Zhang L, Wang W, Tian P, Wang W, Wang K, Gao Z, Liu S, Zhang Y, Irish VF, Huang T (2020) TCP5 controls leaf margin development by regulating KNOX and BEL-like transcription factors in Arabidopsis. J Exp Bot 72:1809–1821. https://doi.org/10.1093/jxb/eraa569
Yuan TT, Xu HH, Zhang KX, Guo TT, Lu YT (2014) Glucose inhibits root meristem growth via ABA INSENSITIVE 5, which represses PIN1 accumulation and auxin activity in Arabidopsis. Plant Cell Environ 37:1338–1350. https://doi.org/10.1111/pce.12233
Zhang J, Wei B, Yuan R, Wang J, Ding M, Chen Z, Yu H, Qin G (2017) The Arabidopsis RING-type E3 ligase TEAR1 controls leaf development by targeting the TIE1 transcriptional repressor for degradation. Plant Cell 29:243–259. https://doi.org/10.1105/tpc.16.00771
Zhang J, Zhang H, Srivastava AK, Pan Y, Bai J, Fang J, Shi H, Zhu JK (2018) Knockdown of rice microRNA166 confers drought resistance by causing leaf rolling and altering stem xylem development. Plant Physiol 176:2082–2094. https://doi.org/10.1104/pp.17.01432
Zhang H, Zhang F, Yu Y, Feng L, Jia J, Liu B, Li B, Guo H, Zhai J (2020) A comprehensive online database for exploring approximately 20,000 public Arabidopsis RNA-seq libraries. Mol Plant 13:1231–1233. https://doi.org/10.1016/j.molp.2020.08.001
Zhou Y, Xun Q, Zhang D, Lv M, Ou Y, Li J (2019) TCP transcription factors associate with phytochrome interacting factor 4 and cryptochrome 1 to regulate thermomorphogenesis in Arabidopsis thaliana. iScience 15:600–610. https://doi.org/10.1016/j.isci.2019.04.002
Acknowledgements
We thank Kyoko Yoshiwara (JIRCAS) for microarray analysis, and Dr. Yasunari Fujita (JIRCAS) for providing the DNA of AREB1, active form of AREB1 (AREB1ΔQT), and RD29B promoter for transactivation assays. We also thank Saho Mizukado, Hiroko Kobayashi, Kumiko Matsuo, Michie Etoh, and Dr. Fuyuko Shimoda for technical assistance.
Funding
This research was supported by the Program for Promotion of Basic and Applied Researches for Innovations in Bio-oriented Industry (BRAIN; to K.S.); the Ministry of Agriculture, Forestry, and Fisheries (MAFF); and the Ministry of Education, Culture, Sports, Science and Technology (MEXT)/JSPS KAKENHI (20K05833 to K.U.).
Author information
Authors and Affiliations
Contributions
All the authors contributed to the design of the study conception. Materials were prepared and experiments were performed by KU. The first draft of the manuscript was written by KU and KS, and all the authors commented on previous versions of the manuscript. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Accession Numbers
Microarray design and data were deposited at ArrayExpress (Accession No. E-MTAB-9336).
The original online version of this article was revised due to a retrospective Open Access order.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Urano, K., Maruyama, K., Koyama, T. et al. CIN-like TCP13 is essential for plant growth regulation under dehydration stress. Plant Mol Biol 108, 257–275 (2022). https://doi.org/10.1007/s11103-021-01238-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-021-01238-5