Abstract
Genetic improvement of sesame (Sesamum indicum L.), one of the most important oilseed crops providing edible oil, proteins, minerals, and vitamins, is important to ensure a balanced diet for the growing world population. Increasing yield, seed protein, oil, minerals, and vitamins is urgently needed to meet the global demand. The production and productivity of sesame is very low due to various biotic and abiotic stresses. Therefore, various efforts have been made to combat these constraints and increase the production and productivity of sesame through conventional breeding. However, less attention has been paid to the genetic improvement of the crop through modern biotechnological methods, leaving it lagging behind other oilseed crops. Recently, however, the scenario has changed as sesame research has entered the era of “omics” and has made significant progress. Therefore, the purpose of this paper is to provide an overview of the progress made by omics research in improving sesame. This review presents a number of efforts that have been made over past decade using omics technologies to improve various traits of sesame, including seed composition, yield, and biotic and abiotic resistant varieties. It summarizes the advances in genetic improvement of sesame using omics technologies, such as germplasm development (web-based functional databases and germplasm resources), gene discovery (molecular markers and genetic linkage map construction), proteomics, transcriptomics, and metabolomics that have been carried out in the last decade. In conclusion, this review highlights future directions that may be important for omics-assisted breeding in sesame genetic improvement.
Similar content being viewed by others
Data Availability
Not applicable.
References
López-Bellido RJ, López-Bellido L, Castillo JE, López-Bellido FJ (2002) Sunflower response to tillage and soil residual nitrogen in a wheat–sunflower rotation under rainfed Mediterranean conditions. Aust J Agric Res 53(9):1027–1033
Pathak N, Rai A, Kumari R, Bhat K (2014) Value addition in sesame: a perspective on bioactive components for enhancing utility and profitability. Pharmacogn Rev 8(16):147
Anilakumar KR, Pal A, Khanum F, Bawa AS (2010) Nutritional, medicinal and industrial uses of sesame (Sesamum indicum L.) seeds-an overview. Agriculturae Conspectus Scientificus 75(4):159–168
Wang L, Zhang Y, Li P, Wang X, Zhang W, Wei W et al (2012) HPLC analysis of seed sesamin and sesamolin variation in a sesame germplasm collection in China. J Am Oil Chem Soc 89(6):1011–1020
FAOSTAT. FAOSTAT Provides Free Access to Food and Agriculture Data for Over 245 Countries and Territories and Covers All FAO Regional Groupings. Available at: http://faostat.fao.org/ [accessed May 19, 2020].
Dossa K, Diouf D, Wang L, Wei X, Zhang Y, Niang M et al (2017) The emerging oilseed crop Sesamum indicum enters the “Omics” era. Front Plant Sci 8:1154
Singh M, Chahar S, Avtar R, Singh A, Kumar N (2022) Advances in classical and molecular breeding in Sesame (Sesamum indicum L.). Accelerated plant breeding, vol 4. Springer, pp 491–529
Wang L, Zhang Y, Zhu X, Zhu X, Li D, Zhang X et al (2017) Development of an SSR-based genetic map in sesame and identification of quantitative trait loci associated with charcoal rot resistance. Sci Rep 7(1):1–8
Zhang Y-, Hu F, Zhang J-G, Wei P-C, Wei Z-J (2020) Integrated small RNA and degradome sequencing provide insights into salt tolerance in sesame (Sesamum indicum L). BMC Genomics 21:494
Dossa K, Niang M, Assogbadjo AE, Cissé N, Diouf D (2016) Whole genome homology-based identification of candidate genes for drought tolerance in sesame (Sesamum indicum L). Afr J Biotechnol 15(27):1464–1475
Dossa K, Mmadi MA, Zhou R, Zhang T, Su R, Zhang Y et al (2019) Depicting the core transcriptome modulating multiple abiotic stresses responses in sesame (Sesamum indicum L). Int J Mol Sci 20(16):3930
Li D, Dossa K, Zhang Y, Wei X, Wang L, Zhang Y et al (2018) GWAS uncovers differential genetic bases for drought and salt tolerances in sesame at the germination stage. Genes 9(2):87
Zhang H, Miao H, Wang L, Qu L, Liu H, Wang Q et al (2013) Genome sequencing of the important oilseed crop Sesamum indicumL. Genome Biol 14(1):401
Dossa K, Wei X, Li D, Fonceka D, Zhang Y, Wang L et al (2016) Insight into the AP2/ERF transcription factor superfamily in sesame and expression profiling of DREB subfamily under drought stress. BMC Plant Biol 16(1):171
Dossa K, Diouf D, Cissé N (2016) Genome-wide investigation of hsf genes in sesame reveals their segmental duplication expansion and their active role in drought stress response. Front Plant Sci 7:1522
Li D, Liu P, Yu J, Wang L, Dossa K, Zhang Y et al (2017) Genome-wide analysis of WRKY gene family in the sesame genome and identification of the WRKY genes involved in responses to abiotic stresses. BMC Plant Biol 17(1):152
Mmadi MA, Dossa K, Wang L, Zhou R, Wang Y, Cisse N et al (2017) Functional characterization of the versatile MYB gene family uncovered their important roles in plant development and responses to drought and waterlogging in sesame. Genes 8(12):362
Wei M, Liu A, Zhang Y, Zhou Y, Li D, Dossa K et al (2019) Genome-wide characterization and expression analysis of the HD-Zip gene family in response to drought and salinity stresses in sesame. BMC Genomics 20(1):748
Wang Y, Zhang Y, Zhou R, Dossa K, Yu J, Li D et al (2018) Identification and characterization of the bZIP transcription factor family and its expression in response to abiotic stresses in sesame. PLoS One. ;13(7)
Zhang Y, Li D, Wang Y, Zhou R, Wang L, Zhang Y et al (2018) Genome-wide identification and comprehensive analysis of the NAC transcription factor family in Sesamum indicum. PLoS ONE. ;13(6)
Zhang H, Miao H, Ju M (2019) Potential for adaptation to climate change through genomic breeding in sesame. Genomic designing of climate-smart oilseed crops. Springer, pp 371–440
Wei PZF, Wang Z, Wang Q, Chai X, Hou G, Meng Q (2022) Sesame (Sesamum indicum L.): a Comprehensive Review of Nutritional Value, Phytochemical Composition, Health benefits, development of Food, and Industrial Applications. Nutrients 14:4079
Yadav RKS, Rangan P, Pradheep K, Rao GP, Kaur V, Pandey R, Rai V, Vasimalla CC, Langyan S, Sharma S, Thangavel B, Rana VS, Vishwakarma H, Shah A, Saxena A, Kumar A (2022) Singh K and Siddique KHM. Current research Trends and prospects for yield and quality improvement in Sesame, an important oilseed crop. Front Plant Sci 13:863521
Teklu DHSH, Abady S (2022) Genetic improvement in Sesame (Sesamum indicum L.): Progress and Outlook: a review. Agronomy 12:2144
Singh V, Kumar S, Singh A, Bhaduri NP, Bhat KV, Lakhanpaul S (2016) Unlocking the potential of genetic resources for improvement of sesame (Sesamum indicum L.): the current scenario. Springer, Gene Pool Diversity and Crop Improvement, pp 447–479
KOBAYASHI T (1991) Cytogenetics of sesame (Sesamum indicum). Developments in Plant Genetics and breeding, vol 2. Elsevier, pp 581–592
Ekta S, Shah TI, Fatima K (2014) A review enlightening genetic divergence in Sesamum indicum based on morphological and molecular studies. Int J Agric Crop Sci (IJACS) 7(1):1–9
Bedigian D (2003) Sesame in Africa: origin and dispersals. Food, fuel and Fields—Progress in African Archaeobotany, Afr Prae-Historica. :17–36
Bedigian D (2004) History and lore of sesame in Southwest Asia. Econ Bot 58(3):329–353
Zeven AC, Zhukovsky PM (1975) Dictionary of cultivated plants and their centres of diversity: excluding ornamentals, forest trees and lower plants. Pudoc
Zhang H, Wei L, Miao H, Zhang T, Wang C (2012) Development and validation of genic-SSR markers in sesame by RNA-seq. BMC Genomics 13(1):316
Morris J (2009) Characterization of sesame (Sesamum indicum L.) germplasm regenerated in Georgia, USA. Genet Resour Crop Evol 56(7):925
Bisht I, Mahajan R, Loknathan T, Agrawal R (1998) Diversity in indian sesame collection and stratification of germplasm accessions in different diversity groups. Genet Resour Crop Evol 45(4):325–335
Park J-H, Suresh S, Raveendar S, Baek H-J, Kim C-K, Lee S et al (2015) Development and evaluation of core collection using qualitative and quantitative trait descriptor in sesame (Sesamum indicum L.) germplasm. Korean J crop Sci 60(1):75–84
IPGRI N (2004) Descriptors for sesame. Sesamum
Hodgkin T, Brown AH, Van Hintum TJ, Morales EAV (1995) Core collections of plant genetic resources. John Wiley & Sons
Zhang Y, Zhang X, Che Z, Wang L, Wei W, Li D (2012) Genetic diversity assessment of sesame core collection in China by phenotype and molecular markers and extraction of a mini-core collection. BMC Genet 13(1):102
Dossa K, Wei X, Zhang Y, Fonceka D, Yang W, Diouf D et al (2016) Analysis of genetic diversity and population structure of sesame accessions from Africa and Asia as major centers of its cultivation. Genes 7(4):14
Basak M, Uzun B, Yol E (2019) Genetic diversity and population structure of the Mediterranean sesame core collection with use of genome-wide SNPs developed by double digest RAD-Seq. PLoS ONE 14(10):e0223757
Uncu AO, Frary A, Karlovsky P, Doganlar S (2016) High-throughput single nucleotide polymorphism (SNP) identification and mapping in the sesame (Sesamum indicum L.) genome with genotyping by sequencing (GBS) analysis. Mol Breeding 36(12):173
Zhang H, Miao H, Li C, Wei L, Duan Y, Ma Q et al (2016) Ultra-dense SNP genetic map construction and identification of SiDt gene controlling the determinate growth habit in Sesamum indicum L. Sci Rep 6(1):1–13
Zhang Y, Wang L, Xin H, Li D, Ma C, Ding X et al (2013) Construction of a high-density genetic map for sesame based on large scale marker development by specific length amplified fragment (SLAF) sequencing. BMC Plant Biol 13(1):141
Wei L, Miao H, Li C, Duan Y, Niu J, Zhang T et al (2014) Development of SNP and InDel markers via de novo transcriptome assembly in Sesamum indicum L. Mol Breeding 34(4):2205–2217
Wu K, Liu H, Yang M, Tao Y, Ma H, Wu W et al (2014) High-density genetic map construction and QTLs analysis of grain yield-related traits in Sesame (Sesamum indicum L.) based on RAD-Seq techonology. BMC Plant Biol 14(1):274
Wang L, Xia Q, Zhang Y, Zhu X, Zhu X, Li D et al (2016) Updated sesame genome assembly and fine mapping of plant height and seed coat color QTLs using a new high-density genetic map. BMC Genomics 17(1):31
Wei X, Wang L, Zhang Y, Qi X, Wang X, Ding X et al (2014) Development of simple sequence repeat (SSR) markers of sesame (Sesamum indicum) from a genome survey. Molecules 19(4):5150–5162
Wang L, Zhang Y, Zhu X, Zhu X, Li D, Zhang X et al (2017) Development of an SSR-based genetic map in sesame and identification of quantitative trait loci associated with charcoal rot resistance. Sci Rep 7(1):8349
El Harfi M, Charafi J, Houmanat K, Hanine H, Nabloussi A (2021) Assessment of genetic diversity in moroccan sesame (Sesamum indicum) using ISSR molecular markers. OCL 28:3
Sehr EM, Okello-Anyanga W, Hasel-Hohl K, Burg A, Gaubitzer S, Rubaihayo PR et al (2016) Assessment of genetic diversity amongst Ugandan sesame (Sesamum indicum L.) landraces based on agromorphological traits and genetic markers. J Crop Sci Biotechnol 19(1):117–124
Yi D-K, Kim K-J (2012) Complete chloroplast genome sequences of important oilseed crop Sesamum indicum L. PLoS ONE. ;7(5)
Dossa K (2016) A physical map of important QTLs, functional markers and genes available for sesame breeding programs. Physiol Mol Biology Plants 22(4):613–619
Yu J, Dossa K, Wang L, Zhang Y, Wei X, Liao B et al (2017) PMDBase: a database for studying microsatellite DNA and marker development in plants. Nucleic Acids Res 45(D1):D1046–D53
Wang L, Zhang Y, Qi X, Gao Y, Zhang X (2012) Development and characterization of 59 polymorphic cDNA-SSR markers for the edible oil crop Sesamum indicum (Pedaliaceae). Am J Bot 99(10):e394–e8
Stavridou E, Lagiotis G, Kalaitzidou P, Grigoriadis I, Bosmali I, Tsaliki E et al (2021) Characterization of the genetic diversity present in a diverse sesame landrace collection based on phenotypic traits and EST-SSR markers coupled with an HRM analysis. Plants 10(4):656
Adéoti K, Rival A, Dansi A, Santoni S, Brown S, Beule T et al (2011) Genetic characterization of two traditional leafy vegetables (Sesamum radiatum Thonn. Ex Hornem and Ceratotheca sesamoides Endl.) Of Benin, using flow cytometry and amplified fragment length polymorphism (AFLP) markers. Afr J Biotechnol 10(65):14264–14275
Dar AA, Mudigunda S, Mittal PK, Arumugam N (2017) Comparative assessment of genetic diversity in Sesamum indicum L. using RAPD and SSR markers. 3 Biotech 7:1–12
Suh MC, Kim MJ, Hur C-G, Bae JM, Park YI, Chung C-H et al (2003) Comparative analysis of expressed sequence tags from Sesamum indicum and Arabidopsis thaliana developing seeds. Plant Mol Biol 52(6):1107–1123
Wei W, Qi X, Wang L, Zhang Y, Hua W, Li D et al (2011) Characterization of the sesame (Sesamum indicum L.) global transcriptome using Illumina paired-end sequencing and development of EST-SSR markers. BMC Genomics 12(1):451
Wu K, Yang M, Liu H, Tao Y, Mei J, Zhao Y (2014) Genetic analysis and molecular characterization of chinese sesame (Sesamum indicum L.) cultivars using insertion-deletion (InDel) and simple sequence repeat (SSR) markers. BMC Genet 15(1):35
Wei X, Zhu X, Yu J, Wang L, Zhang Y, Li D et al (2016) Identification of sesame genomic variations from genome comparison of landrace and variety. Front Plant Sci 7:1169
Du H, Zhang H, Wei L, Li C, Duan Y, Wang H (2019) A high-density genetic map constructed using specific length amplified fragment (SLAF) sequencing and QTL mapping of seed-related traits in sesame (Sesamum indicum L). BMC Plant Biol 19(1):588
Yu J, Golicz AA, Lu K, Dossa K, Zhang Y, Chen J et al (2019) Insight into the evolution and functional characteristics of the pan-genome assembly from sesame landraces and modern cultivars. Plant Biotechnol J 17(5):881–892
Verma P, Goyal R, Chahota R, Sharma TR, Abdin M, Bhatia S (2015) Construction of a genetic linkage map and identification of QTLs for seed weight and seed size traits in lentil (Lens culinaris Medik). PLoS ONE 10(10):e0139666
Wei L-B, Zhang H-Y, Zheng Y-Z, Miao H-M, Zhang T-Z, Guo W-Z (2009) A genetic linkage map construction for sesame (Sesamum indicum L). Genes & genomics 31(2):199–208
Zhang H, Miao H, Wei L, Li C, Zhao R, Wang C (2013) Genetic analysis and QTL mapping of seed coat color in sesame (Sesamum indicum L.). PLoS ONE. ;8(5)
Cui C, Mei H, Liu Y, Zhang H, Zheng Y (2017) Genetic diversity, population structure, and linkage disequilibrium of an association-mapping panel revealed by genome-wide SNP markers in sesame. Front Plant Sci 8:1189
Zhang Y, Wang L, Xin H, Li D, Ma C, Ding X et al (2013) Construction of a high-density genetic map for sesame based on large scale marker development by specific length amplified fragment (SLAF) sequencing. BMC Plant Biol 13(1):1–12
Zhang H, Miao H, Li C, Wei L, Duan Y, Ma Q et al (2016) Ultra-dense SNP genetic map construction and identification of SiDt gene controlling the determinate growth habit in Sesamum indicum L. Sci Rep 6:31556
Ferreira A, Silva MFd, Cruz CD (2006) Estimating the effects of population size and type on the accuracy of genetic maps. Genet Mol Biology 29(1):187–192
Mei H, Liu Y, Du Z, Wu K, Cui C, Jiang X et al (2017) High-density genetic map construction and gene mapping of basal branching habit and flowers per leaf axil in sesame. Front Plant Sci 8:636
Asekova S, Oh E, Kulkarni KP, Lee MH, Kim JI, Pae S-B et al (2020) A Combinatorial Approach of Biparental QTL Mapping and Genome-Wide Association Analysis Identifies Candidate Genes for Phytophthora Blight Resistance in Sesame. bioRxiv.
Berhe M, Dossa K, You J, Mboup PA, Diallo IN, Diouf D et al (2021) Genome-wide association study and its applications in the non-model crop Sesamum indicum. BMC Plant Biol 21(1):283
Wei X, Liu K, Zhang Y, Feng Q, Wang L, Zhao Y et al (2015) Genetic discovery for oil production and quality in sesame. Nat Commun 6(1):8609
Zhou R, Dossa K, Li D, Yu J, You J, Wei X et al (2018) Genome-wide association studies of 39 seed yield-related traits in sesame (Sesamum indicum L). Int J Mol Sci 19(9):2794
He Q, Xu F, Min M-H, Chu S-H, Kim K-W, Park Y-J (2019) Genome-wide association study of vitamin E using genotyping by sequencing in sesame (Sesamum indicum). Genes & genomics 41:1085–1093
Dossa K, Li D, Zhou R, Yu J, Wang L, Zhang Y et al (2019) The genetic basis of drought tolerance in the high oil crop Sesamum indicum. Plant Biotechnol J 17(9):1788–1803
Cui C, Liu Y, Liu Y, Cui X, Sun Z, Du Z et al (2021) Genome-wide association study of seed coat color in sesame (Sesamum indicum L). PLoS ONE 16(5):e0251526
Dossa K, Zhou R, Li D, Liu A, Qin L, Mmadi MA et al (2021) A novel motif in the 5’-UTR of an orphan gene ‘Big Root Biomass’ modulates root biomass in sesame. Plant Biotechnol J 19(5):1065–1079
Tesfaye T, Tesfaye K, Keneni G, Alemu T, Alemu A (2022) Genome-wide association study for yield‐related traits in sesame (Sesamum Indicum). Plant Breeding 141(2):246–256
Dossa K, Yu J, Liao B, Cisse N, Zhang X (2017) Development of highly informative genome-wide single sequence repeat markers for breeding applications in sesame and construction of a web resource: SisatBase. Front Plant Sci 8:1470
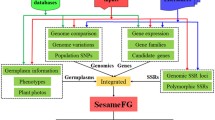
Wei X, Gong H, Yu J, Liu P, Wang L, Zhang Y et al (2017) SesameFG: an integrated database for the functional genomics of sesame. Sci Rep 7(1):1–10
Ke T, Yu J, Dong C, Mao H, Hua W, Liu S (2015) ocsESTdb: a database of oil crop seed EST sequences for comparative analysis and investigation of a global metabolic network and oil accumulation metabolism. BMC Plant Biol 15(1):19
Wei X, Liu K, Zhang Y, Feng Q, Wang L, Zhao Y et al (2015) Genetic discovery for oil production and quality in sesame. Nat Commun 6:8609
Wang L, Yu J, Li D, Zhang X (2015) Sinbase: an integrated database to study genomics, genetics and comparative genomics in Sesamum indicum. Plant Cell Physiol 56(1):e2–e
Wang L, Yu J, Zhang Y, You J, Zhang X, Wang L (2021) Sinbase 2.0: an updated database to study multi-omics in Sesamum indicum. Plants 10(2):272
Mei H, Liu Y, Cui C, Hu C, Xie F, Zheng L et al (2021) QTL mapping of yield-related traits in sesame. Mol Breeding 41(7):43
Xu F, Zhou R, Dossou SSK, Song S, Wang L (2021) Fine mapping of a major pleiotropic QTL associated with sesamin and sesamolin variation in sesame (Sesamum indicum L). Plants 10(7):1343
Yol E, Basak M, Kızıl S, Lucas SJ, Uzun B (2021) A high-density SNP genetic map construction using ddRAD-seq and mapping of capsule shattering trait in sesame. Front Plant Sci. :917
Li C, Duan Y, Miao H, Ju M, Wei L, Zhang H (2021) Identification of candidate genes regulating the seed coat color trait in sesame (Sesamum indicum L.) using an integrated approach of QTL mapping and transcriptome analysis. Front Genet 12:700469
Liang J, Sun J, Ye Y, Yan X, Yan T, Rao Y et al (2021) QTL mapping of PEG-induced drought tolerance at the early seedling stage in sesame using whole genome re-sequencing. PLoS ONE 16(2):e0247681
Teboul N, Gadri Y, Berkovich Z, Reifen R, Peleg Z (2020) Genetic architecture underpinning yield components and seed mineral–nutrients in sesame. Genes 11(10):1221
Zhang Y, Wang L, Gao Y, Li D, Yu J, Zhou R et al (2018) Genetic dissection and fine mapping of a novel dt gene associated with determinate growth habit in sesame. BMC Genet 19(1):38
Rao P, Prasuna K, Anuradha G, Srividya A, Vemireddy LR, Shankar VG et al (2014) Molecular mapping of important agro-botanic traits in sesame. Electron J Plant Breed 5(3):475–488
Zhang Y, Wang L, Li D, Gao Y, Lu H, Zhang X (2014) Mapping of sesame waterlogging tolerance QTL and identification of excellent waterlogging tolerant germplasm. Sci Agric Sin 47:422–430
Dossa K, Mmadi MA, Zhou R, Liu A, Yang Y, Diouf D et al (2020) Ectopic expression of the sesame MYB transcription factor SiMYB305 promotes root growth and modulates ABA-mediated tolerance to drought and salt stresses in Arabidopsis. AoB Plants 12(1):plz081
Mehmood M, Khan MJ, Khan MJ, Akhtar N, Mughal F, Shah STA et al (2022) Systematic analysis of HD-ZIP transcription factors in sesame genome and gene expression profiling of SiHD-ZIP class I entailing drought stress responses at early seedling stage. Mol Biol Rep 49(3):2059–2071
You J, Zhang Y, Liu A, Li D, Wang X, Dossa K et al (2019) Transcriptomic and metabolomic profiling of drought-tolerant and susceptible sesame genotypes in response to drought stress. BMC Plant Biol 19(1):267
You J, Wang Y, Zhang Y, Dossa K, Li D, Zhou R et al (2018) Genome-wide identification and expression analyses of genes involved in raffinose accumulation in sesame. Sci Rep 8(1):4331
Su RDS, Dossa K, Zhou R, Liu A, Zhong Y, Fang S, Zhang X, Wu Z, You J (2022) Genome-wide characterization and identifcation of candidate ERF genes involved in various abiotic stress responses in sesame (Sesamum indicum L). BMC Plant Biol 22:256
Dossa K, Li D, Wang L, Zheng X, Liu A, Yu J et al (2017) Transcriptomic, biochemical and physio-anatomical investigations shed more light on responses to drought stress in two contrasting sesame genotypes. Sci Rep 7(1):8755
Dossa K, Li D, Wang L, Zheng X, Yu J, Wei X et al (2017) Dynamic transcriptome landscape of sesame (Sesamum indicum L.) under progressive drought and after rewatering. Genomics data 11:122–124
Wang L, Dossa K, You J, Zhang Y, Li D, Zhou R et al (2021) High-resolution temporal transcriptome sequencing unravels ERF and WRKY as the master players in the regulatory networks underlying sesame responses to waterlogging and recovery. Genomics 113(1):276–290
Zhou W, Song S, Dossou SSK, Zhou R, Wei X, Wang Z et al (2022) Genome-wide association analysis and transcriptome reveal novel loci and a candidate regulatory gene of fatty acid biosynthesis in sesame (Sesamum indicum L). Plant Physiol Biochem 186:220–231
Zoclanclounon YABRM, Chung NJ, Mo Y, Karlovsky P, Dossa K (2022) Characterization of peroxidase and laccase gene families and in Silico Identification of potential genes involved in Upstream Steps of Lignan formation in Sesame. Life 12:1200
Wang L, Dossou SSK, Wei X, Zhang Y, Li D, Yu J et al (2020) Transcriptome dynamics during black and white sesame (Sesamum indicum L.) seed development and identification of candidate genes associated with black pigmentation. Genes 11(12):1399
Marakali S (2018) Identification and functional analyses of new sesame miRNAs (Sesamum indicum L.) and their targets. Mol Biol Rep 45:2145–2155
Zhang Y-P, ZY-Y TK, Zhang F, Hu F, Zhang J-G, Wei P-C, Wei Z-J (2021) Integration of miRNAs, Degradome, and Transcriptome Omics uncovers a Complex Regulatory Network and provides insights into lipid and fatty acid synthesis during Sesame seed development. Front Plant Sci 12:709197
Baghery MAKS, Dahestani A, Mehrabanjoubani P, Naghizadeh MM, Masoudi A (2022) Insight into gene regulatory networks involved in sesame (Sesamum indicum L.) drought response. Biologia 77:1181–1196
Sheng CSS, Zhou R, Li D, Gao Y, Cui X, Tang X, Zhang Y, Tu J, Zhang X, Wang L (2021) QTL-Seq and transcriptome analysis disclose major QTL and candidate genes Controlling Leaf size in Sesame (Sesamum indicum L). Front Plant Sci 12:580846
Song Q, Joshi M, Wang S, Johnson CD, Joshi V (2021) Comparative analysis of root transcriptome profiles of sesame (Sesamum indicum L.) in response to osmotic stress. J Plant Growth Regul 40(4):1787–1801
Zhang Y, Wei M, Liu A, Zhou R, Li D, Dossa K et al (2019) Comparative proteomic analysis of two sesame genotypes with contrasting salinity tolerance in response to salt stress. J Proteom 201:73–83
Pamei I, Makandar R (2022) Comparative proteome analysis reveals the role of negative floral regulators and defense-related genes in phytoplasma infected sesame. Protoplasma. :1–13
Weckwerth W (2003) Metabolomics in systems biology. Annu Rev Plant Biol 54(1):669–689
Brennan L (2017) The nutritional metabolomics crossroads: how to ensure success for dietary biomarkers. Oxford University Press
Kanu PJ, Zhu K, Kanu JB, Zhou H, Qian H, Zhu K (2007) RETRACTED: biologically active components and nutraceuticals in sesame and related products: a review and prospect. Trends Food Sci Technol 18(12):599–608
Bose U, Hewavitharana AK, Ng YK, Shaw PN, Fuerst JA, Hodson MP (2015) LC-MS-Based metabolomics study of marine bacterial secondary metabolite and antibiotic production in Salinispora arenicola. Mar Drugs 13(1):249–266
Wang D, Zhang L, Huang X, Wang X, Yang R, Mao J et al (2018) Identification of nutritional components in black sesame determined by widely targeted metabolomics and traditional chinese medicines. Molecules 23(5):1180
Dossou SKWZ, Zhou W, Zhou R, Zhang Y, Li D, Liu A, Dossa K, You J, Wang L The Dark Pigment in the Sesame (Sesamum indicum L.) seed coat: isolation, characterization, and its potential precursors. Front Nutr 2022;Front Nutr 9: 858673
Dossou SSKXF, You J, Zhou R, Li D, Wang L (2022) Widely targeted metabolome profiling of different colored sesame (Sesamum indicum L.) seeds provides new insight into their antioxidant activities. Food Res Int 151:110850
Mekky RHA-SE, Segura-Carretero A (2021). Contreras MdM. Metabolic profiling of the oil of Sesame of the egyptian Cultivar ‘Giza 32’ employing LC-MS and Tandem MS-Based untargeted method. Foods. ;10(298)
Song S, Zhang L, Zhao Y, Sheng C, Zhou W, Dossou SSK et al (2022) Metabolome genome-wide association study provides biochemical and genetic insights into natural variation of primary metabolites in sesame. Plant J.
Dossou SSKXF, Cui X, Sheng C, Zhou R, You J, Tozo K, Wang L (2021) Comparative metabolomics analysis of different sesame (Sesamum indicum L.) tissues reveals a tissue-specific accumulation of metabolites. BMC Plant Biol 21:352
FD. HSaN. Cloning, sequencing, and bioinformatics study of CYP81Q1 gene of iranian sesame (Sesamum indicum L.) cultivar, Modares. J Biotechnol. ;9 277–84
Chandra K, Sinha A, Arumugam N (2019) Gene isolation, heterologous expression, purification and functional confirmation of sesamin synthase from Sesamum indicum L. Biotechnol Rep 22:e00336
Murata J, Ono E, Yoroizuka S, Toyonaga H, Shiraishi A, Mori S et al (2017) Oxidative rearrangement of (+)-sesamin by CYP92B14 co-generates twin dietary lignans in sesame. Nat Commun 8:2155
Zhou R, Liu P, Li D, Zhang X, Wei X (2018) Photoperiod response-related gene SiCOL1 contributes to flowering in sesame. BMC Plant Biol 18(1):1–16
Wei X, Qiu J, Yong K, Fan J, Zhang Q, Hua H et al (2021) A quantitative genomics map of rice provides genetic insights and guides breeding. Nat Genet 53(2):243–253
You J, Li D, Yang L, Dossou SSK, Zhou R, Zhang Y et al (2022) CRISPR/Cas9-Mediated efficient targeted mutagenesis in Sesame (Sesamum indicum L.). Front Plant Sci. ;13
Dossa KZR, Li D, Liu A, Qin L, Mmadi MA, Su R, Zhang Y, Wang J, Gao Y, Zhang X, You J (2021) A novel motif in the 5’-UTR of an orphan gene ‘Big Root Biomass’ modulates root biomass in sesame. Plant Biotechnol J 19:1065–1079
Acknowledgements
The authors express gratitude to the vice president for research and community engagement office of Mekelle University for providing the necessary support of the present work.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
Weldemichael MY conceived and designed the present research, analyzed the data and wrote the manuscript. Gebremeddhn HM wrote and review the manuscript. All the authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not Applicable.
Research involving human participants and/or animals
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors have declared that no competing interests exist.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Weldemichael, M.Y., Gebremedhn, H.M. Omics technologies towards sesame improvement: a review. Mol Biol Rep 50, 6885–6899 (2023). https://doi.org/10.1007/s11033-023-08551-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08551-w