Abstract
Purpose
To explain the differences in protein-protein interactions (PPI) of concentrated versus dilute formulations of a model antibody.
Methods
High frequency rheological measurements from pH 3.0 to 12.0 quantitated viscoelasticity and PPI at high concentrations. Dynamic light scattering (DLS) characterized PPI in dilute solutions.
Results
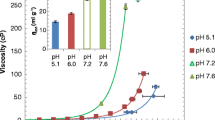
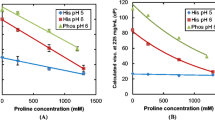
For concentrated solutions at low ionic strength, the storage modulus, a viscosity component and a measure of PPI, is highest at the isoelectric point (pH 9.0) and lowest at pH 5.4. This profile flattens at higher ionic strength but not completely, indicating PPI consist of long-range electrostatics and other short-range attractions. At low concentrations, PPI are near zero at pI but become repulsive as the pH is shifted. Higher salt concentrations completely flatten this profile to zero, indicating that these PPI are mainly electrostatic.
Conclusions
This discrepancy occurs because long-range interactions are significant at low concentrations, whereas both long- and short-range interactions are significant at higher concentrations. Computer modeling was used to calculate antibody properties responsible for long- and short-range interactions, i.e. net charge and dipole moment. Charge-charge interactions are repulsive while dipole-dipole interactions are attractive. Their net effect correlated with the storage modulus profile. However, only charge-charge repulsions correlated with PPI determined by DLS.







Similar content being viewed by others
Abbreviations
- B 22 :
-
second osmotic virial coefficient
- B′ 22 :
-
second osmotic virial coefficient multiplied by solute molecular weight
- CD:
-
circular dichroism
- DLS:
-
dynamic light scattering
- E B22 :
-
pairwise energetic interaction term
- G′ :
-
storage modulus of complex viscosity
- G″ :
-
loss modulus of complex viscosity
- IgG:
-
immunoglobulin G
- k D :
-
interaction parameter from DLS
- mAb:
-
monoclonal antibody
- |η*|:
-
magnitude of complex viscosity
- PDB:
-
Protein Databank
- PPI:
-
protein-protein interaction(s)
- V B22 :
-
excluded volume term
References
Fulton AB. How crowded is the cytoplasm? Cell. 1982;30:345–7.
Zimmerman SB, Minton AP. Macromolecular crowding: biochemical, biophysical, and physiological consequences. Annu Rev Biophys Biomol Struct. 1993;22:7–65.
Minton AP. Influence of macromolecular crowding upon the stability and state of association of proteins: Predictions and observations. J Pharm Sci. 2005;94:1668–75.
Wang W, Singh S, Zeng DL, King K, Nema S. Antibody structure, instability, and formulation. J Pharm Sci. 2006;96:1–26.
Shire SJ, Shahrokh Z, Liu J. Challenges in the development of high protein concentration formulations. J Pharm Sci. 2004;2004:1390–402.
Liu J, Nguyen MD, Andya JD, Shire SJ. Reversible self-association increases the viscosity of a concentrated monoclonal antibody in aqueous solution. J Pharm Sci. 2005;94:1928–40.
Saluja A, Kalonia DS. Nature and consequences of protein-protein interactions in high protein concentration solutions. Int J Pharm. 2008;358:1–15.
Zhang J, Liu XY. Effect of protein–protein interactions on protein aggregation kinetics. J Chem Phys. 2003;119:10972–6.
Kanai S, Liu J, Patapoff TW, Shire SJ. Reversible self-association of a concentrated monoclonal antibody solution mediated by Fab-Fab interaction that impacts solution viscosity. J Pharm Sci. 2008;97:4219–27.
Meehan S, Berry Y, Luisi B, Dobson CM, Carver JA, MacPhee CE. Amyloid fibril formation by lens crystallin proteins and its implications for cataract formation. J Biol Chem. 2004;279:3413–9.
Harper JD, Lansbury PT. Models of amyloid seeding in Alzheimer’s disease and scrapie: mechanistic truths and physiological consequences of the time-dependent solubility of amyloid proteins. Annu Rev Biochem. 1997;66:385–407.
Koo EH, Lansbury PT, Kelly JW. Amyloid diseases: abnormal protein aggregation in neurodegeneration. Proc Natl Acad Sci USA. 1999;96:9989–90.
Ganeval D, Noël LH, Preud’homme JL, Droz D, Grünfeld JP. Light-chain deposition disease: its relation with AL-type amyloidosis. Kidney Int. 1984;26:1–9.
Buxbaum JN, Chuba JV, Hellman GC, Solomon A, Gallo GR. Monoclonal immunoglobulin deposition disease: light chain and light and heavy chain deposition diseases and their relation to light chain amyloidosis. Clinical features, immunopathology, and molecular analysis. Ann Intern Med. 1990;112:455–64.
Zimm BH. Applications of the methods of molecular distribution to solutions of large molecules. J Chem Phys. 1946;14:164–79.
Teraoka I. Polymer solutions: an introduction to physical properties. New Jersey, USA: Wiley-IEEE; 2002.
George A, Chiang Y, Guo B, Arabshahi A, Cai Z, Wilson WW. Second virial coefficient as predictor in protein crystal growth. Methods Enzymol. 1997;276:100–10.
Saluja A, Kalonia DS. Measurement of fluid viscosity at microliter volumes using quartz impedance analysis. AAPS PharmSciTech. 2004;5:e47.
Saluja A, Kalonia DS. Application of ultrasonic shear rheometer to characterize rheological properties of high protein concentration solutions at microliter volume. J Pharm Sci. 2005;94:1161–8.
Saluja A, Badkar AV, Zeng DL, Nema S, Kalonia DS. Application of high-frequency rheology measurements for analyzing protein-protein interactions in high protein concentration solutions using a model monoclonal antibody (IgG2). J Pharm Sci. 2006;95:1967–83.
Hackley VA, Ferraris CF. Guide to rheological nomenclature for liquid-based particle systems. Maryland, USA: NIST; 2001.
Saluja A, Badkar AV, Zeng DL, Nema S, Kalonia DS. Ultrasonic storage modulus as a novel parameter for analyzing protein-protein interactions in high protein concentration solutions: correlation with static and dynamic light scattering measurements. Biophys J. 2007;92:234–44.
Hiemenz PC, Rajagopalan R. Principles of colloid and surface chemistry. New York, USA: Marcel Dekker; 1997.
Curtis RA, Prausnitz JM, Blanch HW. Protein-protein and protein-salt interactions in aqueous protein solutions containing concentrated electrolytes. Biotechnol Bioeng. 1998;57:11–21.
Elcock AH, McCammon JA. Calculation of weak protein-protein interactions: the pH dependence of the second virial coefficient. Biophys J. 2001;80:613–25.
Torshin IY. Bioinformatics in the post-genomic era: the role of biophysics. New York, USA: Nova Science; 2006.
Al-Shakhshira RH, Regnierb FE, Whitec JL, Hema SL. Contribution of electrostatic and hydrophobic interactions to the adsorption of proteins by aluminium-containing adjuvants. Vaccine. 1995;13:41–4.
Harris LJ, Larson SB, Hasel KW, McPherson A. Refined structure of an intact IgG2a monoclonal antibody. Biochem. 1997;36:1581–97.
RCSB Protein Data Bank. http://www.pdb.org (accessed December 2008).
Hyperchem Professional 7.5.1 (Hypercube, Inc., Gainesville, Florida, USA).
Discovery Studio 2.1 (Accelrys, Inc., San Diego, California, USA).
Nezlin R. The immunoglobulins: structure and function. London, UK: Academic; 1998.
Li H, Robertson AD, Jensen JH. Very fast empirical prediction and interpretation of protein pKa values. Proteins. 2005;61:704–21.
Bas DC, Rogers DM, Jensen JH. Very fast prediction and rationalization of pKa values for protein-ligand complexes. Proteins. 2008;73:765–83.
Brooks BR, Bruccoleri RE, Olafson BD, States DJ, Swaminathan S, Karplus M. CHARMM: a program for macromolecular energy, minimization, and dynamics calculations. J Comp Chem. 1983;4:187–17.
Gilson MK, Gilson HSR, Potter MJ. Fast assignment of accurate partial atomic charges. An electronegativity equalization method that accounts for alternate resonance forms. J Chem Inf Comput Sci. 2003;43:1982–97.
Chen W, Huang J, Gilson MK. Identification of symmetries in molecules and complexes. J Chem Inf Comput Sci. 2004;44:1301–13.
Felder CE, Prilusky J, Silman I, Sussman JL. A server and database for dipole moments of proteins. Nucleic Acids Res. 2007;35:W512–21.
Baker NA, Sept D, Joseph S, Holst MJ, McCammon JA. Electrostatics of nanosystems: Application to microtubules and the ribosome. Proc Natl Acad Sci USA. 2001;98:10037–41.
Dolinsky TJ, Nielsen JE, McCammon JA, Baker NA. PDB2PQR: an automated pipeline for the setup, execution, and analysis of Poisson-Boltzmann electrostatics calculations. Nuc Acids Res. 2004;32:W665–7.
Humphrey W, Dalke A, Schulten K. VMD—visual molecular dynamics. J Molec Graphics. 1996;14:33–8.
JAVA 1.4.2 programming language (Sun Microsystems, Inc., Santa Clara, California, USA).
Shaw KL, Grimsley GR, Yakovlev GI, Makarov AA, Pace CN. The effect of net charge on the solubility, activity, and stability of ribonuclease Sa. Protein Sci. 2001;10:1206–15.
Neal BL, Asthagiri D, Lenhoff AM. Molecular origins of osmotic second virial coefficients of proteins. Biophys J. 1998;75:2469–77.
Papp E, Fricsovszky G, Meszéna G. Electrodichroism of purple membrane. Biophys J. 1986;49:1089–100.
Acknowledgements
The authors thank Pfizer Inc. for donating the mAb for this study and for partial financial support of this work.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Materials
Below is the link to the electronic supplementary material.
ESM 1
(DOC 269 kb)
Rights and permissions
About this article
Cite this article
Chari, R., Jerath, K., Badkar, A.V. et al. Long- and Short-Range Electrostatic Interactions Affect the Rheology of Highly Concentrated Antibody Solutions. Pharm Res 26, 2607–2618 (2009). https://doi.org/10.1007/s11095-009-9975-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11095-009-9975-2