Abstract
Background
Major depressive disorder (MDD) is a polygenic, and highly prevalent disorder affecting 322 million people globally. It results in several psychological changes which adversely affect different dimensions of life and may lead to suicide.
Methods
Whole exome sequencing of 15 MDD patients, enrolled at the Dr. A. Q. Khan Institute of Behavioral Sciences, Karachi, was performed using NextSeq500. Different bioinformatics tools and databases like ANNOVAR, ALoFT, and GWAS were used to identify both common and rare variants associated with the pathogenesis of MDD.
Results
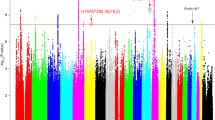
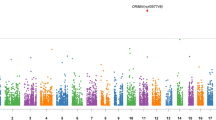
A total of 1985 variations were identified in 479 MDD-related genes. Several SNPs including rs1079610, rs11750538, rs1799913, rs1801131, rs2230267, rs2231187, rs3819976, rs4314963, rs56265970, rs587780434, rs6330, rs75111588, rs7596487, and rs9624909 were prioritized due to their deleteriousness and frequency difference between the patients and the South Asian population. A non-synonymous variation rs56265970 (BCR) had 26% frequency in patients and was not found in the South Asian population; a multiallelic UTR-5′ insertion rs587780434 (RELN) was present with an allelic frequency of 70% in patients whereas 22% in the SAS population. Genetic alterations in PABPC1 genes, a stress-associated gene also had higher allele frequency in the cases than in the normal population.
Conclusion
This present study identifies both common and rare variants in the genes associated with the pathogenesis of MDD in Pakistani patients. Genetic variations in BCR, RELN, and stress-associated PABPC1 suggest potential roles in the pathogenesis of MDD.



Similar content being viewed by others
Data availability
The raw data that support the findings of this study has been deposited in the NCBI SRA archive under the BioProject PRJNA774142.
References
World Health Organization (2017) Depression and other common mental disorders: global health estimates. World Health Organization, Geneva
Kessler RC, Berglund P, Demler O, Jin R, Merikangas KR, Walters EE (2005) Lifetime prevalence and age-of-onset distributions of DSM-IV disorders in the national comorbidity survey replication. Arch Gen Psychiatry 62(6):593–602
Merikangas KR, He J-P, Burstein M, Swanson SA, Avenevoli S, Cui L, Benjet C, Georgiades K, Swendsen J (2010) Lifetime prevalence of mental disorders in US adolescents: results from the national comorbidity survey replication-adolescent supplement (NCS-A). J Am Acad Child Adolesc Psychiatry 49(10):980–989
Han K-M, Han M-R, Kim A, Kang W, Kang Y, Kang J, Tae W-S, Cho Y, Ham B-J (2020) A study combining whole-exome sequencing and structural neuroimaging analysis for major depressive disorder. J Affect Disord 262:31–39
Ahmed B, Enam SF, Iqbal Z, Murtaza G, Bashir S (2016) Depression and anxiety: a snapshot of the situation in Pakistan. Int J Neurosci Behav Sci 4(2):32–36
Neitzke AB (2016) An illness of power: gender and the social causes of depression. Cult Med Psychiatry 40(1):59–73
Howard DM, Adams MJ, Clarke T-K, Hafferty JD, Gibson J, Shirali M, Coleman JR, Hagenaars SP, Ward J, Wigmore EM (2019) Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat Neurosci 22(3):343–352
Tombácz D, Maróti Z, Kalmár T, Csabai Z, Balázs Z, Takahashi S, Palkovits M, Snyder M, Boldogkői Z (2017) High-coverage whole-exome sequencing identifies candidate genes for suicide in victims with major depressive disorder. Sci Rep 7(1):1–11
Zhou W, Chen L, Jiang B, Sun Y, Li M, Wu H, Zhang N, Sun X, Qin S (2021) Large-scale whole-exome sequencing association study identifies Foxh1 gene and sphingolipid metabolism pathway influencing major depressive disorder. CNS Neurosci Ther 27(11):1425–1428
Boda E (2021) Myelin and oligodendrocyte lineage cell dysfunctions: new players in the etiology and treatment of depression and stress-related disorders. Eur J Neurosci 53(1):281–297
Yang J, Chen C, Jin X, Liu L, Lin J, Kang X, Zhu S (2021) Wfs1 and related molecules as key candidate genes in the hippocampus of depression. Front Genet 11:1760
Dunn EC, Brown RC, Dai Y, Rosand J, Nugent NR, Amstadter AB, Smoller JW (2015) Genetic determinants of depression: recent findings and future directions. Harv Rev Psychiatry 23(1):1–18
Smoller JW (2016) The genetics of stress-related disorders: PTSD, depression, and anxiety disorders. Neuropsychopharmacology 41(1):297–319
Levchenko A, Vyalova NM, Nurgaliev T, Pozhidaev IV, Simutkin GG, Bokhan NA, Ivanova SA (2020) Nrg1, Pip4k2a, and Htr2c as potential candidate biomarker genes for several clinical subphenotypes of depression and bipolar disorder. Front Genet. https://doi.org/10.3389/fgene.2020.00936
Maul S, Giegling I, Fabbri C, Corponi F, Serretti A, Rujescu D (2020) Genetics of resilience: implications from genome-wide association studies and candidate genes of the stress response system in posttraumatic stress disorder and depression. Am J Med Genet B 183(2):77–94
Yang C, Li S, Ma JX, Li Y, Zhang A, Sun N, Wang Y, Xu Y, Zhang K (2019) Whole exome sequencing identifies a novel predisposing gene, MAPKAP1, for familial mixed mood disorder. Front Genet 10:74
Kang H-J, Kim K-T, Park Y, Yoo K-H, Kim J-W, Lee J-Y, Kim S-W, Shin I-S, Kim JH, Kim J-M (2021) Genetic markers for depressive disorders with earlier age at onset. Prog Neuro-Psychopharmacol Biol Psychiatry 108:110176
Tian R, Ge T, Liu JZ, Lam M, Levey DF, Gelernter J, Stein MB, Tsai EA, Huang H, Lencz T (2021) Whole exome sequencing in the UK biobank reveals risk gene Slc2a1 and biological insights for major depressive disorder. medRxiv. https://doi.org/10.1016/j.euroneuro.2021.07.173
Winnepenninckx B, Backeljau T, Mackey LY, Brooks JM, De Wachter R, Kumar S, Garey JR (1995) 18S rRNA data indicate that Aschelminthes are polyphyletic in origin and consist of at least three distinct clades. Mol Biol Evol 12(6):1132–1137
Andrew, S. (2018). FastQC: a quality control tool for high throughput sequence data. Accessed http://www.bioinformatics.bbsrc.ac.uk/projects/fastqc
Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26(5):589–595
Board Institute. (2019). Picard tools. Accessed http://broadinstitute.github.io/picard/
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M (2010) The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20(9):1297–1303
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T, O’Donnell-Luria AH, Ware JS, Hill AJ, Cummings BB (2016) Analysis of protein-coding genetic variation in 60,706 humans. Nature 536(7616):285–291
Wang K, Li M, Hakonarson H (2010) ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38(16):e164
Ng PC, Henikoff S (2003) SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res 31(13):3812–3814
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, Kondrashov AS, Sunyaev SR (2010) A method and server for predicting damaging missense mutations. Nat Methods 7(4):248–249
Schwarz JM, Cooper DN, Schuelke M, Seelow D (2014) MutationTaster2: mutation prediction for the deep-sequencing age. Nat Methods 11(4):361–362
Balasubramanian S, Fu Y, Pawashe M, McGillivray P, Jin M, Liu J, Karczewski KJ, MacArthur DG, Gerstein M (2017) Using ALoFT to determine the impact of putative loss-of-function variants in protein-coding genes. Nat Commun 8(1):1–11
Buniello A, MacArthur JAL, Cerezo M, Harris LW, Hayhurst J, Malangone C, McMahon A, Morales J, Mountjoy E, Sollis E (2019) The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res 47(D1):D1005–D1012
Cortes A, Albers PK, Dendrou CA, Fugger L, McVean G (2020) Identifying cross-disease components of genetic risk across hospital data in the UK Biobank. Nat Genet 52(1):126–134
Piñero J, Ramírez-Anguita JM, Saüch-Pitarch J, Ronzano F, Centeno E, Sanz F, Furlong LI (2020) The DisGeNET knowledge platform for disease genomics: 2019 update. Nucleic Acids Res 48(D1):D845–D855
Landrum MJ, Lee JM, Riley GR, Jang W, Rubinstein WS, Church DM, Maglott DR (2014) ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res 42(D1):D980–D985
Brennand K, Simone A, Tran N, Gage F (2012) Modeling psychiatric disorders at the cellular and network levels. Mol Psychiatry 17(12):1239–1253
Hamosh A, Scott AF, Amberger JS, Bocchini CA, McKusick VA (2005) Online mendelian inheritance in man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res 33(suppl_1):D514–D517
Kopanos C, Tsiolkas V, Kouris A, Chapple CE, Aguilera MA, Meyer R, Massouras A (2019) VarSome: the human genomic variant search engine. Bioinformatics 35(11):1978–1980
Poleszak E, Wlaź P, Szewczyk B, Wlaź A, Kasperek R, Wróbel A, Nowak G (2011) A complex interaction between glycine/NMDA receptors and serotonergic/noradrenergic antidepressants in the forced swim test in mice. J Neural Transm 118(11):1535–1546
Knox C, Law V, Jewison T, Liu P, Ly S, Frolkis A, Pon A, Banco K, Mak C, Neveu V (2010) DrugBank 3.0: a comprehensive resource for ‘omics’ research on drugs. Nucleic Acids Res 39(suppl_1):D1035–D1041
Sangkuhl K, Klein T, Altman R (2009) Selective serotonin reuptake inhibitors (SSRI) pathway. Pharm Genomics 19(11):907–909
Whirl-Carrillo M, McDonagh EM, Hebert J, Gong L, Sangkuhl K, Thorn C, Altman RB, Klein TE (2012) Pharmacogenomics knowledge for personalized medicine. Clin Pharmacol Ther 92(4):414–417
The 1000 Genome Project Consortium (2015) A global reference for human genetic variation. Nature 526(7571):68–74
Fang P, He J-Y, Han A-X, Lan T, Dai D-P, Cai J-P, Hu G-X (2017) Effects of CYP2C19 variants on fluoxetine metabolism in vitro. Pharmacology 100(1–2):91–97
Zhang L-S, Li H-B, Zeng J, Yang Y, Ding C (2018) Knobloch syndrome caused by homozygous frameshift mutation of the COL18A1 gene in a Chinese pedigree. Int J Ophthalmol 11(6):918–922
Brest P, Lapaquette P, Souidi M, Lebrigand K, Cesaro A, Vouret-Craviari V, Mari B, Barbry P, Mosnier J-F, Hébuterne X (2011) A synonymous variant in IRGM alters a binding site for miR-196 and causes deregulation of IRGM-dependent xenophagy in Crohn’s disease. Nat Genet 43(3):242–245
Kochar B, Barnes EL, Long MD, Cushing KC, Galanko J, Martin CF, Raffals LE, Sandler RS (2018) Depression is associated with more aggressive inflammatory bowel disease. Am J Gastroenterol 113(1):80–85
Barberio B, Zamani M, Black CJ, Savarino EV, Ford AC (2021) Prevalence of symptoms of anxiety and depression in patients with inflammatory bowel disease: a systematic review and meta-analysis. Lancet Gastroenterol Hepatol. https://doi.org/10.1016/S2468-1253(21)00014-5
Kohl S, Kitiratschky V, Grau T, Schaich S, Wissinger B, Group ACS (2008) ABCA4 gene analysis in patients with autosomal recessive cone and cone rod dystrophies. Investig Ophthalmol Vis Sci 49(13):3098–3098
Baselmans BM, Jansen R, Ip HF, van Dongen J, Abdellaoui A, van de Weijer MP, Bao Y, Smart M, Kumari M, Willemsen G (2019) Multivariate genome-wide analyses of the well-being spectrum. Nat Genet 51(3):445–451
Turley P, Walters RK, Maghzian O, Okbay A, Lee JJ, Fontana MA, Nguyen-Viet TA, Wedow R, Zacher M, Furlotte NA (2018) Multi-trait analysis of genome-wide association summary statistics using MTAG. Nat Genet 50(2):229–237
Wigner P, Czarny P, Synowiec E, Bijak M, Białek K, Talarowska M, Galecki P, Szemraj J, Sliwinski T (2018) Association between single nucleotide polymorphisms of TPH1 and TPH2 genes, and depressive disorders. J Cell Mol Med 22(3):1778–1791
Rathje M, Waxman H, Benoit M, Tammineni P, Leu C, Loebrich S, Nedivi E (2021) Genetic variants in the bipolar disorder risk locus syne1 that affect Cpg2 expression and protein function. Mol Psychiatry 26(2):508–523
Loebrich S, Rathje M, Hager E, Ataman B, Harmin DA, Greenberg ME, Nedivi E (2016) Genomic mapping and cellular expression of human Cpg2 transcripts in the syne1 gene. Mol Cell Neurosci 71:46–55
Green EK, Grozeva D, Forty L, Gordon-Smith K, Russell E, Farmer A, Hamshere M, Jones IR, Jones L, McGuffin P (2013) Association at syne1 in both bipolar disorder and recurrent major depression. Mol Psychiatry 18(5):614–617
Lussier AL, Lebedeva K, Fenton EY, Guskjolen A, Caruncho HJ, Kalynchuk LE (2013) The progressive development of depression-like behavior in corticosterone-treated rats is paralleled by slowed granule cell maturation and decreased reelin expression in the adult dentate gyrus. Neuropharmacology 71:174–183
Hashimoto R, Okada T, Kato T, Kosuga A, Tatsumi M, Kamijima K, Kunugi H (2005) The breakpoint cluster region gene on chromosome 22q11 is associated with bipolar disorder. Biol Psychiatry 57(10):1097–1102
Kaviani M, Nikooyeh B, Zand H, Yaghmaei P, Neyestani TR (2020) Effects of vitamin D supplementation on depression and some involved neurotransmitters. J Affect Disord 269:28–35
Kilicaslan DY, Cumaogullari O, Emiral E, Tezer N, Oncu B, Ozdag H, Canturk N, Tufan NLS, Satiroglu L (2021) Investigation of polymorphic variants of Slc6a4, Tph-1, and Tph-2 genes in cases of completed suicide. J Men Health. https://doi.org/10.31083/jomh.2021.116
Varinthra P, Liu IY (2019) Molecular basis for the association between depression and circadian rhythm. Tzu-Chi Med J 31(2):67–72
Utge SJ, Soronen P, Loukola A, Kronholm E, Ollila HM, Pirkola S, Porkka-Heiskanen T, Partonen T, Paunio T (2010) Systematic analysis of circadian genes in a population-based sample reveals association of TIMELESS with depression and sleep disturbance. PLoS ONE 5(2):e9259
Markmiller S, Soltanieh S, Server KL, Mak R, Jin W, Fang MY, Luo E-C, Krach F, Yang D, Sen A (2018) Context-dependent and disease-specific diversity in protein interactions within stress granules. Cell 172(3):590–604
Shyn SI, Shi J, Kraft J, Potash JB, Knowles J, Weissman M, Garriock H, Yokoyama J, McGrath P, Peters E (2011) Novel loci for major depression identified by genome-wide association study of sequenced treatment alternatives to relieve depression and meta-analysis of three studies. Mol Psychiatry 16(2):202–215
Kao A, Kuzman MR, Tiwari A, Zivkovic M, Chowdhury N, Medved V, Kekin I, Zai C, Lieberman J, Meltzer HY (2014) Methylenetetrahydrofolate reductase gene variants and antipsychotic-induced weight gain and metabolic disturbances. J Psychiatr Res 54:36–42
Gupta M, Neavin D, Liu D, Biernacka J, Hall-Flavin D, Bobo WV, Frye MA, Skime M, Jenkins GD, Batzler A (2016) TSPAN5, ERICH3 and selective serotonin reuptake inhibitors in major depressive disorder: pharmacometabolomics-informed pharmacogenomics. Mol Psychiatry 21(12):1717–1725
Fonseka TM, Tiwari AK, Gonçalves VF, Lieberman JA, Meltzer HY, Goldstein BI, Kennedy JL, Kennedy SH, Müller DJ (2015) The role of genetic variation across IL-1β, IL-2, IL-6, and BDNF in antipsychotic-induced weight gain. World J Biol Psychiatry 16(1):45–56
Lane H-Y, Liu Y-C, Huang C-L, Chang Y-C, Wu P-L, Lu C-T, Chang W-H (2006) Risperidone-related weight gain: genetic and nongenetic predictors. J Clin Psychopharmacol 26(2):128–134
Lawal HO, Krantz DE (2013) SLC18: Vesicular neurotransmitter transporters for monoamines and acetylcholine. Mol Aspects Med 34(2–3):360–372
Zhou L, Ma S, Yeung PKK, Wong YH, Tsim KWK, So K, Lam L, Chung S (2016) Anxiety and depression with neurogenesis defects in exchange protein directly activated by cAMP 2-deficient mice are ameliorated by a selective serotonin reuptake inhibitor, Prozoc. Transl Psychiatry 6(9):e881–e881
Wong M-L, Whelan F, Deloukas P, Whittaker P, Delgado M, Cantor RM, McCann SM, Licinio J (2006) Phosphodiesterase genes are associated with susceptibility to major depression and antidepressant treatment response. Proc Natl Acad Sci USA 103(41):15124–15129
Acknowledgements
We would like to extend our special gratitude to the medical staff and authorities of Dr. Abdul Qadir Khan Institute of Behavioral Sciences, Karachi for permitting us to collect blood samples on their premises.
Funding
The study was performed with the funding provided by The Searle Company Limited (TSCL), Karachi, Pakistan to the institute (Research Project ID 8211).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors had nothing to disclose that might construe a conflict of interest.
Ethical approval
The procedures performed in this study involving human participants were in accordance with the Institutional Ethical Committee (Study accession number IEC-046-HB-2019), and with the 1964 Helsinki declaration and its later amendments.
Informed consent
Informed consent was obtained from all the individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Qazi, S.R., Irfan, M., Ramzan, Z. et al. Identification of putative genetic variants in major depressive disorder patients in Pakistan. Mol Biol Rep 49, 2283–2292 (2022). https://doi.org/10.1007/s11033-021-07050-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-07050-0