Abstract
Accumulating molecular evidence suggests that insulin resistance, rather than SARS-CoV-2- provoked beta-cell impairment, plays a major role in the observed rapid metabolic deterioration in diabetes, or new-onset hyperglycemia, during the COVID-19 clinical course. In order to clarify the underlying complexity of COVID-19 and diabetes mellitus interactions, we propose the imaginary diabetes-COVID-19 molecular tetrahedron with four lateral faces consisting of SARS-CoV-2 entry via ACE2 (lateral face 1), the viral hijacking and replication (lateral face 2), acute inflammatory responses (lateral face 3), and the resulting insulin resistance (lateral face 4). The entrance of SARS-CoV-2 using ACE2 receptor triggers an array of multiple molecular signaling beyond that of the angiotensin II/ACE2-Ang-(1–7) axis, such as down-regulation of PGC-1 α/irisin, increased SREBP-1c activity, upregulation of CD36 and Sirt1 inhibition leading to insulin resistance. In another arm of the molecular cascade, the SARS-CoV-2 hijacking and replication induces a series of molecular events in the host cell metabolic machinery, including upregulation of SREBP-2, decrement in Sirt1 expression, dysregulation in PPAR-ɣ, and LPI resulting in insulin resistance. The COVID-19-diabetes molecular tetrahedron may suggest novel targets for therapeutic interventions to overcome insulin resistance that underlies the pathophysiology of worsening metabolic control in patients with diabetes mellitus or the new-onset of hyperglycemia in COVID-19.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The well-known association of viral infections with metabolic disorders has been further highlighted following the emergence of the novel coronavirus disease 2019 (COVID-19) caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). More recently, evidence has been accumulated to suggest potential cross-talk between diabetes mellitus (DM) and COVID-19 [1]. Diabetes mellitus has been described as a risk factor for severe COVID-19 with an increased risk of mortality [1, 2]. Interestingly, COVID-19 has been associated with new onset diabetes mellitus [3], the precipitation of acute complications of DM, such as diabetic ketoacidosis and hyperosmolar hyperglycemic state [4] and with an increased requirement of insulin in patients with diabetes mellitus [4].
In COVID-19, the entity of “new-onset hyperglycemiaˮ can be classified as “stress- induced hyperglycemia,ˮ “new-onset DMˮ in previously unrecognized pre-diabetes patients, hyperglycemia following pancreatic damage caused by SARS-CoV-2, and drug-induced hyperglycemia or “ secondary diabetes ˮ following use of glucocorticoids [5].
The recruitment of host metabolic pathways by SARS-COV-2 has been suggested for its replication and pathogenesis. This viral interaction with intracellular tissue and organ metabolic processes in conjunction with the adoptive host cellular responses may be responsible for the observed metabolic derangements in COVID-19 and diabetes mellitus [6]. Nowadays, accumulating clinical evidence on COVID-19 demonstrates that, in addition to DM, other components of the metabolic syndrome such as hypertension and obesity could be considered as risk factors for developing severe COVID-19 [7]. These findings may imply that unhealthy metabolic conditions produce a more permissive milieu for the pathogenesis of SARS-CoV-2.
It has been postulated that the COVID-19-induced new onset DM might be mediated by virus-induced damage of β-cells following the viral invasion and entrance through ACE2 receptors [8]. The observed high levels of pancreatic enzymes in COVID-19-induced new onset DM [9] reinforce this hypothesis. Although several lines of evidence suggest that the interaction between ACE2, ADAM17, and TMPRSS2 is pivotal for SARS-CoV-2 related pancreatic β cells injury [10], data on the pancreatic β cells pathologic alterations and the resulting long-term changes remain inconclusive. However, more recent studies showed that the systemic proinflammatory milieu following the COVID-19 cytokine storm may play a major role in insulin resistance, beta cell hyperstimulation and disruption of insulin signaling, eventually leading to SARS-CoV-2- induced DM [11, 12].
Undoubtedly, the omics-derived data will reveal the underlying mysterious molecular mechanisms of the COVID-19 and DM interactions. Recently, the large-scale multi-omics landscape of COVID-19 has clarified 219 biomolecules that have been strongly involved in COVID-19 status and its clinical severity [13]. These underlying metabolic alterations during COVID-19 may be involved in the development of DM or precipitation of its acute metabolic complications.
The mutual interplay between diabetes mellitus and COVID-19 involves a series of overlapping complex networks, each with a degree of redundancy and with alternative pathways which might cross-talk with each other in multiple known and unknown interfaces. The dynamic equilibrium of the network components of underlying hyperglycemia and glycometabolic control could be disturbed by SARS-CoV-2 at multiple sites to emerge an untoward behavior, such as insulin resistance. In order to understand the interactions among the different elements of the complex network of the COVID-19 and diabetes mellitus, we provide a tetrahedron molecular model.
The tetrahedron is one kind of triangular pyramid composed of four triangular faces. The base of this tetrahedron is a triangle, and the lateral faces are also triangles; three triangular faces connecting the base to a common point. In the proposed COVID-19-diabetes mellitus molecular tetrahedron, the molecular pathways for SARS-CoV-2 entry via ACE2 (lateral face 1), the viral hijacking and replication (lateral face 2), acute inflammatory responses (lateral face 3) and the resulting insulin resistance (lateral face 4) constitute the four lateral faces of the tetrahedron. Although the perturbation of glycometabolic complex system is usually initiated via attachment of the SARS-CoV-2 spike protein to its receptor, ACE2, followed by the viral hijacking and replication and pro-inflammatory reactions that create insulin resistance in the host, the numerical sequence of lateral faces of the tetrahedron molecular model is arbitrary here. At the present, it could not be demonstrated which of the molecular pathways of the lateral faces of the tetrahedron may have important role in pathogenesis when compare with the others. All four lateral faces of a tetrahedron are the same size and shape (congruent) and all edges are the same length, all the components (e.g. molecular pathways in the lateral faces) in our proposed COVID-19-diabetes mellitus molecular tetrahedron have the same specific weight in the pathogenesis. Since the lateral faces of a tetrahedron are congruent triangles, the vertex (diabetes mellitus) is directly above the center of the triangular base (insulin resistance) (Fig. 1).
The COVID-19—Diabetes Mellitus molecular tetrahedron: The molecular pathways for SARS-CoV-2 entry via ACE2 (lateral face 1), the viral hijacking and replication (lateral face 2), acute inflammatory responses (lateral face 3) and the resulting insulin resistance (lateral face 4) constitute the four lateral faces of the tetrahedron. Since the lateral faces of a tetrahedron are congruent triangles, the resulting vertex (diabetes mellitus) is directly above the center of the triangular base (insulin resistance)
This review presents multiple metabolic pathways and potential hubs for pathogenesis of COVID-19-induced new-onset diabetes based on the reported transcriptional and proteomic data from COVID-19 patients and based on previous well-known metabolic pathways and mediators involved in the pathogenesis of pancreatic β-cell damage, insulin hyposecretion, and insulin resistance.
Viral entrance (lateral face 1)
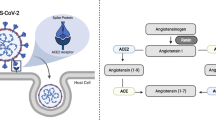
Angiotensin II/ACE2/Ang-(1–7) axis
The attachment of SARS-CoV-2 with ACE2 as its cellular receptor triggers internalization of the complex into the target cells leading to down-regulation of the ACE2 receptor [14]. Cao et al. reported that knockout (ACE2−/Y) mice showed impaired glucose tolerance, reduced first phase secretion of insulin and a significantly inhibited level of hepatic Akt, as one of the pivotal mediators in glucose and lipid metabolism [15].
The ACE2 down-regulation following the viral attachment would potentially result in an increased overall ratio of Ang II to Ang-(1–7). This imbalance results in attenuation of the MasR function [16] leading to hepatic insulin resistance because Cao, et al. showed that the ACE2/ Ang-(1–7)/MasR axis improved hepatic insulin sensitivity via activation of the Akt/PI3K/IRS1/JNK pathway [15] (Fig. 2A). In addition, in an experimental study, ACE2/Ang-(1–7) ameliorated free fatty acid (FFA)-induced oxidative stress and inflammation, leading to decreased levels of reactive oxygen species (ROS) [15], lesser hepatic lipid accumulation [17] and better insulin sensitivity [18]. In an animal model, Ang-1–7 resulted in better insulin sensitivity [19] and improved insulin production in pancreatic β-cells [20].
A The ACE2/ Ang-(1–7)/MasR axis improves insulin sensitivity via activation of the Akt/PI3K/IRS1/JNK pathway. Ang II-induced AT1R activation disrupts insulin-stimulated IRS-1/IRS-2-associated PI3-kinase pathway activities through p38 MAPK, thereby impairing GLUT-4 translocation into membrane of insulin-sensitive tissues. Increased activity of AT1R also has inhibitory effects on insulin receptor signaling and PPARɣ expression through the activation of p38MAPK. B CD36 orchestrates an intracellular metabolic cascade which is involved in metabolism of lipids, leading to overexpression of inflammatory enzymes which inhibit intracellular signaling of insulin receptor. CD36 also induces overexpression of inflammatory cytokines and DPP4 through interaction with TLR2/4. Inflammatory cytokines such as NF-κB, TNF-α, IL6, IL12 have inhibitory effects on insulin signaling. DPP4 inactivates GLP-1/GIP, resulting to disruption of glucose metabolism
The SARS-CoV-2-induced down-regulation of ACE2 and the resulting increased Ang II may be a critical promoter of observed insulin resistance in COVID-19 [11, 12] because increased levels of Ang II has been associated with deterioration of β-cell function [21]. In fact, in vivo studies showed that Ang II disrupts insulin-stimulated IRS-1/IRS-2-associated PI3-kinase pathway activities, thereby impairing GLUT-4 translocation into membrane of insulin-sensitive tissues such as muscle, liver and adipose tissue [22, 23]. Therefore, it can be hypothesized that one of contributing pathologic factor in the observed systemic insulin resistance of COVID-19 may be the disruption of insulin signaling via SARS-CoV-2-induced down-regulation of ACE2 that leads to Ang II hyperactivity, accompanied with insulin signaling impairment.
Taken together, in COVID-19, the increased level of Ang II which is accompanied with an Ang-(1–7) decrement may be accompanied with β-cell dysfunction and insulin hyposecretion, resulting in SARS-COV-2-induced DM.
Dipeptidyl peptidase-4 (DPP4)
MERS-CoV binds to human dipeptidyl peptidase-4 (DPP4/CD26) [24]. The transgenic mice expressing human DPP4 with diet-induced T2D developed a prolonged phase of severe disease and delayed recovery following MERS-CoV infection, indicating a pathological role for DPP4 in the association of type 2 diabetes and MERS-CoV [25]. The docked complex model of COVID-19 spike glycoprotein and DPP4/CD26 suggests that DPP4/CD26 may also act as receptor for SARS-CoV-2 [26]. Hence, DPP4 has been introduced as a potential link between type 2 diabetes and severity of clinical presentations in patients with COVID 19 [27]. Since DPP4 is also a contributing agent in inflammatory and immune reactions, targeting it might be a good modality to control severity of COVID 19. Although, a multicenter, case–control, retrospective, observational study showed that sitagliptin treatment in patients with type 2 diabetes and COVID-19 was associated with reduced mortality and improved clinical prognosis in comparison to standard-of-care treatment [28], contradictory results for the impact of DPP4 on COVID-19 have been reported in some observational clinical studies [29]. Therefore, the DPP4 inhibitors treatment in patients with type 2 diabetes in order to reduce SARS-Co-2 entry and replication should be addressed in an ongoing randomized, placebo-controlled trial.
Viral hijacking and replication (lateral face 2)
Peroxisome proliferator-activator receptor ɣ (PPARɣ)
PPARs are transcription factors belonging to a ligand-activated nuclear hormone (NR) receptor superfamily including three main subtypes [30]. PPARɣ, as one member of the PPAR family, has pleotropic functions on lipid and glucose metabolism [31] through eliciting the glucose transporter4 (Glut4) and c-Cbl-associated protein (CAP) transcription [32]. PPARɣ exerts its anti-inflammatory properties through modulation of M2 macrophages with inhibition of inflammatory cytokine bursts [33], and inactivation of NF-κB via silencing pro-inflammatory genes and p65 [34]. It has been well known that PPARɣ is down-regulated during influenza A infection [33]. Furthermore, a recent study on biopsied lung tissue from COVID-19 patients revealed a significant dysregulation of genetic programs in PPARɣ-dependent pathways [35]. Hence, dysregulated PPARɣ-dependent unbalanced inflammatory responses and disturbed post-receptor signaling of PPARɣ might induce insulin resistance during SARS-CoV-2 infection.
Pioglitazone is a potent PPAR-γ agonist that reduces chronic inflammation in T2DM patients, which is manifested in a decrease in the secretion of various pro-inflammatory cytokines, including TNF-α, IL-1, and IL-6, and other inflammatory markers. Therefore, it has been hypothesized that administering pioglitazone to T2DM patients who have SARSCoV-2-driven hyperinflammation and cytokine storm syndrome will improve their clinical prognosis [36]. Currently, two randomized clinical trial have been designed to investigate the effects of pioglitazone on type 2 diabetes mellitus with COVID-19 (https://clinicaltrials.gov/ct2/show/NCT04604223) and https://clinicaltrials.gov/ct2/show/NCT04535700).
Lysophosphatidylinositol (LPI)
Lysophosphatidylinositol (LPI) has been introduced as an active molecule involved in the regulation of glucose homeostasis and insulin signaling [37]. It triggers insulin release from pancreatic β-cells via activation of its G protein-coupled receptor 55 (GPR55) [38] and subsequent intracellular Ca2+ mobilization with an increased intracellular level of cAMP [39]. GPR55 is highly expressed in pancreatic islets cells. A disruption in the activity of LPI has been reported in the recovery phase of SARS-CoV [37]. Therefore, the LPI activity during COVID-19 infection should be investigated to reveal the involvement of LPI dysfunction in the pathogenesis of insulin resistance and hyposecretion of insulin during COVID-19.
Sterol response element-binding protein 1 (SREBP-1)
SARS-CoV-2 relies on the host lipid machinery for membrane fusion, envelopment, and transformation for viral replication, endocytosis, and exocytosis [40]. Although the mechanisms and induced perturbation of lipid complex pathways mediated by SARS-CoV-2 infection has yet to be revealed, targeting lipid metabolism in COVID-19 to interfere with the viral replication has been recently noticed.
Sterol regulatory element binding protein (SREBP) may be a potential target for the modulation of cellular lipid metabolism in COVID-19 [40, 41] as an antiviral therapy. The SREBP family consists of three transcription factors: SREBP-1a, SREBP-1c, and SREBP-2. SREBP-1a and SREBP-1c are mainly involved in the activation of transcription for fatty acid and triglyceride synthesis, whereas SREBP-2 mediates the transcription of genes in cholesterol biosynthetic pathways. Sterol depletion activates SREBP-2 expression, while insulin glucose and fatty acids control SREBP-1c expression [42].
SARS-CoV-2-mediated ACE2 decrement may lead to an increase in SREBP-1c activity. In fact, increased SREBP-1c protein has been observed in ACE2 knockdown mice [15]. The dysregulated lipid metabolism following the entrance of the virus with an increase in SREBP also has been suggested [43]. Shimomura et al., who produced aP2-SREBP-1c transgenic mice overexpressing SREBP-1c in adipose tissue, showed marked insulin resistance with 60-fold elevation in the plasma levels of insulin. These aP2-SREBP-1c mice were markedly resistant to the injected insulin in order to decrease elevated blood glucose (300 mg/dl) [44]. Therefore, the observed insulin resistance and new onset of DM following COVID-19 may be ascribed by dysregulation in lipid metabolism which is accompanied by SREPB-1c overexpression following the virus entrance.
The SREBP-1c overexpression with resulting elevated lipogenesis and hyperinsulinemia may aggravate insulin resistance via down-regulation of the mRNA for insulin receptor protein 2 (IRS2) [45]. Thus, SARS-CoV-2-induced SREBP-1c overexpression may accentuate insulin resistance through down-regulation of IRS2, a pivotal component of the insulin-signaling pathway. Previously, it has been reported that SREBPs interfere with the effect of the insulin response element on the IRS2 promoter leading to suppression of the down-stream insulin signaling pathway [45]. The possible effect of SARS-CoV-2-mediated SREBP overexpression on other components of the insulin signaling cascade, including the mammalian target of rapamycin complex 1 (mTORC1), should be investigated because the mTOR is a major effector of PI3K/Akt signaling in the insulin-dependent regulation of SREBP-1c [46].
Interestingly, SARS-CoV-2-induced SREBP-2 up-regulation was reported in the peripheral blood mononuclear cells of infected patients. This up-regulation was accompanied with inflammatory responses and the observed cytokine storm in patients with COVID-19 [47]. The involvement of COVID-19-activated SREBP-2 in insulin resistance via inflammatory complex pathways warrants more investigations.
Fatty acid translocase (CD36)
Fatty acid translocase, so-called CD36, is a member of the class B scavenger receptor family, and a multifunctional transmembrane glycoprotein receptor which has attracted attention for its involvement in DM and its complications [48]. It is highly expressed in fat tissue and plays as a co-receptor for the toll-like receptor 2 (TLR2) ligand that is essential for some TLR2/6 responses which are evoked by pathogens, leading to a classic TLR signaling cascade and the subsequent production of TNF-α, IL-6, and IL-12 [49] (Fig. 2B). In obesity it is proposed that this molecule mediates a cross-talk between adipocytes and macrophages [50]. The obesity-induced CD36 up-regulation is closely related to chronic inflammation which might induce insulin resistance [48].
In fact, CD36 knockout mice had better insulin sensitivity profiles and lower pro-inflammatory markers such as IFN-ɣ, MCP-1, and TNF-α in response to oxidized LDL (ox LDL) in adipose tissue [51]. Goudrian et al. demonstrated that CD36−/− mice had better whole-body insulin-mediated glucose uptake and better insulin sensitivity in muscle but more hepatic insulin resistance [52].
The activation of TLR4 and CD36-dependent pathways results in up-regulation of dipeptidyl peptidase 4 (DPP4) [53]. It has been revealed that DPP4, as a trans-membrane glycoprotein with catalytic activity, interacts with other ligands, such as the MERS-CoV spike protein [54] and its activity has been associated with the degree of insulin resistance [53]. TLR4, as an important component of the immune system, has direct interaction with CD36 in macrophages and monocytes [53]. The overactivity of TLR4 signaling inflammatory pathways in circulating mononuclear cells of patients with COVID-19 has been shown [55]. Therefore, it could be postulated that TLR4-induced up-regulation of CD36 may result in the increased activity of DPP4, which may subsequently lead to abnormal insulin secretion and insulin resistance in COVID-19.
An animal study demonstrated that ACE2 knockout mice had increased expression of CD36 in the liver [56]. Additionally, angiotensin II resulted in indirect increased expression of CD36 through down-regulation of sirtuin 1 (which negatively regulates CD36) [56]. Thus, SARS-CoV-2 induced ACE2 attenuation may also result in increased levels of CD36 that might switch on inflammatory processes, leading to insulin resistance. In a compelling and very recent in vitro study, Gomez Diaz et al. reported that SRAS-CoV-2 infection modulates lipid metabolism via CD36 pathways in monocytes. The observed CD36 overexpression in monocytes suggests an increment of lipid uptake and hijacking of the host LDLs to enhance the viral replication [57].
Taken together, the observed ACE2/Ang II imbalance and decreased level of sirt1 may result in overexpression of CD36 leading to inflammatory states accompanied with insulin resistance.
Sirtuin1
The ancient protein sirtuin1 (Sirt1), as one of the most conserved mammalian nicotinamide adenosine dinucleotide (NAD+)-dependent histone deacetylases, has significant roles in metabolic pathways, in the protection of organism against oxidative stress, and the promotion of DNA stability and attenuation of age-related disorders such as neurodegenerative diseases and cancers [58,59,60]. This protein modulates intracellular metabolic pathways by targeting cellular proteins such as NF-κB and peroxisome proliferator-activated receptor ɣ (PPARɣ) [61].
The insulin signaling and PI3K/Akt pathway activation are positively regulated by sirt1 (Fig. 3A). Yoshizaki et al. reported that sirt1 knockdown in adipocyte resulted in inhibition of insulin-stimulated glucose uptake and increased serine phosphorylation of insulin receptor substrate1 (IRS1) [62]. The modulation of sirt1 on insulin-induced tyrosine phosphorylation of IRS2 was also reported [63]. The sirt1-dependent Akt activation signaling has been shown to be pivotal in promoting PI3 binding and activation [64]. Lower expression of sirt1 disrupts insulin secretion [65] through an increased expression of uncoupling protein 2 (UCP2) in pancreatic β-cells [66]. Taken together, sirt1 exerts its effect on energy metabolism and insulin resistance through a variety of molecular mechanisms [67].
A The MasR-induced expression of sirtuin1 results in increased expression of PPARɣ and enhanced intracellular signaling of insulin receptor through Akt activation. Sirtuin1 has an inhibitory effect on SREBP-1c expression through negative regulation of P53. In β-cells, sirtuin 1 inhibits UCP2 in which prevents the release of insulin. SREBP1 interferes with the effect of the insulin response element on the IRS2 promoter leading to suppression of the down-stream insulin signaling pathway. B In skeletal muscles, irisin is derived from FNDC5 under the stimulatory effect of MasR through the PGC1-α activation. It leads to overexpression of GLUT4 through activation of p38MAPK in insulin-sensitive tissues. Irisin also induces production of betatrophin which induces proliferation of β-cells. During COVID-19, the possible alteration of irisin production may induce inflammatory pathway through the activation of p-JNK and p-ERK. This will lead to overproduction of NF-κB, TNF-α, IL6, IL1β and MCP1 and disturbed insulin signaling
The sirtuins machinery system is used by the host cells as a defense mechanism against DNA and RNA viral pathogens. Therefore, sirtuins have been defined as critical regulatory hubs in a variety of metabolic and regulatory processes, such as fatty acid oxidation and tricarboxylic acid cycle metabolism, which are affected during viral infections [68, 69]. Hence, SARS-CoV-2 may struggle for control over sirtuin functions to promote its replication and protein assemblage. Interestingly, Bordoni et al. demonstrated a significant decrease in the sirt1 expression of peripheral blood mononuclear cells of COVID-19 patients resulting in the maintenance of p53 active form and a dysregulated p53/sirt1 axis and a provoked inflammatory stat [70]. The observed repressed sirt1 in COVID-19 may be involved in insulin resistance and metabolic derangement.
However, COVID-19 mediated ACE2 depletion, and the resulting Ang-(1–7)/AngII imbalance which results in sirt1 inhibition in liver and up-regulation of CD36 [56], leading to increased insulin resistance, should also be considered.
Irisin
Irisin, which is derived from fibronectin type III domain-containing protein 5 (FNDC5) [71], is highly produced by skeletal muscle in response to exercise [72] (Fig. 3B). This dependent PPARɣ co-activator 1-α (PGC1-α) myokine has a pivotal role in the induction of subcutaneous fat browning through increased expression of uncoupling protein 1 (UCP1), leading to higher energy expenditure, higher weight loss, and improvement of glucose metabolism [73,74,75]. Irisin reduces the expression of TLR4 protein and subsequently reduces the activation of NF-κB through down-regulation of the MAPK signaling pathway that can subsequently improve insulin sensitivity. Additionally, this molecule reduces the phosphorylation of p-JNK and p-ERK, leading to a reduction of crucial inflammatory cytokines such as TNF-α, MCP-1, IL-6, and IL-1β that are involved in the pathogenesis of inflammation-induced insulin resistance [76]. Irisin induces the expression of UCP1 and betatrophin through the activation of MAPK pathways (P38 and ERK) [77]. The new hormone betatrophin resulted in regeneration of pancreatic β-cells and an increase in the mass of pancreatic β-cells of mice [72]. Furthermore, the ROS-mediated activation of AMPK which is caused by irisin results in more glucose uptake through higher expression of GLUT4 in muscles [78]. As a result, irisin may ameliorate insulin resistance by targeting peripheral muscles through a different metabolic pathway [72].
Bernardi et al. demonstrated that ACE2 knockout mice had significantly decreased expression of the PGC1-α protein [79]. In agreement with this finding, Cao et al. also reported down-regulation of PGC-1α in the liver of ACE2 knockdown mice [15]. These findings may support the potential involvement of ACE2/Ang-(1–7)/Mas axis in irisin production via PGC-1α activity. Therefore, it could be hypothesized that SARS-CoV-2-mediated ACE2 down-regulation, which is accompanied with down-regulation of PGC-1α, would result in the lower production of irisin in COVID-19, leading to insulin resistance through multiple metabolic pathways. Curiously, transcriptional analysis revealed that irisin down-regulates the genes that are involved in the pathogenesis of severe forms of COVID-19, such as FURIN and ADAM10, which are also involved in virus replication [80].
Inflammation (lateral face 3) and insulin resistance (lateral face 4)
From the aforementioned molecular mechanisms relating COVID-19 and diabetes mellitus or hyperglycemia, it appears that the common root of new-onset hyperglycemia or its worsening in preexisting diabetes mellitus during multiple clinical phases of COVID-19 could be SARS-CoV-2-induced insulin resistance. In fact, the entrance of SARS-CoV-2 to host cells using ACE2 or DPP4 as its entry receptors and the viral hijacking-replication mechanisms trigger insulin resistance via perturbation in insulin signaling primarily in the IRS/PI3K/AKT/mTORC2 axis through multiple diverse pathways. Therefore, the observed SARS-CoV-2–induced hyperglycemia or ketoacidosis in ketosis-prone diabetes and the association of diabetes mellitus/obesity with severe COVID-19 may be primarily described by the appearance or worsening of insulin resistance during this catastrophic viral infection [1, 81]. However, the modulation of immune reactions, provocation of inflammatory responses, and the cytokine storm of COVID-19 should also be considered in the aggravation of insulin resistance and dysregulation of glucose metabolism in patients with severe COVID-19 [82].
In an Italian cohort study with hospitalized COVID-19 patients without pre-existing history of diabetes mellitus, presence of new-onset hyperglycemia, insulin resistance, abnormal cytokine profile, beta cell hyperstimulation including higher insulin, proinsulin, C-peptide levels as compared to the controls were reported during acute and post recovery phases of COVID-19 [11]. These findings demonstrate that SARS-CoV-2 induces an insulin resistance state per se leading to aberrant glycometabolic control, which can persist in the patients who recovered from COVID-19. In another 6-month prospective study, it was shown that SARS-CoV-2 induced the risk of insulin resistance in patients without prior history of diabetes, which persisted at both 3-month and 6-month post COVID-19 phases [12]. Therefore, SARS-CoV-2 may induce and aggravate an insulin resistance state based on a proinflammatory milieu, resembling that which could be seen in patients with type 2 diabetes mellitus.
Ang II via Ang II type 1A receptor (AT1R) signalling contributes to a powerful inflammatory response. The up-regulation of AT1R through Ang II elicits inflammation by both immune cells and tissue-resident cells. This interaction creates an enhanced vascular permeability that is associated with production of reactive oxygen species (ROS), proinflammatory cytokines and chemokines including prostaglandins, VEGF, NFκB, TNFα, IL-1β, IL-6, and IFNγ [83]. In fact, initial plasma concentrations of IL1B, IL1RA, IL7, IL8, IL9, IL10,, IFNγ, TNFα, interferon gamma-induced protein 10 (IP10), monocyte chemoattractant protein 1 (MCP1), macrophage inflammatory protein 1 α (MIP1A), MIP1B, and VEGF were higher in patients with COVID-19 than in the healthy controls [2].
Furthermore, immune cell recruitment [84] and overexpression of TLR4, TLR2, CD40, and matrix metalloproteinase 9 (MMP9) provoke a hyperinflammatory state [85, 86], leading to the cytokine storm. Thus COVID-19 can be classified as an inflammatory condition in nature. It must be considered that the end product of almost all aforementioned pathways is over-activity of the inflammatory state. Inflammation can induce insulin resistance and β-cell dysfunction through different mechanisms which are mediated by inflammatory mediators such as IL-1, IL-6, TNFα, INF-ɣ, and MCP1.
Pro-inflammatory cytokines block the insulin signaling receptors in β-cells [87]. IL-1β as master pro-inflammatory mediator down-regulates IRS-1 through ERK-dependent and ERK-independent mechanisms in post-transcriptional levels leading to insulin resistance [88].
The mechanisms which are involved in the pathogenesis of IL-6 mediated insulin resistance are versatile and consist of the prevention of non-oxidative glucose metabolism [89], suppression of lipoprotein lipase that increases triglycerides [90], and activation of SOCS [91] that is opposing the action of insulin [92].
TNF-α phosphorylates serine 307 in IRS-1 and induces insulin resistance through activation of NF-κB and Jun NH2-terminal kinase (JNK) [87]. Overproduction of MCP-1, which resulted in an increased proliferation of macrophages in adipose tissue [93], in combination with an increased production of CCR5 [94], may induce insulin resistance. Oxidative stress caused by inflammation over-activates stress signaling pathways, such as JNK and NF-κB [87], and promotes secretion of TNF-α and IL-6—the condition that interferes with glucose uptake in peripheral tissues.
Thus, it can be concluded that inflammation may be the beating heart for the dysregulation of glucose metabolism and hyperglycemia which is occurring during COVID-19.
The imbalance of other potent cytokines which modulate low grade chronic inflammation in adipose tissue may contribute to SARS-CoV-2-provoked insulin resistance. In a very recent study, Reiterer et al. showed lower levels of adiponectin as a protective adipocytokine against diabetes in hyperglycemic COVID-19 patients [95]. They also suggested adipose tissue dysfunction and resulting insulin resistance as a feature of SARS-CoV-2 infection [95]. The complex interactions among preexisting chronic low grade inflammation and alterations in ACE2 [80], PPAR-ɣ [34], SREBP [96], CD36 [97], sirtuin [98], irisin [99] in obesity, and the dysregulation in secreted adipocytokines from adipose tissue in SARS-CoV-2 infection need further investigations in severe COVID-19.
The next step would be to determine whether other molecular mechanisms underlying insulin resistance, such as integrated stress response (ISR), free radical overload, and mitochondrial dysfunction, could be involved in SARS-CoV-2-induced hyperglycemia [81]. The involvement of ISR pathways in the down-regulation of insulin signaling via serine phosphorylation of insulin receptor substrates by fragments of viral RNA has been suggested in COVID-19 [81].
Conclusions
Experimental, clinical, and epidemiologic evidence has accrued for a bidirectional relationship between diabetes mellitus and COVID-19. On one hand, preexisting diabetes is accompanied with COVID-19 severity, and on the other, worsening metabolic control in patients with diabetes leading to diabetic ketoacidosis, hyperglycemic hyperosmolar syndrome, or new-onset of hyperglycemia have been reported in COVID-19 [1].
The results of recently published studies that have been reflected in the COVID-19-diabetes mellitus molecular tetrahedron provide support for the hypothesis of a potential diabetogenic effect of COVID-19, beyond stress response associated with severe illness, pre-existing undiagnosed diabetes, steroid-induced hyperglycemia, and SARS-CoV-2-induced β-cell impairment [100]. Thus, it is plausible that COVID-19 could create a perturbation in the glycometabolic complex system with resulting hyperglycemia and insulin resistance that can not only complicate the pathophysiology of pre-existing diabetes but also may lead to new-onset diabetes. Very recent studies showed that aberrant glycometabolic control and aggravated insulin resistant in COVID-19 patients without pre-existing diabetes persisted for at least 6 months after the infection [11, 12]. Whether elevated glucose concentrations (in a non-diabetes range) or new-onset diabetes, is likely to remain permanent is not known at the present time.
The more recent molecular evidence suggests that insulin resistance, rather than SARS-CoV-2-provoked beta-cell impairment, plays a major role in the observed rapid metabolic deterioration in diabetes or the new-onset hyperglycemia during the COVID-19 clinical course.
The entrance of SARS-CoV-2 using ACE2 receptor triggers an array of multiple molecular signaling beyond that of the angiotensin II/ACE2-Ang-(1–7) axis such as down-regulation of PGC-1 α/irisin, increased SREBP-1c activity, upregulation of CD36 and Sirt1 inhibition leading to insulin resistance. In another arm of the molecular cascade, the SARS-CoV-2 hijacking and replication induces a series of molecular events in the host cell metabolic machinery including upregulation of SREBP-2, decrement in Sirt1 expression, dysregulation in PPAR-ɣ, and LPI resulting in insulin resistance.
Taken together, it could be argued that insulin resistance is the predominant common cascade for a diverse cytokine, molecular signaling pathways, and SARS-CoV-2-induced inflammatory responses that mediate the diabetes-COVID-19 relationship. The complexity of these molecular interactions that are triggered by SARS-CoV-2 entry via ACE2 (lateral face 1), the viral hijacking and replication (lateral face 2), and inflammatory responses (lateral face 3), leading to insulin resistance (lateral face 4), can be imagine as a mysterious tetrahedron with ACE2 at its center, which we call the COVID-19-diabetes molecular tetrahedron (Fig. 4).
This imaginary molecular tetrahedron presents an array of intertwined molecular interactions at its four lateral faces. Clarification of hierarchy of the complex network components across different organizational levels may expand our armamentarium of treatments to tackle SARS-CoV-2 via different therapeutic strategies to overcome insulin resistance underlying the pathophysiology of worsening metabolic control in patients with diabetes mellitus or new-onset hyperglycemia seen in COVID-19.
Data availability
Not applicable.
Abbreviations
- COVID-19:
-
Coronavirus disease 2019
- SARS-CoV-2:
-
Severe acute respiratory syndrome coronavirus 2
- DM:
-
Diabetes mellitus
- SREBP:
-
Sterol regulatory element binding protein
- SOCS 3:
-
The suppressor of cytokine signaling 3
- FFA:
-
Free fatty acid
- ROS:
-
Reactive oxygen species
- AMPK:
-
AMP-activated protein kinase
- P38MAPK:
-
Phosphorylation of p38 mitogen-activated protein kinase
- PPARɣ:
-
Peroxisome proliferator-activator receptor ɣ
- NR:
-
Nuclear hormone
- Glut4:
-
Glucose transporter4
- CAP:
-
C-Cbl-associated protein
- LPI:
-
Lysophosphatidylinositol
- GPR55:
-
G protein-coupled receptor 55
- IRS:
-
Insulin receptor protein
- mTORC1:
-
Mammalian target of rapamycin complex 1
- TLR:
-
Toll-like receptor
- DPP4:
-
Dipeptidyl peptidase 4
- NAD+ :
-
Nicotinamide adenosine dinucleotide
- UCP2:
-
Uncoupling protein 2
- FNDC5:
-
Fibronectin type III domain-containing protein 5
- VEGF:
-
Vascular endothelial growth factor
- MMP9:
-
Matrix metalloproteinase 9
- JNK:
-
Jun NH2-terminal kinase
- ISR:
-
Integrated stress response
- LPA:
-
Lysophosphatidic acid
- DAG:
-
Diacylglycerol
- GLP-1:
-
Glucagon-like peptide-1
- FNDC5:
-
Fibronectin type III domain containing 5
- Ag:
-
Angiotensin
- ACE:
-
Angiotensin converting enzyme
- IRS1:
-
Insulin receptor substrate
- PI3K:
-
Phosphoinositide 3-kinase
- PDK1:
-
Phosphoinositide-dependent kinase 1
- RGC:
-
Ral-GAP complex
- AS160:
-
Akt substrate of 160 kDa
- MCP1:
-
Monocyte chemoattractant protein-1
- TNF-α:
-
Tumor necrosis factor α
- IL-6:
-
Interleukin 6
- NF-κB:
-
Nuclear factor kappa B
- PGC1:
-
Peroxisome proliferator-activated receptor γ coactivator 1
References
Rubino F, Amiel SA, Zimmet P, Alberti G, Bornstein S, Eckel RH, Mingrone G, Boehm B, Cooper ME, Chai Z (2020) New-onset diabetes in Covid-19. N Engl J Med 383:789–790
Zhou F, Yu T, Du R, Fan G, Liu Y, Liu Z, Xiang J, Wang Y, Song B, Gu X (2020) Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet 395:1054–1062
Li J, Wang X, Chen J, Zuo X, Zhang H, Deng A (2020) COVID-19 infection may cause ketosis and ketoacidosis. Diabetes Obes Metab 22:1935–1941
Jeon JY (2020) Acute hyperglycemic crises with coronavirus disease-19: case reports. Diabetes Metab J 44:349–353
Singh AK, Singh R (2020) Hyperglycemia without diabetes and new-onset diabetes are both associated with poorer outcomes in COVID-19. Diabetes Res Clin Pract 167:108382
Ayres JS (2020) Immunometabolism of infections. Nat Rev Immunol 20:79–80
Bornstein SR, Dalan R, Hopkins D, Mingrone G, Boehm BO (2020) Endocrine and metabolic link to coronavirus infection. Nat Rev Endocrinol 16:297–298
Yang J-K, Lin S-S, Ji X-J, Guo L-M (2010) Binding of SARS coronavirus to its receptor damages islets and causes acute diabetes. Acta Diabetol 47:193–199
Ashok A, Faghih M, Singh VK (2020) Mild pancreatic enzyme elevations in COVID-19 pneumonia: synonymous with injury or noise? Gastroenterology 160:1872
El-Huneidi W, Hamad M, Taneera J (2021) Expression of SARS-CoV-2 receptor “ACE2” in human pancreatic β cells: to be or not to be! Islets 13:106–114
Montefusco L, Nasr MB, D’Addio F, Loretelli C, Rossi A, Pastore I, Daniele G, Abdelsalam A, Maestroni A, Dell’Acqua M (2021) Acute and long-term disruption of glycometabolic control after SARS-CoV-2 infection. Nat Metab 2021:1–12
Chen M, Zhu B, Chen D, Hu X, Xu X, Shen W-J, Hu C, Li J, Qu S (2021) COVID-19 may increase the risk of insulin resistance in adult patients without diabetes: a 6-month prospective study. Endocr Pract 27:834
Overmyer KA, Shishkova E, Miller IJ, Balnis J, Bernstein MN, Peters-Clarke TM, Meyer JG, Quan Q, Muehlbauer LK, Trujillo EA (2021) Large-scale multi-omic analysis of COVID-19 severity. Cell Syst 12:23-407.e7
Hoffmann M, Kleine-Weber H, Schroeder S, Krüger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu N-H, Nitsche A (2020) SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181:271-280.e8
Cao X, Yang F, Shi T, Yuan M, Xin Z, Xie R, Li S, Li H, Yang J-K (2016) Angiotensin-converting enzyme 2/angiotensin-(1–7)/Mas axis activates Akt signaling to ameliorate hepatic steatosis. Sci Rep 6:1–11
Shete A (2020) Urgent need for evaluating agonists of angiotensin-(1–7)/Mas receptor axis for treating patients with COVID-19. Int J Infect Dis 96:348–351
Wei Y, Clark SE, Morris EM, Thyfault JP, Uptergrove GM, Whaley-Connell AT, Ferrario CM, Sowers JR, Ibdah JA (2008) Angiotensin II-induced non-alcoholic fatty liver disease is mediated by oxidative stress in transgenic TG (mRen2) 27 (Ren2) rats. J Hepatol 49:417–428
Fromenty B, Robin M, Igoudjil A, Mansouri A, Pessayre D (2004) The ins and outs of mitochondrial dysfunction in NASH. Diabetes Metab 30:121–138
Giani JF, Mayer MA, Muñoz MC, Silberman EA, Hocht C, Taira CA, Gironacci MM, Turyn D, Dominici FP (2009) Chronic infusion of angiotensin-(1–7) improves insulin resistance and hypertension induced by a high-fructose diet in rats. Am J Physiol-Endocrinol Metab 296:E262–E271
He J, Yang Z, Yang H, Wang L, Wu H, Fan Y, Wang W, Fan X, Li X (2015) Regulation of insulin sensitivity, insulin production, and pancreatic β cell survival by angiotensin-(1–7) in a rat model of streptozotocin-induced diabetes mellitus. Peptides 64:49–54
Yuan L, Li X, Xu G-L, Qi C-J (2010) Effects of renin-angiotensin system blockade on islet function in diabetic rats. J Endocrinol Investig 33:13–19
Folli F, Saad M, Velloso L, Hansen H, Carandente O, Feener E, Kahn C (1999) Crosstalk between insulin and angiotensin II signalling systems. Exp Clin Endocrinol Diabetes 107:133–139
Olivares-Reyes JA, Arellano-Plancarte A, Castillo-Hernandez JR (2009) Angiotensin II and the development of insulin resistance: implications for diabetes. Mol Cell Endocrinol 302:128–139
Raj VS, Mou H, Smits SL, Dekkers DH, Müller MA, Dijkman R, Muth D, Demmers JA, Zaki A, Fouchier RA (2013) Dipeptidyl peptidase 4 is a functional receptor for the emerging human coronavirus-EMC. Nature 495:251–254
Kulcsar KA, Coleman CM, Beck SE, Frieman MB (2019) Comorbid diabetes results in immune dysregulation and enhanced disease severity following MERS-CoV infection. J CI insight 4:e131774
Vankadari N, Wilce JA (2020) Emerging COVID-19 coronavirus: glycan shield and structure prediction of spike glycoprotein and its interaction with human CD26. Emerg Microbes Infect 9:601–604
Valencia I, Peiró C, Lorenzo Ó, Sánchez-Ferrer CF, Eckel J, Romacho T (2020) DPP4 and ACE2 in diabetes and COVID-19: therapeutic targets for cardiovascular complications? Front Pharmacol 11:1161
Solerte SB, D’Addio F, Trevisan R, Lovati E, Rossi A, Pastore I, Dell’Acqua M, Ippolito E, Scaranna C, Bellante R (2020) Sitagliptin treatment at the time of hospitalization was associated with reduced mortality in patients with type 2 diabetes and COVID-19: a multicenter, case-control, retrospective, observational study. Diabetes Care 43:2999–3006
Sun B, Huang S, Zhou J (2021) Perspectives of antidiabetic drugs in diabetes with coronavirus infections. Front Pharmacol 11:2424
Berger J, Moller DE (2002) The mechanisms of action of PPARs. Annu Rev Med 53:409–435
Grygiel-Górniak B (2014) Peroxisome proliferator-activated receptors and their ligands: nutritional and clinical implications-a review. Nutr J 13:1–10
Wu Z, Xie Y, Morrison RF, Bucher N, Farmer SR (1998) PPARgamma induces the insulin-dependent glucose transporter GLUT4 in the absence of C/EBPalpha during the conversion of 3T3 fibroblasts into adipocytes. J Clin Investig 101:22–32
Ayres JS (2020) A metabolic handbook for the COVID-19 pandemic. Nat Metab 2:572–585
Hou Y, Moreau F, Chadee K (2012) PPARγ is an E3 ligase that induces the degradation of NFκB/p65. Nat Commun 3:1300. https://doi.org/10.1038/ncomms2270
Desterke C, Turhan AG, Bennaceur-Griscelli A, Griscelli F (2020) PPARγ cistrome repression during activation of lung monocyte-macrophages in severe COVID-19. Iscience 23:101611
Subir R (2020) Use of pioglitazone in people with type 2 diabetes mellitus with coronavirus disease 2019 (COVID-19): boon or bane? Diabetes Metab Syndr 14:829–831
Wu Q, Zhou L, Sun X, Yan Z, Hu C, Wu J, Xu L, Li X, Liu H, Yin P (2017) Altered lipid metabolism in recovered SARS patients twelve years after infection. Sci Rep 7:1–12
Arifin SA, Falasca M (2016) Lysophosphatidylinositol signalling and metabolic diseases. Metabolites 6:6
Metz SA (1988) Mobilization of cellular Ca2+ by lysophospholipids in rat islets of Langerhans. Biochim Et Biophys Acta (BBA) 968:239–252
Abu-Farha M, Thanaraj TA, Qaddoumi MG, Hashem A, Abubaker J, Al-Mulla F (2020) The role of lipid metabolism in COVID-19 virus infection and as a drug target. Int J Mol Sci 21:3544
Yuan S, Chu H, Chan JF-W, Ye Z-W, Wen L, Yan B, Lai P-M, Tee K-M, Huang J, Chen D (2019) SREBP-dependent lipidomic reprogramming as a broad-spectrum antiviral target. Nat Commun 10:1–15
DeBose-Boyd RA, Ye J (2018) SREBPs in lipid metabolism, insulin signaling, and beyond. Trends Biochem Sci 43:358–368
Al Heialy S, Hachim MY, Senok A, Gaudet M, Abou Tayoun A, Hamoudi R, Alsheikh-Ali A, Hamid Q (2020) Regulation of angiotensin-converting enzyme 2 in obesity: implications for COVID-19. Front Physiol 11:555039
Shimomura I, Hammer RE, Richardson JA, Ikemoto S, Bashmakov Y, Goldstein JL, Brown MS (1998) Insulin resistance and diabetes mellitus in transgenic mice expressing nuclear SREBP-1c in adipose tissue: model for congenital generalized lipodystrophy. Genes Dev 12:3182–3194
Ide T, Shimano H, Yahagi N, Matsuzaka T, Nakakuki M, Yamamoto T, Nakagawa Y, Takahashi A, Suzuki H, Sone H (2004) SREBPs suppress IRS-2-mediated insulin signalling in the liver. Nat Cell Biol 6:351–357
Matsuzaka T, Shimano H (2013) Insulin-dependent and-independent regulation of sterol regulatory element-binding protein-1c. J Diabetes Investig 4:411
Lee W, Ahn JH, Park HH, Kim HN, Kim H, Yoo Y, Shin H, Hong KS, Jang JG, Park CG (2020) COVID-19-activated SREBP2 disturbs cholesterol biosynthesis and leads to cytokine storm. Signal transduct Target Ther 5:1–11
Puchałowicz K, Rać ME (2020) The multifunctionality of CD36 in Diabetes mellitus and its complications: update in pathogenesis, treatment and monitoring. Cells 9:1877
Febbraio M, Silverstein RL (2007) CD36: implications in cardiovascular disease. Int J Biochem Cell Biol 39:2012–2030
Kennedy DJ, Kuchibhotla S, Westfall KM, Silverstein RL, Morton RE, Febbraio M (2011) A CD36-dependent pathway enhances macrophage and adipose tissue inflammation and impairs insulin signalling. Cardiovasc Res 89:604–613
Vallerie SN, Hotamisligil GS (2010) The role of JNK proteins in metabolism. Sci Transl Med 2:605
Goudriaan JR, Dahlmans VE, Teusink B, Ouwens DM, Febbraio M, Maassen JA, Romijn JA, Havekes LM, Voshol PJ (2003) CD36 deficiency increases insulin sensitivity in muscle, but induces insulin resistance in the liver in mice. J Lipid Res 44:2270–2277
Rao X, Zhao S, Braunstein Z, Mao H, Razavi M, Duan L, Wei Y, Toomey AC, Rajagopalan S, Zhong J (2019) Oxidized LDL upregulates macrophage DPP4 expression via TLR4/TRIF/CD36 pathways. EBioMedicine 41:50–61
Lu G, Hu Y, Wang Q, Qi J, Gao F, Li Y, Zhang Y, Zhang W, Yuan Y, Bao J (2013) Molecular basis of binding between novel human coronavirus MERS-CoV and its receptor CD26. Nature 500:227–231
Sohn KM, Lee S-G, Kim HJ, Cheon S, Jeong H, Lee J, Kim IS, Silwal P, Kim YJ, Paik S (2020) COVID-19 patients upregulate toll-like receptor 4-mediated inflammatory signaling that mimics bacterial sepsis. J Korean Med Sci 35:e343
Nunes-Souza V, Alenina N, Qadri F, Penninger JM, Santos RAS, Bader M, Rabelo LA (2016) CD36/sirtuin 1 axis impairment contributes to hepatic steatosis in ACE2-deficient mice. Oxid Med Cell Longev 2016:6487509
Dias SSG, Soares VC, Ferreira AC, Sacramento CQ, Fintelman-Rodrigues N, Temerozo JR, Teixeira L, da Silva MAN, Barreto E, Mattos M (2020) Lipid droplets fuel SARS-CoV-2 replication and production of inflammatory mediators. PLoS Pathog 16:e1009127
Guarantee L, Lecture FHE (2011) Sirtuins, aging, and medicine. N Engl J Med 364:2235–2244
Longo VD, Kennedy BK (2006) Sirtuins in aging and age-related disease. Cell 126:257–268
Jęśko H, Wencel P, Strosznajder RP, Strosznajder JB (2017) Sirtuins and their roles in brain aging and neurodegenerative disorders. Neurochem Res 42:876–890
Elibol B, Kilic U (2018) High levels of SIRT1 expression as a protective mechanism against disease-related conditions. Front Endocrinol (Lausanne) 9:614. https://doi.org/10.3389/fendo.2018.00614
Yoshizaki T, Milne JC, Imamura T, Schenk S, Sonoda N, Babendure JL, Lu JC, Smith JJ, Jirousek MR, Olefsky JM (2009) SIRT1 exerts anti-inflammatory effects and improves insulin sensitivity in adipocytes. Mol Cell Biol 29:1363–1374. https://doi.org/10.1128/mcb.00705-08
Zhang J (2007) The direct involvement of SirT1 in insulin-induced insulin receptor substrate-2 tyrosine phosphorylation. J Biol Chem 282:34356–34364
Pillai VB, Sundaresan NR, Gupta MP (2014) Regulation of Akt signaling by sirtuins: its implication in cardiac hypertrophy and aging. Circul Res 114:368–378
Moynihan KA, Grimm AA, Plueger MM, Bernal-Mizrachi E, Ford E, Cras-Méneur C, Permutt MA, Imai S-i (2005) Increased dosage of mammalian Sir2 in pancreatic β cells enhances glucose-stimulated insulin secretion in mice. Cell Metab 2:105–117
Bordone L, Motta MC, Picard F, Robinson A, Jhala US, Apfeld J, McDonagh T, Lemieux M, McBurney M, Szilvasi A (2005) Sirt1 regulates insulin secretion by repressing UCP2 in pancreatic β cells. PLoS Biol 4:e31
Zhou S, Tang X, Chen H-Z (2018) Sirtuins and insulin resistance. Front Endocrinol 9:748
Alqarni MH, Foudah AI, Muharram MM, Labrou NE (2021) The pleiotropic function of human sirtuins as modulators of metabolic pathways and viral infections. Cells 10:460
Budayeva HG, Rowland EA, Cristea IM (2016) Intricate roles of mammalian sirtuins in defense against viral pathogens. J Virol 90:5–8
Bordoni V, Tartaglia E, Sacchi A, Fimia GM, Cimini E, Casetti R, Notari S, Grassi G, Marchioni L, Bibas M (2021) The unbalanced p53/SIRT1 axis may impact lymphocyte homeostasis in COVID-19 patients. Int J Infect Dis 105:49–53
Boström P, Wu J, Jedrychowski MP, Korde A, Ye L, Lo JC, Rasbach KA, Boström EA, Choi JH, Long JZ (2012) A PGC1-α-dependent myokine that drives brown-fat-like development of white fat and thermogenesis. Nature 481:463–468
Rabiee F, Lachinani L, Ghaedi S, Nasr-Esfahani MH, Megraw TL, Ghaedi K (2020) New insights into the cellular activities of Fndc5/Irisin and its signaling pathways. Cell Biosci 10:1–10
Huh JY, Panagiotou G, Mougios V, Brinkoetter M, Vamvini MT, Schneider BE, Mantzoros CS (2012) FNDC5 and irisin in humans: I. Predictors of circulating concentrations in serum and plasma and II. mRNA expression and circulating concentrations in response to weight loss and exercise. Metabolism 61:1725–1738
Stengel A, Hofmann T, Goebel-Stengel M, Elbelt U, Kobelt P, Klapp BF (2013) Circulating levels of irisin in patients with anorexia nervosa and different stages of obesity–correlation with body mass index. Peptides 39:125–130
Villarroya F (2012) Irisin, turning up the heat. Cell Metab 15:277–278
Mazur-Bialy AI, Pocheć E, Zarawski M (2017) Anti-inflammatory properties of irisin, mediator of physical activity, are connected with TLR4/MyD88 signaling pathway activation. Int J Mol Sci 18:701
Crunkhorn S (2013) Betatrophin boosts β-cells. Nat Rev Drug Discov 12:504–505
Lee HJ, Lee JO, Kim N, Kim JK, Kim HI, Lee YW, Kim SJ, Choi J-I, Oh Y, Kim JH (2015) Irisin, a novel myokine, regulates glucose uptake in skeletal muscle cells via AMPK. Mol Endocrinol 29:873–881
Bernardi S, Tikellis C, Candido R, Tsorotes D, Pickering RJ, Bossi F, Carretta R, Fabris B, Cooper ME, Thomas MC (2015) ACE2 deficiency shifts energy metabolism towards glucose utilization. Metabolism 64:406–415
de Oliveira M, De Sibio MT, Mathias LS, Rodrigues BM, Sakalem ME, Nogueira CR (2020) Irisin modulates genes associated with severe coronavirus disease (COVID-19) outcome in human subcutaneous adipocytes cell culture. Mol Cell Endocrinol 515:110917
Santos A, Magro DO, Evangelista-Poderoso R, Saad MJA (2021) Diabetes, obesity, and insulin resistance in COVID-19: molecular interrelationship and therapeutic implications. Diabetol Metab Syndr 13:1–14
Lim S, Bae JH, Kwon H-S, Nauck MA (2020) COVID-19 and diabetes mellitus: from pathophysiology to clinical management. Nat Rev Endocrinol 17:1–20
Iwasaki M, Saito J, Zhao H, Sakamoto A, Hirota K, Ma D (2021) Inflammation triggered by SARS-CoV-2 and ACE2 augment drives multiple organ failure of severe COVID-19: molecular mechanisms and implications. Inflammation 44:13–34
Nataraj C, Oliverio MI, Mannon RB, Mannon PJ, Audoly LP, Amuchastegui CS, Ruiz P, Smithies O, Coffman TM (1999) Angiotensin II regulates cellular immune responses through a calcineurin-dependent pathway. J Clin Investig 104:1693–1701
Ji Y, Liu J, Wang Z, Liu N (2009) Angiotensin II induces inflammatory response partly via toll-like receptor 4-dependent signaling pathway in vascular smooth muscle cells. Cell Physiol Biochem 23:265–276
Wu J, Yang X, Zhang Y-F, Zhou S-F, Zhang R, Dong X-Q, Fan J-J, Liu M, Yu X-Q (2009) Angiotensin II upregulates Toll-like receptor 4 and enhances lipopolysaccharide-induced CD40 expression in rat peritoneal mesothelial cells. Inflamm Res 58:473–482
Rehman K, Akash MSH (2016) Mechanisms of inflammatory responses and development of insulin resistance: how are they interlinked? J Biomed Sci 23:1–18
Jager J, Grémeaux T, Cormont M, Le Marchand-Brustel Y, Tanti JF (2007) Interleukin-1beta-induced insulin resistance in adipocytes through down-regulation of insulin receptor substrate-1 expression. Endocrinology 148:241–251. https://doi.org/10.1210/en.2006-0692
Kirwan JP, Jing M (2002) Modulation of insulin signaling in human skeletal muscle in response to exercise. Exer Sport Sci Rev 30:85–90
Kern PA, Ranganathan S, Li C, Wood L, Ranganathan G (2001) Adipose tissue tumor necrosis factor and interleukin-6 expression in human obesity and insulin resistance. Am J Physiol-Endocrinol Metab 280:E745–E751
Hilton D, Emanuelli B, Peraldi P, Filloux C, Sawka-Verhelle D, Van Obberghen E (2000) SOCS-3 is an insulin-induced negative regulator of insulin signaling. J Biol Chem 275:15985–15991
Tilg H, Moschen AR (2008) Inflammatory mechanisms in the regulation of insulin resistance. Mol Med 14:222–231
Kanda H, Tateya S, Tamori Y, Kotani K, Hiasa K-i, Kitazawa R, Kitazawa S, Miyachi H, Maeda S, Egashira K (2006) MCP-1 contributes to macrophage infiltration into adipose tissue, insulin resistance, and hepatic steatosis in obesity. J Clin Investig 116:1494–1505
Kitade H, Sawamoto K, Nagashimada M, Inoue H, Yamamoto Y, Sai Y, Takamura T, Yamamoto H, K-i M, Ginsberg HN (2012) CCR5 plays a critical role in obesity-induced adipose tissue inflammation and insulin resistance by regulating both macrophage recruitment and M1/M2 status. Diabetes 61:1680–1690
Reiterer M, Rajan M, Gomez-Banoy N, Lau JD, Gomez-Escobar LG, Gilani A, Alvarez-Mullet S, Sholle ET, Chandar V, Bram Y (2021) Hyperglycemia in acute COVID-19 is characterized by adipose tissue dysfunction and insulin resistance. medRxiv
Ohno H, Matsuzaka T, Tang N, Sharma R, Motomura K, Shimura T, Satoh A, Han S-i, Takeuchi Y, Aita Y (2018) Transgenic mice overexpressing SREBP-1a in male ob/ob mice exhibit lipodystrophy and exacerbate insulin resistance. Endocrinol 159:2308–2323
Zhao L, Varghese Z, Moorhead J, Chen Y, Ruan XZ (2018) CD36 and lipid metabolism in the evolution of atherosclerosis. Br Med Bull 126:101–112
Miller R, Wentzel A, Richards G (2020) COVID-19: NAD+ deficiency may predispose the aged, obese and type2 diabetics to mortality through its effect on SIRT1 activity. Med Hypoth 144:110044
Korta P, Pocheć E, Mazur-Biały A (2019) Irisin as a multifunctional protein: implications for health and certain diseases. Medicina 55:485
Khunti K, Del Prato S, Mathieu C, Kahn SE, Gabbay RA, Buse JB (2021) COVID-19, hyperglycemia, and new-onset diabetes. Diabetes Care 44:2645
Acknowledgements
The technical assistance of Mr. Dara Joukar in drawing of figures is gratefully acknowledged. The authors also thank Ms. Melissa T. Yang, PhD for her kind language editing of the manuscript.
Funding
There was no funding for this review.
Author information
Authors and Affiliations
Contributions
MM, IN, KV and MK interpreted and wrote the manuscript. IN and MM supervised the study. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare the following financial interests/personal relationships, which may be considered as potential competing interests: An award was given to Iraj Nabipour by “Novo Nordisk Pars” (The Best Innovator of Diabetes in Iran 2013). For the remaining authors none were declared.
Ethical approval
This article does not contain any studies with human participants performed by any of the authors.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mahmudpour, M., Vahdat, K., Keshavarz, M. et al. The COVID-19-diabetes mellitus molecular tetrahedron. Mol Biol Rep 49, 4013–4024 (2022). https://doi.org/10.1007/s11033-021-07109-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-07109-y