Abstract
Background
SARS-CoV-2 which causes COVID-19 disease has started a pandemic episode all over the world infecting millions of people and has created medical and economic crisis. From December 2019, cases originated from Wuhan city and started spreading at an alarming rate and has claimed millions of lives till now. Scientific studies suggested that this virus showed genomic similarity of about 90% with SARS-CoV and is found to be more contagious as compared to SARS-CoV and MERS-CoV. Since the pandemic, virus has undergone constant mutation and few strains have raised public concern like Delta and Omicron variants of SARS-CoV-2.
Objective
This review focuses on the structural features of SARS-CoV-2 proteins and host proteins as well as their mechanism of action. We have also elucidated the repurposed drugs that have shown potency to inhibit these protein targets in combating COVID-19. Moreover, the article discusses the vaccines approved so far and those under clinical trials for their efficacy against COVID-19.
Conclusion
Using cryo-electron microscopy or X-ray diffraction, hundreds of crystallographic data of SARS-CoV-2 proteins have been published including structural and non-structural proteins. These proteins have a significant role at different aspects in the viral machinery and presented themselves as potential target for drug designing and therapeutic interventions. Also, there are few host cell proteins which helps in SARS-CoV-2 entry and proteolytic cleavage required for viral infection.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coronavirus is a new name for chaos that has developed since December 2019 with pandemic features and proved to be a threat to humans claiming millions of lives till now. The 2019 new coronavirus marked its onset from the Huanan Seafood Wholesale Market, Wuhan city, a Hubei province in China. 27 cases of pneumonia with unknown etiology were reported from this city on 31st December 2019. These patients who worked at or live around the seafood market showed the clinical symptoms of fever, dyspnea, dry cough and bilateral lung infiltrates on imaging [1]. The virus was identified from swab samples of patient’s throat conducted by the Chinese Centre for Disease Control and Prevention (CCDC) as novel beta coronavirus (a member of beta group of coronavirus) on 7th January 2020 and initially named as 2019-nCoV (2019 novel coronavirus) by World Health Organisation (WHO) [2]. Later, International Committee of Taxonomy of Viruses (ICTV) named this novel virus as severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) [3]. As the infectious diseases gradually spread worldwide and became a massive global outbreak, WHO declared the virus as sixth Public Health Emergency of International Concern on January 30th, 2020. WHO officially named this disease as COVID-19 (where CO stands for corona, VI for virus, D for disease and 19 represents the year 2019 in which it first emerged) on February 2020 [2].
Origin and progress of COVID-19
Wuhan is an emerging business hub of China and the seafood market trades in fish and variety of live animal species including poultry, bats, snakes and marmots. The outbreak of novel coronavirus gradually affected more than seventy thousand individual and killed more than eighteen hundred within the first fifty days of pandemic [4]. It was suggested from the genetic sequence of virus that the patients infected with coronavirus in Wuhan, China may have visited this market or may have consumed infected animals as source of food. While cases started increasing exponentially with no record of visiting the seafood market, thus, suggesting the virus having strong potential for human to human transmission [5]. Environmental samples from the Huanan sea food market was taken and tested positive, suggesting that the virus originated from there and likely there is a chance of transmission of pathogens from animals to human [6]. According to a genomic study, it was claimed that the role of Huanan seafood market in propagating of disease is not clear and suggested that the virus may be introduced from an unknown location into seafood market where it spread rapidly [7]. SARS-CoV-2 reported to be phylogenetically related to SARS-like bat CoV, with a sequence similarity of more than 90%, thus, suggesting that bats could be the key reservoir or zoonotic source [8]. Until recently, Lam et al. isolated Malayan pangolin CoV genomes and found 85.5–92.4% similarity to SARS-CoV-2, hence, concluded that it may be the intermediate host for SARS-CoV-2 [9]. Nonetheless, bats either directly transmit SARS-CoV-2 virus or requires an intermediate host to cause infection- this theory needs to be confirmed so that zoonotic transmission patterns could be established and understood [10]. The novel coronavirus has since spread overseas in other regions in Asia, North America, South America, Europe, Africa and Oceania and thus, making it global pandemic. The new coronavirus outbreak has not only caused the downfall of economy in all countries but also brought down medical and public health infrastructure in a tight spot [2]. The novel coronavirus has proven to be more contagious having enhanced transmission rate than SARS and MERS (middle eastern respiratory syndrome) [11].
Novel coronavirus 2019 (SARS-CoV-2)
Coronavirus are large group of viruses that mainly causes infection in respiratory and gastrointestinal tract and present in various species of birds, bats, snakes and other mammals. Coronavirus are named so due to the presence of crown-like bulbous appearance (“corona” means crown) [3]. SARS-CoV-2 belong to the subfamily Orthocoronavirinae in Coronaviridae family and order Nidovirales that consists of enveloped, positive sense ssRNA (single-stranded) genome [12]. They are spherical in shape with club-shaped spikes and a particle size of 125 nm as shown in Fig. 1a.
a Structure of respiratory syndrome causing human coronavirus. This is an Open Access article distributed under the terms of the Creative Commons Attribution Licence (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution and reproduction in any medium provided the original work is properly cited. Copyright@ Elsevier [11]. b The infection cycle of SARS-CoV-2 inside the host cell. The sequence of events, from host cell recognition through the release of new virion, is represented graphically as steps 1 to 12. Reprinted with permission from [91]. c Genome organization of the SARS-CoV-2. The viral genome encodes 16 Non-structural proteins (Nsps) required for replication/transcription along with the structural proteins required for the assembly of new virions. The proteins are marked below the genome with their respective coding regions. A short description of the functions of different proteins is also shown. Reprinted with permission from [91]
The subfamily coronavirinae are further subdivided on the basis of serological pattern into four genera: (a) alphacoronavirus that includes human coronavirus (HCoV)-229E and HCoV-NL63; (b) betacoronavirus includes severe acute respiratory syndrome (SARS-HCoV), HCoV-OC43, HCoV-HKU1 and MERS-CoV; (c) gamma-coronavirus contains viruses of whales and birds and; (d) delta-coronavirus consists of viruses of pigs and birds. SARS-CoV-2 along with SARS-CoV and MERS belong to beta-coronavirus [13]. The life cycle of SARS-CoV-2 infection is illustrated in Fig. 1b.
On the basis of genomic analysis, SARS-CoV-2 showed > 90% homology with bat SARS-like -CoVZXC21, 82% with human SARS-CoV and 50% with MERS-CoV [14]. SARS spread out in 2002 from Guangdong province of south-eastern China, an epidemic with unusual pneumonia and acute respiratory distress syndrome (ARDS) cases affecting 26 countries and affected 8096 people with 774 deaths by 2004. A decade later, similar case of respiratory tract infection came to light in middle east countries (Saudi Arabia, UAE) in 2012 affecting 27 countries in total including US and Malaysia with 2428 cases and 838 deaths [11]. Table 1 summarizes few differences between SARS-CoV-2, SARS and MERS.
SARS-CoV-2 consists a genome of about 20–30 kb in size, encoding a large non-structural polyprotein which are proteolytically cleaved and thus, generate 15/16 proteins, accessory proteins (ORF3a, ORF6, ORF7, ORF8, ORF9) as well as four structural proteins. It also contains 14 open reading frames [14]. At the 5’ terminal region of the genome, ORF1 and ORF2 encode non-structural proteins (nsps) important for virus replication and 3’ terminal encodes structural proteins. The 4 structural proteins are outer spike glycoprotein (S), membrane (M), envelope (E) and nucleoprotein (N) which are important for virus assembly and infection. SARS-CoV-2 consists of 16 nsps encoded by largest gene, orf1ab as well as by orf1a gene [15]. SARS-CoV-2 genomic organisation is depicted in Fig. 1c.
It also expresses other polyproteins and membrane proteins, such as RNA polymerase, papain-like protease (PLpro), 3-chymotrypsin-like protease (3CLpro) and helicase [14]. To gain entry into the human cell, SARS-CoV-2 and SARS-CoV recognizes angiotensin-converting enzyme 2 (ACE2) as a key receptor while MERS requires dipeptidyl peptidase 4 (DPP4). Some other variations have been seen by researchers between SARS-CoV-2 and SARS-CoV. It is marked by the absence of 8a protein and change in number of amino acids in 3c and 8b protein in SARS-CoV-2 [16].
Current status of COVID-19
Antarctica was the only continent free of novel coronavirus but at the end of December 2020, 36 COVID-19 cases were reported. Scientists and WHO are now accessing the risk of potential transmission of coronavirus from humans to Antarctic wildlife and taking appropriate measures to protect the wildlife population (seabirds, penguins, seal, dolphins, whale) [17]. Since its emergence, the virus has undergone some mutations to adapt itself to various environmental factors like weather and population. These mutations of concern have raised public alarm. These mutations are related to proteins such as spike, envelope and membrane. Center for Disease Control and Prevention has classified few of them as variants of concern (VOC) and variants of interests (VOI). The D614G mutation at amino acid position 614 in S protein was reported in the early phase of the pandemic raised public concern. D614G mutant transmitted quickly enough to became globally dominant by June-July 2020 [18]. Due to high evolutionary rate of the virus, its transmissibility has reported to be increased. Around September 2020, another mutation emerged from South Africa with a new variant designated as B.1.351 (Beta variant, VOC) or 501Y.V2 and number of COVID-19 cases started to rise rapidly. In this variant, the mutation, N501Y, was seen in the receptor binding domain (RBD) of S protein along with E484K mutation. It was also reported that due to mutation, the binding efficiency of the virus to cell surface receptor has drastically increased and the virus can also reduce its neutralization against LY-CoV016 (monoclonal antibody); this strain has already spread over 20 countries [19]. Another new variant B.1.1.7 (Alpha variant, VOC) with N501Y mutation similar to 501Y.V1 reported to have arisen independently in SARS-CoV-2 strain in the UK and found to have 70% more transmission capacity [20]. In B.1.1.7 variant, besides N501Y, mutations in 17 other amino acids (including 8 in S protein) and a deletion at amino acid positions 69 and 70 in S protein was also reported. In early 2021, this variant rapidly spread around more than 100 countries in Europe, America and Asia [20]. Scientists reported that antibodies are created against multiple parts of S protein of SARS-CoV-2, so there is a high chance that the vaccines will retain efficacy for these variants [21]. Brazil reported another variant P.1 (Gamma variant, VOC) derived from the lineage B.1.1.28 with mutation present in the S protein primarily responsible for entry of viral particles into human cells [22]. Mutations N501Y and E484K was found in this variant and has high transmissibility. Zeta variant (lineage P.2) carrying E484K mutation derived from gamma emerged in Brazil but has low transmission rate than gamma [22]. In India, two VOIs B.1.617 and B.1.618 as well as B.1.617.2 (Delta variant, VOC), a sub-lineage of B.1.617 was reported and found to be more infectious and transmissible [23]. It has 50% more transmission rate than B.1.1.7 and has already spread across many countries. Delta variant has E484Q and L452R mutations which is involved in increased interaction of virus particles with human receptor cells and hence, increased rate of infection. Shortly after Delta variant, a new strain B.1.617 + (Delta plus variant) was emerged in some part of India during second wave of COVID-19 with a new P618R mutant which is responsible for reduced antibody binding capability as well as evasion of natural immunity along with E484Q and L452R. B.1.617 + has spread four times faster than alpha variant and rapidly expanded to other countries [23]. Another variant of concern known as Omicron (B.1.1.529) has emerged with a large number of mutations including at least 34 mutation in S protein and is more transmissible than Delta variant in late 2021 [24]. This variant has two lineages, BA.1 and BA.2 and has raised major concerns due to its ability to evade protection conferred by therapeutics monoclonal antibodies and vaccines [24]. As per WHO, total COVID-19 cases of 534,245,759 have been reported with 6,317,736 deaths and 505,168,553 recovered worldwide in 228 countries and territories till this date [25].
In this review, all the structural proteins and non-structural proteins (nsps) of SARS-CoV-2 that are involved in causing COVID-19 infection in humans have been comprehensively discussed. The interaction of these proteins with the cell receptor (ACE2) to gain entry into the host and its role in processes such as proteolytic cleavage, fusion, transcription, translation, viral packaging, assembly and exocytosis in life cycle of the virus have been explained in extensive detail. Repurposing of the antiviral drugs which have shown the efficacy to target proteins in any step of the viral life cycle have been outlined as well as illustrated. Other therapeutic options such as immunotherapy and cellular therapy as well as vaccines approved have been summarised as well.
Key proteins responsible for SARS-CoV-2 and possible druggable sites
Spike protein and its interaction with ACE2
The spike (S) protein of coronavirus is a large glycoprotein of about 180 kDa containing approximately 1273 amino acids and 20 asparagine-linked glycans. Spike glycoprotein present on the surface of novel coronavirus (SARS-CoV-2) as a homotrimer, plays an important role in the attachment of virus to receptors of the host cells as well as membrane fusion. The trimers are formed from S monomers in the endoplasmic reticulum (ER) of virus producing cells [26]. S protein consists of three segments: large ectodomain, single-pass transmembrane anchor and short intracellular tail. Ectodomain are composed of two functional domains/subunits: S1 domain is responsible for receptor binding and S2 domain is responsible for fusing viral and host cell membrane. The S1 domain (14-685 residues) is further divided into N-terminal subdomain (NTD) and C-terminal subdomain (or C-domain) [27]. S2 domain (686-1273 residues) consists of fusion peptide, heptapeptide repeat sequence (HR1, HR2), transmembrane domain and cytoplasmic domain. S protein trimer is located on the surface of viral envelope and have large number of N-linked glycans that are essential for proper S folding and for controlling the accessibility of host proteases and neutralizing antibodies of host. There are 22N-linked glycosylation sequons per protomer in SARS-CoV-2 and out of which 20 are in homology with SARS-CoV S protein [28]. Studies have reported that the degree of amino acid sequence homology between S protein of SARS-CoV-2 and SARS-CoV is 76% [29]. Researchers have found that the receptor binding motif (RBM) of S glycoprotein of SARS-CoV-2 is similar to that of pangolin coronavirus S protein and it has been suggested that pangolin is involved in evolution of SARS-CoV-2 [30]. As compared to other coronavirus, SARS-CoV-2 have been found to contain 27 mutations in genes encoding viral S protein [26]. SARS-CoV-2 recognizes exopeptidases while other coronavirus also recognizes exopeptidases as well as aminopeptidases or carbohydrate as receptor for entry into human cells. SARS-CoV-2’s S protein was confirmed to have increased binding affinity to human ACE2 receptor than 2002 strain of SARS due to the single N501T mutation in S protein of SARS-CoV-2 which significantly enhanced its binding efficiency [31]. However, Othman et al. stated that mutation of residue Q493 to N493 rather than N501T is the main reason for higher binding affinity as it satisfies Van der Waals due to the presence of longer side chain of asparagine [32].
S protein is multifunctional which controls the invasion of virus into host cells and ultimately causing infection. Receptor recognition is the initial step in viral infection and the key determinant of host cell and tissue tropism. ACE2 is present on plasma membrane of cell of various tissues, especially on lower respiratory tract, heart, lung, small intestine, kidney, and gastrointestinal tract. It is a type I transmembrane metallo-glycoprotein which consists of an extracellular catalytic domain. An 805-amino acids long ACE2 has amino-terminal as well as carboxy-terminal domain. The major function of ACE2 is to degrade angiotensin II (potent vasoconstrictor) to form angiotensin-(1–7) and therefore, negatively regulates renin-angiotensin system (RAS). A single amino acid is cleaved by ACE2; hence they are called mono carboxypeptidase. It plays protective function in cardiovascular system and other organs [33]. A study conducted by Zhou et al. showed that SARS-CoV-2 uses ACE2 from humans, horseshoe bats, pigs and civet cats to enter the ACE2-expressing HeLa cells [8]. An in vitro study revealed that cytopathic effects occurs when SARS-CoV-2 is present on the surface of human airway epithelial cells followed by cessation of cilia movements [18]. Downregulation of this ACE2 is responsible for pathogenesis of acute lung injury and ultimately ARDS [34].
During infection, the viral S protein is processed at the boundary between S1 and S2 subunits (S1/S2 cleavage site) by the host cell, furin-like proteases. This step is known as priming which divides the S protein into N-terminal S1 subunit that recognises cell receptor and membrane-bound C-terminal S2 region that helps in viral entry. Both the subunits are reported to be non-covalently bound in prefusion conformation. A conserved receptor binding domain (RBD) is present in S1 region consisting of approximately 193 amino acid fragment which recognizes and binds ACE2 receptor [35]. Zhang et al. investigated the amino acid phylogenetic tree concluding that S1 protein of SARS-CoV-2 is more closely related to Pangolin-CoV [36]. The RBD of both are highly conserved and there is one amino acid change only. Five key amino acid residues at positions 442, 472, 479, 487 and 491 in S protein are critical in binding ACE2 as they are present at receptor complex interface with ACE2. Except Tyr491, all other residues in RBD are not conserved when compared to SARS-CoV [37]. When S1 subunit binds with ACE2, it tends to destabilize the prefusion trimer and thus, causes shedding of S1 subunit. This leads to the transition of S2 into highly stable postfusion conformation [28]. The RBD then undergo hinge-like conformational movements which can hide (‘down’ conformation) or expose (‘up’ conformation) the determinants of receptor binding. The ‘down’ conformation is the inaccessible state of the receptor and ‘up’ conformation is the accessible state [38]. The most distinctive feature of SARS-CoV-2 S protein is the presence of multibasic furin-like cleavage site (S1/S2) which has been reported to be absent in other SARS-like CoV of beta-coronaviruses. Other SARS-like coronaviruses only contain TMPRSS2 (transmembrane protease, serine 2) or trypsin cleavage site [39].
After receptor binding, S2 domain helps in fusing the viral-host membrane by exposing highly conserved fusion peptide (FP). The S2 domain contains this fusion peptide which is proteolytically cleaved at a site found immediately upstream (S2’). It is the second priming event. FP contains 15–20 conserved amino acids, mainly glycine or alanine which helps in anchoring of virus to target membrane. The S2 domain also contains internal fusion peptide (IFP). Somehow FP and IFP both are likely to be involved in viral entry process, hence, the priming event at both S1/S2 and S2’ sites are necessary [40]. The priming event at S2’ by host cell protease plays a key role in the final activation of the S protein and also regulates viral tropism and pathogenesis. It has also been proposed that one or more furin-like enzymes are involved in S2’ cleavage [35]. After the priming of second site, the S2 subunit undergoes conformational rearrangement when it inserts FP into host membrane and as a result, HR1 and HR2 (Heptad repeat) interacts to form six-helical bundle (6-HB) which brings viral envelope and host membrane in close proximity and marks the completion of fusion followed by entry of virus into cell, release of its content, viral replication and infection of other cells. Apart from proteases involved in cleaving and activation, ionic interactions (H+ and Ca2+) also dictate the entry of virus into host, hence, controlling viral stability and transmission. However, the whole molecular pathway involved in viral entry into host cell are still unclear and needs to be understood completely [41]. Once the virus enters the alveolar epithelial cells, it replicates quickly and triggers strong immune response, thus, causing cytokine storm syndromes (also called hypercytokinaemia) and pulmonary tissue damage [42]. SARS-CoV-2 S-protein has binding free energy of − 50.6 kcal/mol which is more as compared to − 78.6 kcal/mol for SARS-CoV. The reason being loss of hydrogen bond interaction by replacing Arg426 with Asn426 in Wuhan SARS-CoV-2 S-protein [37]. The affinity with which SARS-CoV-2 RBD binds with ACE2 is 10- to 20-fold higher as compared to other CoVs RBD. This may justify the rapid development and enhanced human-to-human transmissibility in COVID-19 [38]. Since the outbreak of pandemic, 96.5% of the mutation has been reported in the S protein sequence [43]. This protein has been an area of interest to target for the development of therapeutic treatment as blocking them will inhibit the growth of infection.
Role of viral proteases
3-chymotrypsin-like protease (3CLpro) is the primary protease of coronavirus that has been characterized as important drug target site. It is the main protease of coronavirus, hence called Mpro or 3C-like protease (also known as nsp5). It is a 33 kDa cysteine protease that plays a key role in replication cycle of the virus. Mpro targets the viral polyproteins, pp1a and pp1ab translated from viral RNA and digests them at 11 conserved sites. As a result, it produces 12 functional nsps (nsp 4, 6–16) and these nsps further responsible for viral replication and viral assembly. Mpro contains 11 cleavage site and the recognition sequence is Leu-Gln (cleavage site) (Ser, Ala, Gly) [44]. The crystal structure and amino acid analysis revealed that Mpro of SARS-CoV-2 showed approximately 96% sequence similarity with other members of coronavirus family and most of the residues are conserved in Mpro. The co-crystallized structure showed that Mpro functions as an active homodimer with 303 amino acid residues and divide into three domains. Domain I (residues 8–101) and II (residues 102–184) has chymotrypsin-like six-stranded antiparallel β-sheets while domain III (residues 201–303) contains 5 antiparallel α-helices and are connected to domain II via loop region (residues 185–200). The substrate binding site is located between domain I and II with residues 164–168 and 189–191 residues of loop region, near Cys-His catalytic dyad. Inhibiting the activity of Mpro will stop the viral replication, hence, this enzyme is an attractive drug target site [45].
Along with Mpro, another proteolytic enzyme, papain-like protease (PLpro) is equally important for viral lifecycle. Both the proteases are responsible for processing of pp1a and pp1ab replicase polyprotein precursors into functional proteins. PLpro cleaves the polyproteins at 3 distinct sites, thus, yielding nsp1, nsp2 and nsp3. The proteolytic recognition sequence of PLpro is LXGG↓X [46]. In addition to proteolytic activity, PLpro also represses the innate immune system of the host through de-ubiquitination and de-ISGylation (interferon-stimulated gene product 15) events as they hijack the ubiquitin (Ub). Ubiquitin (Ub) and ISG15 both carries the recognition sequence LXGG at the C-terminus [47]. PLpro hampers the host anti-viral reactions as ubiquitin plays an important role in host defence mechanism. It shuts down the crucial pathway by cleaving ISG15 (two domain Ub-like protein) and Lys48- linked poly Ub chains and thus, inhibiting the activation of interferon regulatory factor-3 pathway. Notably, the overall sequence similarity found between PLpro of SARS-CoV and SARS-CoV-2 is 83% and both the PLpro differs by 54 residues. The ubiquitin-like domain and catalytic domain of SARS-CoV-2 PLpro are distinct and well separated from each other. PLpro is a cysteine protease with zinc ion as central atom coordinated by four cysteine residues. The active site of SARRS-CoV-2 PLpro is conserved and has a catalytic triad (cysteine, histidine and aspartic acid). With dual function, PLpro proves to be an important drug target and inhibition of this protein will improve the antiviral response of host immune system [48]. Many studies have revealed and reported the structures of proteases (PLpro and Mpro) of SARS-CoV-2 and drug screening has been reported to find out the potent candidate for inhibition of proteases.
RNA-dependent RNA polymerase
RNA dependent RNA Polymerase (RdRp) is the nsp12 which is 103 kDa long and play a crucial role in the life cycle of viruses like coronaviruses, zika virus, and hepatitis C virus. It is the main component of replication/transcription machinery and hence, participates actively in the replication of viral RNA from RNA template with the help of co-factors nsp-7 and nsp-8 and forms tripartite polymerase complex with them. These nsps activate and confers processivity to the RdRp. Further, this complex gets associated with nsp-14 and confers proofreading exonuclease function [49]. The structure of SARS-CoV-2 RdRp (nsp12)-nsp7-nsp8 complex (160 kDa) was resolved recently by cryo-EM study and found that it is similar to SARS-CoV RdRp complex. SARS-CoV-2 RdRp has two domains: NiRAN (N-terminal extension nidovirus-unique RdRp-associated nucleotidyl transferase) domain from residues 60–249 and RdRp domain from residues 366–920. Both the domains are interconnected by interface domain. NiRAN domain contains kinase-like fold and N-terminal β-hairpin region located between NiRAN and RdRp domain. Two Zn2+ ions are located in the NiRAN domain away from RdRp catalytic site. It has been reported that 96% sequence similarity was found between SARS-CoV RdRp and SARS-CoV-2 RdRp [50]. So, it forms an interesting target for docking studies and molecules used against SARS-CoV RdRp can be effective against SARS-CoV-2 RdRp. Table 2 summarises the functional feature of nsps and structural protein of COVID-19.
Role of other viral structural protein
Nucleocapsid (N) protein
The nucleocapsid (N) protein (46 kDa) of coronaviruses are important structural protein which are highly expressed and abundant in the virus after infection. It is a multifunctional RNA-binding protein that is essential for RNA transcription, and replication [51]. After entering host cell, its main function includes binding the RNA of virus, packing them into a long helical ribonucleoprotein (RNP) complex and processing the viral particle assembly and release. They also modulate infected cell metabolism. They are highly conserved and consists of three parts: N-terminal RNA-binding domain (NTD) responsible for RNA binding, C-terminal dimerization domain (CTD) for oligomerization, and an intrinsically disordered central Ser/Arg (SR)-rich linker for primary phosphorylation [52]. Both NTD and CTD are rich in β-strands with some short helices and surface of both the domain contains highly positively charged regions thus, enhancing the ability of N protein to bind non-specific nucleic acids. It was reported that N protein of SARS-CoV-2 show 90.52% homology with that of SARS-CoV N protein [53]. They are highly immunogenic as they can induce strong immune response against SARS-CoV-2 during infection. It could be a suitable drug-targeting candidate as they play a pivotal role in genome packing, viral transcription and assembly in the host cell [54]. A study conducted by Guo et al. showed that IgG in serum of COVID-19 infected patients can bind with N antigen [55]. Another study found the presence of IgG, IgA and IgM antibodies in COVID-19 recovering patients against N antigen [56].
Membrane and envelope protein
The membrane (M) and envelope (E) structural proteins plays an important role in regulating the assembly of virus particles. S, M and E proteins contains trafficking signal sequences and gets incorporated in endoplasmic reticulum along with ribonucleoprotein complex which is essential for maturation and budding of new virus particles. Both M and E proteins show more than 90% sequence similarity with SARS-CoV proteins and are conserved [57]. M protein of SARS-CoV-2 is a type-III transmembrane glycoprotein which is 230 amino acid long and the most abundant of all structural proteins. It consists of 3 domains: N-terminal ectodomain, C-terminal endo-domain and transmembrane helices (TMH1, TMH2, TMH3). The M protein shows homotypic (interaction with itself) as well as heterotypic (interaction with other structural proteins) interaction. These interactions induces membrane bending (budding) and hence, serve as checkpoint for assembly of new virus particles. Homotypic interaction occurs through TM region of M protein while heterotypic interaction with N and E protein occurs through C-terminal endo-domain. M protein can also induce humoral immune response as the antigenic epitopes are located in the TM1 and TM2 region of SARS-CoV-2 [58].
E protein is the smallest structural protein of SARS-CoV-2 with 75 amino acids long and contains 3 domains: hydrophilic N-terminal ectodomain, transmembrane domain (hydrophobic) and hydrophilic C-terminal endo-domain. According to NMR study, E protein display structure similar to viroporins with pentameric helix bundle that surrounds a narrow hydrophilic cationic central pore. These pores or ion channels activates the host inflammasome by causing the loss of membrane potential. The cysteine residues present in C-terminal domain undergoes palmitoylation that are essential for subcellular trafficking and membrane binding. The last four amino acids (Asp-Leu-Leu-Val) ae involved in the interaction of E protein with host associated proteins (PALS1 and syntenin) thus, causing viral dissemination. These events have been reported to induce cytokine storm. Inhibition of E protein can hinder viral maturation as well as viral propagation and thus, developing antiviral drug and vaccine targeting E protein can be good option for treatment [59].
Role of host proteases
Furin-like protease
For cleavage of S protein, the host proteases differ for different coronaviruses which determines the epidemiological as well as pathological features of virus. For example, trypsin, human airway trypsin-like protease (HAT) and TMPRSS2 are some host proteases which are expressed in many essential organs [60]. A study by Wang et al. found that COVID-19 has a novel multibasic unique cleavage site (–RRAR–) in S1/S2 domain, located between residues 682 and 685 which is distinct from other coronaviruses [39]. This site is most likely to be cleaved by convertase furin which enhances viral-host cell membrane fusion. Furin cleaves proteins and peptides precursors and converts them into biologically active state. It is a type 1 membrane-bound serine-protease and a member of calcium-dependent, subtilisin-like proprotein convertase family. It cycles from the trans-Golgi network to the cell membrane and through endosomal system. It specifically recognizes and cleaves the R-X-K/R-R motif in the presence of Ca2+ [61]. This protease is highly expressed in organs and tissues including lung, gastrointestinal tract, brain, pancreas, and reproductive tissues and liver as well. Hence, COVID-19 can infect these organs also resulting in systematic infection of virus in the body and an enhanced transmission and pathogenicity [39]. The binding between the furin protease and S protein is in a clamp-like fashion where furin clips tightly to the cleavage S1/S2 site. Substrate-binding pocket of furin has canyon-like crevice and its key amino acid residues are specifically positioned to interact with S glycoprotein. The presence of 12 additional nucleotides upstream of the exposed -RRAR- sequence in S glycoprotein corresponds to unique canonical furin-like cleavage site. These amino acid residues present between N657 and Q690 of S glycoprotein strongly interacts with furin are well organised in a flexible loop [62]. COVID-19 use this convertase to activate S glycoprotein and thus, provide a gain-of-function to the virus for efficient transmission. Van der waal or hydrogen bonding facilitates the interaction of furin and S glycoprotein. MERS-CoV contains -RSVR- sequence which is most probably cleaved by furin during virus egress whereas SARS-CoV’s S protein remains uncleaved [63].
TMPRSS2 (Transmembrane protease serine 2)
TMPRSS2 is a type II transmembrane enzyme of the host that belongs to serine protease family and encoded by TMPRSS2 gene. Along from S protein-ACE2 interaction, other proteases also play a major role in entry of the virus particle into the host cell. TMPRSS2 helps in the priming of S protein thereby causing the fusion of cellular and viral membranes. TMPRSS2 are found to be localized in epithelial cells of lungs [64]. A study was reported in which TMPRSS2 cells were overexpressed in Vero E6 cell line due to which chances of corona infection elevated in patients as this overexpression of TMPRSS2 in lung made them more vulnerable to SARS-CoV-2 [65]. Much research has proved that entry of novel SARS-CoV-2 depends on TMPRSS2 priming activity and gets blocked when TMPRSS2 inhibitor, camostat was used [66]. So, TMPRSS2 proved to be another protein target for treatment of SARS-CoV-2. A TMPRSS2 knockout mice model was studied and it was reported that knockout mice were immune to coronavirus [67].
Cathepsin L
Cathepsins are host cysteine proteases and play an important role in the entry of SARS-CoV-2 viral particle into the host cells via endocytic pathway and also in protein catabolism in endosomes and lysosomes. Cathepsin L causes the priming of S proteins after the virus enters the endosomes and hence, causing the fusion of viral and endosomal membranes and release of viral genome into cytoplasm. Cathepsin L can be targeted for the treatment of SARS-CoV-2 as its inhibitors have been reported to prevent pulmonary fibrosis. Some cathepsin L inhibitors are SID 26,681,509 and E-64-d [68]. A study reported that a glycopeptide antibiotic, teicoplanin, was able to block cathepsin L activity and thus, inhibits the entry of SARS-CoV and MERS-CoV [69]. Another study reported the inhibition of SARS-CoV-2 pseudoviruses entry into the host cells. It was demonstrated that SARS-CoV-2 pseudo virus uses the cathepsin L specifically to enter into 293/hACE2 cells and E-64-d (a broad spectrum cathepsin inhibitor) proved to reduce the viral entry by 92.5% while SID 26,681,509 inhibitor reduced it by 76% [70]. Different types of cathepsins have been found to play a key role in viral entry to host cells and targeting them along with other target protein can be more beneficial in treating SARS-CoV-2 patients [71].
Therapeutic interventions for COVID-19
COVID-19 is a serious international concern and research teams and health officials from all around the world are working tirelessly to cope with the disease. Since its outbreak, countries have taken measures to slow down the spread of virus by announcing lockdown, testing, isolating, and treating patients with drugs, carrying out contact tracing, limiting travel and quarantining citizens. Some of the methods to treat COVID-19 infected patients are listed below.
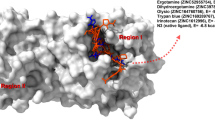
All the proteins associated with SARS-CoV-2 has been proved to be potential catalytic site to target in order to treat the infection and different treatment options has been developed till now. Initially when there were no specific antiviral drugs developed for coronavirus, drugs developed for MERS and SARS showed promising results, so expectation was shifted to them. Table 3 describes the antiviral drugs which have been developed by scientist for SARS-CoV-2 as well as some repurposed drugs evaluated for their potency to counter SARS-CoV-2 infection. Most synthetic small molecules under clinical trials are being repurposed for COVID-19 which are already reported for their efficiency against other disease states [71]. Drugs are being clinically tested to be used against COVID-19 infection such as Favipiravir, Ribavirin, Nafamostat, Nitazoxanide, Penciclovir, Favipiravir, Baricitinib and Arbidol. However, most of them showed moderate results when tested on clinical samples of COVID-19 patients in vitro [11]. Remdesivir, has caught the attention of many researchers due to its promising impact against the virus. It is an adenosine nucleoside analogue and targets viral RNA-dependent RNA polymerase, invades the viral RNA chains and thus, causing pre-mature chain termination. It also evades proofreading viral exoribonuclease. This drug has been used effectively against Ebola virus infection [72]. Sheahan et al. conducted clinical study stating that SARS-CoV-2 replication was significantly blocked in patients and were clinically recovered when remdesivir alone or in combination with cholorquine/ interferon beta was used [73]. Remdesivir was successfully used in treatment of first COVID-19 patient in USA and ameliorated the worsening condition of pneumonia on 7th day of hospitalization in January 2020 [74]. Remdesivir has been found to work effectively with compassionate use in severe cases of COVID-19 though extensive research is still going on. Controlled trials of antiviral drugs are required to determine side effects, if any [73]. Terali et al. reported in silico study to find out drugs which can be repurposed as potential human ACE2 inhibitors like lividomycin, quisinostat, burixafor, fluprofylline, spirofylline, pemetrexed, diniprofylline and edotecarin. They proved to be promising drug candidate which blocks viral entry [75]. Food and Drug administration (FDA) approved drugs like Lopinavir, Ritonavir, Darunavir, Boceprevir, Telaprevir are being investigated as Mpro inhibitors and has shown encouraging result. Compounds like Disulfiram, Baicalein, Ebselen, Carmofur, Shikonin, PX-12, Camostat, Calpeptin, Calpain inhibitor and Tideglusib have been reported to be likeable protease-based drug candidates for SARS-CoV-2 with good potency and selectivity [76]. Scientists are trying to find or develop more effective antiviral drug candidate to treat COVID-19. Figure 2 shows the various inhibitors acting at different stages in SARS-CoV-2 infection cycle.
Use of monoclonal or polyclonal antibodies is another suggested method which can serve as prophylactic and therapeutic tool against viral infection and provide some restitution in this pandemic condition. They are laboratory engineered molecules which are designed to bind antigens. It has been reported that monoclonal antibodies (mAbs) target the S protein of SARS-CoV and prevent virus to enter the host cells [77]. Some of the mAbs have been described in the Table 3. To improve the condition of COVID-19 infected patients, other treatment options were also considered. Convalescent plasma containing IgG, IgA, IgM, IgE and IgD was obtained from patients and has already been used effectively for treating SARS-CoV, poliomyelitis, influenza A (H1N1) and Ebola virus infection. The presence of antiviral antibodies in the plasma of clinically recovered patients helps to suppress viremia [78]. Shen et al. reported that after treatment of critically ill patients of COVID-19 and ARDS with convalescent plasma, there was improvement in their clinical condition. The plasma was obtained from 5 patients recovered from this virus. This transfusion with convalescent plasma leads to the normalization of body temperature of patients within 3 days, resolution of ARDS at 12th day of transfusion, decline in viral load and increase in neutralizing antibody titres. The patients were also administered with antiviral agents and methylprednisolone [79]. It is a form of passive immunization that can be achieved from previously viral infected patients. It was authorized by FDA in August 2020 but newer studies suggest that it is less effective [80].
Natural killer (NK) cells, a type of cytotoxic lymphocytes, are immune cells responsible for providing rapid defense response against virus-infected or malignant cells. Antibody-dependent cellular cytotoxicity (ADCC) is a process initiated by NK cells by causing lysis of antibody-coated virus-infected cells. So, they are specific for all virus-infected cells [81]. It has been reported that NK cells can mediate ADCC against SARS-CoV, cytomegalovirus virus, HIV and show antiviral activity. Sorrento and Celularity disclosed that a clinical collaboration has been undertaken which aims to utilize the CYNK-001 (an allogeneic, umbilical cord blood-derived) NK cell therapy to treat COVID-19 infected patients. NK cell therapy is a very promising strategy which enhances immunity [10]. Mesenchymal stem cells (MSCs) are multipotent cells which are found in bone marrow, are known to have strong immunomodulatory and anti-inflammatory functions. They have a great potential of self-renewal and differentiation. Besides bone marrow, placenta and umbilical cord blood are also considered as a source of MSCs. MSCs have the ability to inhibit abnormal activation of macrophages and T-lymphocytes and causing them to differentiate into anti-inflammatory macrophages and regulator T-cell subsets (Treg) [82, 83]. Studies have been conducted on MSCs and found that it can improve ARDS and acute/chronic lung injury by reducing pro-inflammatory cytokine secretion [84]. They can suppress cytokine storm by inhibiting the secretion of IL-1, IL-6, IL-12, TNF-α and IFN-γ. They also tend to repair tissue rapidly and reduces lung fibrosis by secreting hepatocyte, vascular endothelial and keratinocyte growth factor [85]. Apart from antiviral drugs, ARDS and lung injury needs to be treated in case of severe COVID-19 patients in order to prevent the progression of disease and reduce the level of mortality [86]. MSCs can be proven to become one of the promising therapeutic candidates.
Different vaccines for COVID-19
Understanding the key features, etiology and pathogenesis related to COVID-19 is the main focus of interest for researchers to come up with a suitable vaccine to deal with the current pandemic situation. After the outbreak of SARS and MERS, researchers worldwide began developing vaccines. Due to the major role of S protein of coronavirus and ACE2 human receptor in causing COVID-19, drugs targeting their interaction should be a promising way. Owing to the genomic resemblance and structural similarities of these proteins, therapies and vaccines developed for SARS and MERS may pave way for treatment or prevention of SARS-CoV-2 infection [11]. Vaccines like inactivated or live-attenuated vaccines, viral vector-based protein sub-unit vaccines etc. were produced using different strains of SARS which showed promising result in animal models. Yang et al. developed a DNA vaccine (inactivated whole virus or live-vectored strain of SARS-CoV, AY278741) which has successfully induced the release of neutralizing antibody and reduced viral infection in animal models [87]. Previously developed vaccines for SARS-CoV might be helpful to re-utilize it to facilitate vaccine development for COVID-19.
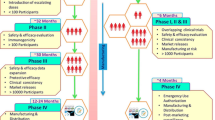
Scientists are working at a breakneck speed to develop vaccines for COVID-19 as early as possible. For this, institutes and pharmaceutical companies are working in collaboration worldwide. More than hundred vaccines are in progress and many of them have even entered clinical trial [88]. Among them, more than 25 have already been gone through clinical trial and has been approved for use till this date. The approved vaccines are being given to people around the world in doses and its efficiency and side effects are being analysed. Till now, some of these vaccines have shown good productivity with little side effects. However, vaccinating people worldwide is a long way to go in completely making world COVID-19 free. Scientists at Gamaleya Research Institute in Russia have successfully made and registered world’s first vaccine, Gam-COVID-Vac, using adenovirus. LNP-encapsulated (lipid nanoparticles) mRNA-based vaccine has been developed by US National Institute of Allergy and Infectious Diseases (NIAID) along with a biotechnology company called Moderna Therapeutics and is currently being administered to people in various countries. The scientists have already investigated the efficiency and side effects of vaccine in adult volunteers. It is a nanoparticle encapsulated vaccine encoding full length, prefusion stabilized S protein [89]. University of Oxford and AstraZeneca has developed a vaccine using Chimpanzee Adenovirus vector (ChAdOx1-5). It has also been tested for safety, reactogenicity and immunogenicity profile as well as tolerability. This vaccine is now called AZD1222/ covishield and is now being given as vaccine in many countries. These adenovirus vectors are weakened version of a common cold virus and is replication-deficient carrying one or few encoded antigens and can stimulate both humoral and cytotoxic T-cell immune responses efficiently [90]. All the vaccines which are approved and those currently in clinical trial in late phase are listed in Table 4.
There are still many challenges which are being faced by scientists working around the clock to fight the battle against COVID-19. Development of vaccines which can perform efficiently can only be done with detailed understanding of the pathway involved in infection. The evolving nature and strain of virus remains the biggest challenge in developing virus-specific vaccine. Many parameters are taken into consideration for rational vaccine design development and for evaluation of vaccine efficacy.
Conclusion and perspective
The novel coronavirus 2019 has challenged the socio-economic, medical and health care foundation worldwide. The deadly virus has already caused devastating impact on human life. The zoonotic source of COVID-19 still needs to be confirmed further which originated from seafood market in Wuhan, China. Bats and pangolins are being considered as key reservoirs according to sequence-based analysis. Research on SARS-CoV-2 has showed many similarities as well genomic variations from SARS and MERS so, a detailed phylogenetic and pathogenetic study will enhance the understanding of virus and help in developing preventive measures. Rapid diagnosis of SARS-COV-2 needs to be done in suspected patients to control the transmission of virus which has already claimed the lives of thousands of people including doctors, health care workers and paramedical staff and infected millions. The human-to-human transmission is a serious concern and a threat to public health. The rigorous surveillance and on-going research will unravel new research findings regrading host adaptation, molecular mechanism, transmissibility, clinical manifestations, evolution, epidemiological pattern and pathogenicity. The new information about COVID-19 is being available every week in scientific journals that will help the public to understand the virus better. Researchers are developing efficient and promising therapeutics strategy to cope with the pandemic situation. Comprehensive measures need to be devised to curb not only this global health emergency but also take care of future outbreak of infection of zoonotic origin.
Data availability
Not applicable.
Abbreviations
- CCDC:
-
Chinese Centre for Disease Control and Prevention
- 2019-nCoV:
-
2019 Novel coronavirus
- WHO:
-
World Health Organisation
- ICTV:
-
International Committee of Taxonomy of Viruses
- SARS-CoV-2:
-
Severe acute respiratory syndrome coronavirus 2
- COVID-19:
-
Corona virus disease 19
- MERS:
-
Middle eastern respiratory syndrome
- VOC:
-
Variants of concern
- VOI:
-
Variants of interests
- RBD:
-
Receptor binding domain
- HCoV:
-
Human coronavirus
- ARDS:
-
Acute respiratory distress syndrome
- S protein:
-
Spike protein
- M protein:
-
Membrane protein
- E protein:
-
Envelope protein
- nsps:
-
Non-structural protein
- PLpro :
-
Papain-like protease
- 3CLpro :
-
3-Chymotrypsin-like protease
- Mpro :
-
Main protease
- ACE2:
-
Angiotensin-converting enzyme 2
- DPP4:
-
Dipeptidyl peptidase 4
- TMPRSS2:
-
Transmembrane protease serine 2
- FP:
-
Fusion peptide
- Ub:
-
Ubiquitin
- FDA:
-
Food and Drug administration
- RdRp:
-
RNA dependent RNA polymerase
- NiRAN:
-
N-terminal extension nidovirus-unique RdRp-associated nucleotidyl transferase
- HAT:
-
Human airway trypsin-like protease
- HIV:
-
Human immunodeficiency virus
- CRISPR:
-
Clustered regularly interspaced short palindromic repeats
- LNP:
-
Lipid nanoparticles
- NIAID:
-
US National Institute of Allergy and Infectious Diseases
- aAPC:
-
Artificial antigen presenting cells
- VLP:
-
Virus-like particles
- CEPI:
-
Coalition for Epidemic Preparedness Innovations
- mAbs:
-
Monoclonal antibodies
References
Lu H, Stratton CW, Tang YW (2020) Outbreak of pneumonia of unknown etiology in Wuhan China: the mystery and the miracle. J Med Virol 92(4):401–402
Sohrabi C, Alsafi Z, O’Neill N et al (2020) World Health Organization declares global emergency: a review of the 2019 novel coronavirus (COVID-19). Int J Surg 76:71–76
Cui J, Li F, Shi ZL (2019) Origin and evolution of pathogenic coronaviruses. Nat Rev Microbiol 17(3):181–192
Lai CC, Shih TP, Ko WC et al (2020) Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and corona virus disease-2019 (COVID-19): the epidemic and the challenges. Int J Antimicrob Agents 55(3):105924
Singhal T (2020) A review of coronavirus disease-2019 (COVID-19). Indian J Pediatr 87:281–286
Huang C, Wang Y, Li X et al (2020) Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395(10223):497–506
Yu WB, Tang GD, Zhang L et al (2020) Decoding the evolution and transmissions of the novel pneumonia coronavirus (SARS-CoV-2/HCoV-19) using whole genomic data. Zool Res 41(3):247–257
Zhou P, Yang XL, Wang XG et al (2020) A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579(7798):270–273
Lam TTY, Jia N, Zhang YW et al (2020) Identifying SARS-CoV-2-related coronaviruses in Malayan pangolins. Nature 583:282–285
Li H, Liu SM, Yu XH et al (2020) Coronavirus disease 2019 (COVID-19): current status and future perspective. Int J Antimicrob Agents 55(5):105951
Shereen MA, Khan S, Kazmi A et al (2020) COVID-19 infection: origin, transmission, and characteristics of human coronaviruses. J Adv Res 24:91–98
Chan JFW, Kok KH, Zhu Z et al (2020) Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg Microbes Infect 9(1):221–236
Harapan H, Itoh N, Yufika A et al (2020) Coronavirus disease 2019 (COVID-19): a literature review. J Infect Public Health 13(5):667–673
Lu R, Zhao X, Li J et al (2020) Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet 395(10224):565–574
Yuan Y, Cao D, Zhang Y et al (2017) Cryo-EM structures of MERS-CoV and SARS-CoV spike glycoproteins reveal the dynamic receptor binding domains. Nat Commun 8(1):1–9
Chen Y, Liu Q, Guo D (2020) Emerging coronaviruses: genome structure, replication, and pathogenesis. J Med Virol 92(4):418–423
Frame B, Hemmings AD (2020) Coronavirus at the end of the world: Antarctica matters. Soc Sci Hum Open 2(1):100054
Zhang L, Jackson CB, Mou H et al (2020) The D614G mutation in the SARS-CoV-2 spike protein reduces S1 shedding and increases infectivity. BioRxiv
Planas D, Bruel T, Grzelak L et al (2021) Sensitivity of infectious SARS-CoV-2 B. 1.1. 7 and B. 1.351 variants to neutralizing antibodies. Nat Med 27(5):917–924
Collier DA, De Marco A, Ferreira IA et al (2021) Sensitivity of SARS-CoV-2 B. 1.1. 7 to mRNA vaccine-elicited antibodies. Nature 593(7857):136–141
Kirby T (2021) New variant of SARS-CoV-2 in UK causes surge of COVID-19. Lancet Respir Med 9(2):e20-21
da Silva JC, Felix VB, Leão SABF et al (2021) New Brazilian variant of the SARS-CoV-2 (P1) of COVID-19 in Alagoas state. Braz J Infect Dis 25(3):101588
Kirola L (2021) Genetic emergence of B. 1.617. 2 in COVID-19. New Microbes New Infect 43:100929
Liu L, Iketani S, Guo Y et al (2022) Striking antibody evasion manifested by the Omicron variant of SARS-CoV-2. Nature 602:676–681
Worldometers.info. COVID-19 CORONAVIRUS PANDEMIC. 2021. worldometers.info/coronavirus/
Belouzard S, Millet JK, Licitra BN et al (2012) Mechanisms of coronavirus cell entry mediated by the viral spike protein. Viruses 4(6):1011–1033
Hulswit RJG, De Haan CAM, Bosch BJ (2016) Coronavirus spike protein and tropism changes. Adv Virus Res 96:29–57
Walls AC, Xiong X, Park YJ et al (2019) Unexpected receptor functional mimicry elucidates activation of coronavirus fusion. Cell 176(5):1026–1039
Walls AC, Park YJ, Tortorici MA et al (2020) Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 181(2):281–292
Wong MC, Cregeen SJJ, Ajami NJ et al (2020) Evidence of recombination in coronaviruses implicating pangolin origins of nCoV-2019. BioRxiv
Wan Y, Shang J, Graham R et al (2020) Receptor recognition by the novel coronavirus from Wuhan: an analysis based on decade-long structural studies of SARS coronavirus. J Virol 94(7):e00127-e220
Othman H, Bouslama Z, Brandenburg JT et al (2020) Interaction of the spike protein RBD from SARS-CoV-2 with ACE2: similarity with SARS-CoV, hot-spot analysis and effect of the receptor polymorphism. Biochem Biophys Res Commun 527(3):702–708
Kuba K, Imai Y, Ohto-Nakanishi T et al (2010) Trilogy of ACE2: a peptidase in the renin–angiotensin system, a SARS receptor, and a partner for amino acid transporters. Pharmacol Ther 128(1):119–128
Imai Y, Kuba K, Ohto-Nakanishi T et al (2010) Angiotensin-converting enzyme 2 (ACE2) in disease pathogenesis. Circ J 74(3):405–410
Coutard B, Valle C, de Lamballerie X et al (2020) The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antivir Res 176:104742
Zhang T, Wu Q, Zhang Z (2020) Probable pangolin origin of SARS-CoV-2 associated with the COVID-19 outbreak. Curr Biol 30(7):1346–1351
Xu X, Chen P, Wang J et al (2020) Evolution of the novel coronavirus from the ongoing Wuhan outbreak and modeling of its spike protein for risk of human transmission. Sci China Life Sci 63(3):457–460
Wrapp D, Wang N, Corbett KS et al (2020) Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367(6483):1260–1263
Wang Q, Qiu Y, Li J et al (2020) A unique protease cleavage site predicted in the spike protein of the novel pneumonia coronavirus (2019-nCoV) potentially related to viral transmissibility. Virol Sin 35(3):337–339
Lu G, Wang Q, Gao GF (2015) Bat-to-human: spike features determining ‘host jump’of coronaviruses SARS-CoV, MERS-CoV, and beyond. Trends Microbiol 23(8):468–478
Jaimes JA, André NM, Chappie JS et al (2020) Phylogenetic analysis and structural modeling of SARS-CoV-2 spike protein reveals an evolutionary distinct and proteolytically-sensitive activation loop. J Mol Biol 432(10):3309–3325
Diao B, Wang C, Tan Y et al (2020) Reduction and functional exhaustion of T cells in patients with coronavirus disease 2019 (COVID-19). Front Immunol 11:827
Gurung AB, Ali MA, Lee J et al (2020) Unravelling lead antiviral phytochemicals for the inhibition of SARS-CoV-2 Mpro enzyme through in silico approach. Life Sci 255:117831
Bacha U, Barrila J, Velazquez-Campoy A et al (2004) Identification of novel inhibitors of the SARS coronavirus main protease 3CLpro. Biochemistry 43(17):4906–4912
Ul-Qamar MT, Alqahtani SM, Alamri MA et al (2020) Structural basis of SARS-CoV-2 3CLpro and anti-COVID-19 drug discovery from medicinal plants. J Pharm Anal 10(4):313-319
Gao X, Qin B, Chen P et al (2021) Crystal structure of SARS-CoV-2 papain-like protease. Acta Pharm Sin B 11(1):237–245
Fu Z, Huang B, Tang J et al (2021) The complex structure of GRL0617 and SARS-CoV-2 PLpro reveals a hot spot for antiviral drug discovery. Nat Commun 12(1):1–12
Garg S, Roy A (2020) In silico analysis of selected alkaloids against main protease (Mpro) of SARS-CoV-2. Chem Biol Interact 332:109309
Hillen HS, Kokic G, Farnung L et al (2020) Structure of replicating SARS-CoV-2 polymerase. Nature 584(7819):154–156
Shang B, Wang XY, Yuan JW et al (2005) Characterization and application of monoclonal antibodies against N protein of SARS-coronavirus. Biochem Biophys Res Commun 336(1):110–117
Hurst KR, Koetzner CA, Masters PS (2009) Identification of in vivo-interacting domains of the murine coronavirus nucleocapsid protein. J Virol 83(14):7221–7234
Saikatendu KS, Joseph JS, Subramanian V et al (2007) Ribonucleocapsid formation of severe acute respiratory syndrome coronavirus through molecular action of the N-terminal domain of N protein. J Virol 81(8):3913–3921
He R, Dobie F, Ballantine M et al (2004) Analysis of multimerization of the SARS coronavirus nucleocapsid protein. Biochem Biophys Res Commun 316(2):476–483
Zeng W, Liu G, Ma H et al (2020) Biochemical characterization of SARS-CoV-2 nucleocapsid protein. Biochem Biophys Res Commun 527(3):618–623
Ahmed SF, Quadeer AA, McKay MR (2020) Preliminary identification of potential vaccine targets for the COVID-19 coronavirus (SARS-CoV-2) based on SARS-CoV immunological studies. Viruses 12(3):254
Guo L, Ren L, Yang S et al (2020) Profiling early humoral response to diagnose novel coronavirus disease (COVID-19). Clin Infect Dis 71(15):778–785
Bianchi M, Benvenuto D, Giovanetti M et al (2020) Sars-CoV-2 envelope and membrane proteins: structural differences linked to virus characteristics? Biomed Res Int 2020:1–6
Thomas S (2020) The structure of the membrane protein of sars-cov-2 resembles the sugar transporter semiSWEET. Pathog Immun 5(1):342–363
Duart G, García-Murria MJ, Grau B et al (2020) SARS-CoV-2 envelope protein topology in eukaryotic membranes. Open Biol 10(9):200209
Millet JK, Whittaker GR (2015) Host cell proteases: Critical determinants of coronavirus tropism and pathogenesis. Virus Res 202:120–134
Hoffmann M, Kleine-Weber H, Pöhlmann S (2020) A multibasic cleavage site in the spike protein of SARS-CoV-2 is essential for infection of human lung cells. Mol Cell 78(4):779–784
Vankadari N (2020) Structural interactions between pandemic SARS-CoV-2 spike glycoprotein and human Furin protease. BioRxiv
Hoffmann M, Kleine-Weber H, Schroeder S et al (2020) SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181(2):271–280
Bestle D, Heindl MR, Limburg H et al (2020) TMPRSS2 and furin are both essential for proteolytic activation of SARS-CoV-2 in human airway cells. Life Sci Alliance 3(9):e202000786
Yamada S, Fukushi S, Kinoshita H et al (2021) Assessment of SARS-CoV-2 infectivity of upper respiratory specimens from COVID-19 patients by virus isolation using VeroE6/TMPRSS2 cells. BMJ Open Respir Res 8(1):e000830
Bittmann S, Luchter E, Moschüring-Alieva E et al (2020) 19: Camostat and the role of serine protease entry inhibitor TMPRSS2. J Regen Biol Med 2(2):1–2
Habtemariam S, Nabavi SF, Ghavami S et al (2020) Possible use of the mucolytic drug, bromhexine hydrochloride, as a prophylactic agent against SARS-CoV-2 infection based on its action on the transmembrane serine protease 2. Pharmacol Res 157:104853
Zhao MM, Yang WL, Yang FY et al (2021) Cathepsin L plays a key role in SARS-CoV-2 infection in humans and humanized mice and is a promising target for new drug development. Signal Transduct Target Ther 6(1):1–12
Zhang J, Ma X, Yu F et al (2020) Teicoplanin potently blocks the cell entry of 2019-nCoV. BioRxiv
Ou X, Liu Y, Lei X et al (2020) Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat Commun 11(1):1–12
Kumar D, Chauhan G, Kalra S et al (2020) A perspective on potential target proteins of COVID-19: comparison with SARS-CoV for designing new small molecules. Bioorg Chem 104:104326
Holshue ML, DeBolt C, Lindquist S et al (2020) First case of 2019 novel coronavirus in the United States. N Engl J Med 382:929–936
Yao X, Ye F, Zhang M et al (2020) In vitro antiviral activity and projection of optimized dosing design of hydroxychloroquine for the treatment of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Clin Infect Dis 71(15):732–739
Grein J, Ohmagari N, Shin D et al (2020) Compassionate use of remdesivir for patients with severe Covid-19. N Engl J Med 382(24):2327–2336
Terali K, Baddal B, Gülcan HO (2020) Prioritizing potential ACE2 inhibitors in the COVID-19 pandemic: insights from a molecular mechanics-assisted structure-based virtual screening experiment. J Mol Graph Model 100:107697
Bhati S (2020) Structure-based drug designing of naphthalene based SARS-CoV PLpro inhibitors for the treatment of COVID-19. Heliyon 6(11):e05558
Beigel JH, Nam HH, Adams PL et al (2019) Advances in respiratory virus therapeutics–A meeting report from the 6th isirv Antiviral Group conference. Antiviral Res 167:45–67
Chen L, Xiong J, Bao L et al (2020) Convalescent plasma as a potential therapy for COVID-19. Lancet Infect Dis 20(4):398–400
Shen C, Wang Z, Zhao F et al (2020) Treatment of 5 critically ill patients with COVID-19 with convalescent plasma. JAMA 323(16):1582–1589
Derebail VK, Falk RJ (2020) ANCA-associated vasculitis: refining therapy with plasma exchange and glucocorticoids. N Engl J Med 382(7):671–673
Hammer Q, Rückert T, Romagnani C (2018) Natural killer cell specificity for viral infections. Nat Immunol 19(8):800–808
Ortiz LA, DuTreil M, Fattman C et al (2007) Interleukin 1 receptor antagonist mediates the antiinflammatory and antifibrotic effect of mesenchymal stem cells during lung injury. Proc Natl Acad Sci 104(26):11002–11007
Gupta N, Su X, Popov B et al (2007) Intrapulmonary delivery of bone marrow-derived mesenchymal stem cells improves survival and attenuates endotoxin-induced acute lung injury in mice. J Immunol 179(3):1855–1863
Matthay MA, Goolaerts A, Howard JP et al (2010) Mesenchymal stem cells for acute lung injury: preclinical evidence. Crit Care Med 38(10 Suppl):S569–S573
Kumamoto M, Nishiwaki T, Matsuo N et al (2009) Minimally cultured bone marrow mesenchymal stem cells ameliorate fibrotic lung injury. Eur Respir J 34(3):740–748
El Agha E, Kramann R, Schneider RK et al (2017) Mesenchymal stem cells in fibrotic disease. Cell Stem Cell 21(2):166–177
Yang ZY, Kong WP, Huang Y et al (2004) A DNA vaccine induces SARS coronavirus neutralization and protective immunity in mice. Nature 428(6982):561–564
Kaul D (2020) An overview of coronaviruses including the SARS-2 coronavirus–Molecular biology, epidemiology and clinical implications. Curr Med Res Pract 10(2):54–64
Wang F, Kream RM, Stefano GB (2020) An evidence based perspective on mRNA-SARS-CoV-2 vaccine development. Med Sci Monit: Int Med J Exp Clin Res 26:e924700–e924701
Le TT, Andreadakis Z, Kumar A et al (2020) The COVID-19 vaccine development landscape. Nat Rev Drug Discov 19(5):305–306
Arya R, Kumari S, Pandey B et al (2021) Structural insights into SARS-CoV-2 proteins. J Mol Biol 433(2):166725
Singh AK, Singh A, Singh R et al (2021) Molnupiravir in COVID-19: a systematic review of literature. Diabetes Metab Syndr 15(6):102329
Fishbane S, Hirsch JS, Nair V (2022) Special considerations for paxlovid treatment among transplant recipients with SARS-CoV-2 infection. Am J Kidney Dis 79(4):480–482
Meini S, Pagotto A, Longo B et al (2020) Role of Lopinavir/Ritonavir in the treatment of Covid-19: a review of current evidence, guideline recommendations, and perspectives. J Clin Med 9(7):2050
Lin C, Li Y, Zhang Y et al (2021) Ceftazidime is a potential drug to inhibit SARS-CoV-2 infection in vitro by blocking spike protein–ACE2 interaction. Signal Transduct Target Ther 6(1):1–4
O’Brien MP, Neto EF, Chen KC et al (2021) Casirivimab with imdevimab antibody cocktail for COVID-19 prevention: interim results. Top Antivir Med 29(1):33–34
Westendorf K, Žentelis S, Wang L et al (2022) LY-CoV1404 (bebtelovimab) potently neutralizes SARS-CoV-2 variants. Cell Rep 39(7):110812
Gottlieb RL, Nirula A, Chen P et al (2021) Effect of bamlanivimab as monotherapy or in combination with etesevimab on viral load in patients with mild to moderate COVID-19: a randomized clinical trial. JAMA 325(7):632–644
Gupta A, Gonzalez-Rojas Y, Juarez E et al (2021) Early treatment for Covid-19 with SARS-CoV-2 neutralizing antibody sotrovimab. N Engl J Med 385(21):1941–1950
Mode D, Stockholm LC AstraZeneca: Evusheld (formerly AZD7442) long-acting antibody combination authorised for emergency use in the US for pre-exposure prophylaxis (prevention) of COVID-19
Funding
The financial support provided by Indian Council of Medical Research (ICMR) to RN is greatly acknowledged.
Author information
Authors and Affiliations
Contributions
Conceptualization and literature search was done by RN. Data analysis and the first draft of the manuscript was written by RN. Reviewing and editing of the manuscript was done by MS.
Corresponding author
Ethics declarations
Competing interest
The authors declare that there are no competing interests.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Noori, R., Sardar, M. An outlook on potential protein targets of COVID-19 as a druggable site. Mol Biol Rep 49, 10729–10748 (2022). https://doi.org/10.1007/s11033-022-07724-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07724-3