Abstract
Suppressing vascular endothelial growth factor (VEGF), its receptor (VEGFR2), and the VEGF/VEGFR2 signaling cascade system to inhibit angiogenesis has emerged as a possible cancer therapeutic target. The present work was designed to discover and evaluate bioactive phytochemicals from the Clerodendrum inerme (L.) Gaertn plant for their anti-angiogenic potential. Molecular docking of twenty-one phytochemicals against the VEGFR-2 (PDB ID: 3VHE) protein was performed, followed by ADMET profiling and molecular docking simulations. These investigations unveiled two hit compounds, cirsimaritin (− 12.29 kcal/mol) and salvigenin (− 12.14 kcal/mol), with the highest binding energy values when compared to the reference drug, Sorafenib (− 15.14 kcal/mol). Furthermore, only nine phytochemicals (cirsimaritin and salvigenin included) obeyed Lipinski’s rule of five and passed ADMET filters. Molecular dynamics simulations run over 100 ns revealed that the protein–ligand complexes remained stable with minimal backbone fluctuations. The binding free energy values of cirsimaritin (− 52.35 kcal/mol) and salvigenin (− 55.89 kcal/mol), deciphered by MM-GBSA analyses, further corroborated the docking interactions. The HOMO–LUMO band energy gap (ΔE) was calculated using density-functional theory (DFT) and substantiated using density of state (DOS) spectra. The chemical reactivity analyses revealed that salvigenin exhibited the highest chemical softness value (6.384 eV), the lowest hardness value (0.07831 eV), and the lowest ΔE value (0.1566 eV), which implies salvigenin was less stable and chemically more reactive than cirsimaritin and sorafenib. These findings provide further evidence that cirsimaritin and salvigenin have the ability to prevent angiogenesis and the development of cancer. Nevertheless, more in vitro and in vivo confirmation is necessary.
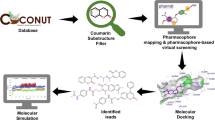
Graphical abstract
Illustration showing the work flow of screening of phytochemicals from C. inerme leaves for their anti-angiogenic potential.









Similar content being viewed by others
Data availability
All the data generated or analyzed during this study are included in this article. Any additional data needed are available upon request to the corresponding author.
References
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144:646–674. https://doi.org/10.1016/j.cell.2011.02.013
Baeriswyl V, Christofori G (2009) The angiogenic switch in carcinogenesis. Semin Cancer Biol 19:329–337. https://doi.org/10.1016/j.semcancer.2009.05.003
Jiang X, Wang J, Deng X et al (2020) The role of microenvironment in tumor angiogenesis. J Exp Clin Cancer Res 39:1–19. https://doi.org/10.1186/s13046-020-01709-5
Sphyris N, King C, Hoar J et al (2021) Carcinoma cells that have undergone an epithelial-mesenchymal transition differentiate into endothelial cells and contribute to tumor growth. Oncotarget 12:823–844. https://doi.org/10.18632/oncotarget.27940
Shibuya M (2011) Vascular Endothelial Growth Factor (VEGF) and Its Receptor (VEGFR) signaling in angiogenesis: a crucial target for anti- and pro-angiogenic therapies. Genes Cancer 2:1097–1105. https://doi.org/10.1177/1947601911423031
Apte RS, Chen DS, Ferrara N (2019) VEGF in signaling and disease: beyond discovery and development. Cell 176:1248–1264. https://doi.org/10.1016/j.cell.2019.01.021
Shibuya M (2013) VEGFR and type-V RTK activation and signaling. Cold Spring Harb Perspect Biol 5:1–13. https://doi.org/10.1101/cshperspect.a009092
Simons M, Gordon E, Claesson-welsh L (2016) Mechanisms and regulation of endothelial VEGF receptor signalling. Nat Rev Mol cell Biol 17:611–625. https://doi.org/10.1038/nrm.2016.87
Zhuang G, Yu K, Jiang Z et al (2013) Phosphoproteomic analysis implicates the mTORC2-FoxO1 axis in VEGF signaling and feedback activation of receptor tyrosine kinases. Cell Biol 6:1–12
Luo M, Hou L, Li J et al (2016) VEGF/NRP-1axis promotes progression of breast cancer via enhancement of epithelial-mesenchymal transition and activation of NF-κB and β-catenin. Cancer Lett 373:1–11. https://doi.org/10.1016/j.canlet.2016.01.010
Ni H, Guo M, Zhang X et al (2021) VEGFR2 inhibition hampers breast cancer cell proliferation via enhanced mitochondrial biogenesis. Cancer Biol Med 18:139–154. https://doi.org/10.20892/j.issn.2095-3941.2020.0151
Al-Muntaser SM, Al-Karmalawy AA, El-Naggar AM et al (2023) Novel 4-thiophenyl-pyrazole, pyridine, and pyrimidine derivatives as potential antitumor candidates targeting both EGFR and VEGFR-2; design, synthesis, biological evaluations, and in silico studies. RSC Adv 13:12184–12203. https://doi.org/10.1039/d3ra00416c
El-Naggar AM, Hassan AMA, Elkaeed EB, Mohamed S, Alesawy AAA (2022) Design, synthesis, and SAR studies of novel 4-methoxyphenyl pyrazole and pyrimidine derivatives as potential dual tyrosine kinase inhibitors targeting both EGFR and VEGFR-2. Bioorg Chem 123
Abdel-Mohsen HT, Abdullaziz MA, El Kerdawy AM et al (2020) Targeting receptor tyrosine kinase VEGFR-2 in hepatocellular cancer: rational design, synthesis and biological evaluation of 1,2-disubstituted benzimidazoles. Molecules 25. https://doi.org/10.3390/molecules25040770
Aziz MA, Serya RAT, Lasheen DS et al (2016) Discovery of potent VEGFR-2 inhibitors based on furopyrimidine and thienopyrimidne scaffolds as cancer targeting agents. Sci Rep 6:1–20. https://doi.org/10.1038/srep24460
Papadopoulos N, Martin J, Ruan Q et al (2012) Binding and neutralization of vascular endothelial growth factor (VEGF) and related ligands by VEGF Trap, ranibizumab and bevacizumab. Angiogenesis 15:171–185. https://doi.org/10.1007/s10456-011-9249-6
Marie S, Ef A, Gyawali B (2020) Assessing the risk-bene fi t pro fi le of ramucirumab in patients with advanced solid tumors: a meta-analysis of randomized controlled trials. EClinicalMedicine 25:1–8. https://doi.org/10.1016/j.eclinm.2020.100458
Liu Z, Chen H, Zheng L et al (2023) Angiogenic signaling pathways and anti-angiogenic therapy for cancer. Signal Transduct Target Ther 8:1–39. https://doi.org/10.1038/s41392-023-01460-1
Dobbin SJH, Cameron AC, Petrie MC et al (2018) Toxicity of cancer therapy: what the cardiologist needs to know about angiogenesis inhibitors. Heart 104:1995–2002. https://doi.org/10.1136/heartjnl-2018-313726
Shaker B, Ahmad S, Lee J et al (2021) In silico methods and tools for drug discovery. Comput Biol Med 137:1–15. https://doi.org/10.1016/j.compbiomed.2021.104851
Agu PC, Afiukwa CA, Orji OU et al (2023) Molecular docking as a tool for the discovery of molecular targets of nutraceuticals in diseases management. Sci Rep 13:1–18. https://doi.org/10.1038/s41598-023-40160-2
Ye N, Yang Z, Liu Y (2022) Applications of density functional theory in COVID-19 drug modeling. Drug Discov Today 27:1411–1419. https://doi.org/10.1016/j.drudis.2021.12.017
Toriyama MY, Ganose AM, Dylla M et al (2022) How to analyse a density of states. Mater Today Electron 1:1–4. https://doi.org/10.1016/j.mtelec.2022.100002
Kar P, Mishra DK, Roy A et al (2021) Elucidation of phytomedicinal efficacies of Clerodendrum inerme (L.) Gaertn. (Wild Jasmine). South African J Bot 140:356–364. https://doi.org/10.1016/j.sajb.2020.07.027
Anitha R, Kannan P (2006) Antifungal activity of Clerodendrum inerme (L). and Clerodendrum phlomidis (L.). Turkish J Biol 30:139–142
Khanam D, Deb D, Dev S et al (2014) Analgesic and anti-inflammatory activities of ethanolic extract of Clerodendrum inerme (L.) Gaertn. Bangladesh Pharm J 17:62–66
Parveen M, Khanam Z, Ali M, Rahman SZ (2010) A novel lupene-type triterpenic glucoside from the leaves of Clerodendrum inerme. Nat Prod Res 24:167–176. https://doi.org/10.1080/14786410902975566
Thirumal M, Srimanthula S, Kishore G et al (2013) Analgesic and antipyretic effects of aqueous extract from Clerodendrum inerme (L.) Gaertn. leaves in animal models. Der Pharm Lettre 5:315–323
Khan AV (2006) Antibacterial potential of Clerodendrum inerme, crude extracts against some human pathogenic bacteria. Orient Pharm Exp Med 6:306–311. https://doi.org/10.3742/opem.2006.6.4.306
Mehdi H, Tan GT, Pezzuto JM et al (1997) Cell culture assay system for the evaluation of natural product-mediated anti-Hepatitis B virus activity. Phytomedicine 3:369–377. https://doi.org/10.1016/s0944-7113(97)80011-6
Wisessombat S, Tayeh M (2021) In vitro wound healing potential and antimicrobial activity of clerodendrum inerme leave extracts. Pharmacogn J 13:1542–1548. https://doi.org/10.5530/PJ.2021.13.196
Li D, Zhou J, Xia J et al (2015) Antioxidant activities of extract and fractions from Clerodendrum inerme 7:875–879
Ly HT, Nguyen TTH, Tran TTL et al (2019) Hypoglycemic and antioxidant activities of Clerodendrum inerme leaf extract on streptozotocin-induced diabetic models in mice. Chinese Herb Med 11:387–393. https://doi.org/10.1016/j.chmed.2019.08.001
Tayeh M, Hiransai P, Kommen H, Watanapokasin R (2020) Anti-migration and anti-invasion abilities of methanolic leaves extract of Clerodendrum inerme on lung cancer cells. Pharmacogn J 12:1024–1031. https://doi.org/10.5530/PJ.2020.12.145
Kalavathi R, Sagayagiri R (2016) Anticancer activity of ethanolic leaf extract of Clerodendrum inerme against lung adenocarcinoma epithelial cell line. Eur J Mol Biol Biochem 3:69–72
Chouhan MK, Hurkadale PJ, Hegde HV (2018) Evaluation of Clerodendrum inerme (L.) Gaertn. on Burkitt’s lymphoma cancer. Indian J Pharm Educ Res 52:241–247. https://doi.org/10.5530/ijper.52.2.27
Berman HM, Westbrook J, Feng Z et al (2000) The Protein Data Bank Helen. Nucleic Acids Res 28:235–242. https://doi.org/10.1093/nar/28.1.235
Oguro Y, Miyamoto N, Okada K et al (2010) Design, synthesis, and evaluation of 5-methyl-4-phenoxy-5H-pyrrolo[3,2-d] pyrimidine derivatives: novel VEGFR2 kinase inhibitors binding to inactive kinase conformation. Bioorganic Med Chem 18:7260–7273. https://doi.org/10.1016/j.bmc.2010.08.017
Ravindranath PA, Sanner MF (2016) AutoSite : an automated approach for pseudo-ligands prediction – from ligand-binding sites identification to predicting key ligand atoms. Bioinformatics 32:3142–3149. https://doi.org/10.1093/bioinformatics/btw367
Tian W, Chen C, Lei X et al (2018) CASTp 3. 0: computed atlas of surface topography of proteins. Nucleic Acids Res 46:363–367. https://doi.org/10.1093/nar/gky473
Morris GM, Ruth H, Lindstrom W et al (2009) Software news and updates AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791. https://doi.org/10.1002/jcc.21256
Vivek-Ananth RP, Mohanraj K, Sahoo AK, Samal A (2023) IMPPAT 2.0: an enhanced and expanded phytochemical atlas of indian medicinal plants. ACS Omega 8:8827–8845. https://doi.org/10.1021/acsomega.3c00156
O’Boyle NM, Banck M, James CA et al (2011) Open Babel: an open chemical toolbox Noel. J Cheminform 3:1–14
Lipinski CA, Lombardo F, Dominy DW, Feeney PJ (1997) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 23:3–25. https://doi.org/10.1016/S0169-409X(96)00423-1
Mullard A (2018) Re-assessing the rule of 5, two decades on. Nat Rev Drug Discov 17:777. https://doi.org/10.1038/nrd.2018.197
Veber DF, Johnson SR, Cheng HY et al (2002) Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem 45:2615–2623. https://doi.org/10.1021/jm020017n
Sander T, Freyss J, Von Korff M, Rufener C (2015) DataWarrior: an open-source program for chemistry aware data visualization and analysis. J Chem Inf Model 55:460–473. https://doi.org/10.1021/ci500588j
Dashti Y, Grkovic T, Quinn RJ (2014) Predicting natural product value, an exploration of anti-TB drug space. Nat Prod Rep 32:1–9. https://doi.org/10.1039/x0xx00000x
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7:1–13. https://doi.org/10.1038/srep42717
Yang H, Chaofeng L, Lixia S et al (2019) AdmetSAR 2.0: web-service for prediction and optimization of chemical ADMET properties. Bioinformatics 35:1067–1069. https://doi.org/10.1093/bioinformatics/bty707/5085368
Pires DEV, Blundell TL, Ascher DB (2015) pkCSM: predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. J Med Chem 58:4066–4072. https://doi.org/10.1021/acs.jmedchem.5b00104
BIOVIA, Dassault Systèmes (2023) BIOVIA Discovery Studio Visualizer
Adasme MF, Linnemann KL, Bolz SN et al (2021) PLIP 2021: expanding the scope of the protein-ligand interaction profiler to DNA and RNA. Nucleic Acids Res 49:W530–W534. https://doi.org/10.1093/nar/gkab294
NY DESRNY (2021) Schrödinger Release 2021-1: Desmond molecular dynamics system. maestro-Desmond interoperability tools. Schrödinger, New York
Bowers KJ, Chow E, Xu H, Dror RO, Gregersen BA, Klepeis JL, Kossvary I, Moraes MA, Salmon JK, Shan Y, Shaw DE (2006) Scalable algorithms for molecular dynamics simulations on commodity clusters. In: Proceedings of the ACM/IEEE Conference on Supercomputing (SC06), pp 1–13
Ayipo YO, Yahaya SN, Babamale HF et al (2021) β-Carboline alkaloids induce structural plasticity and inhibition of SARS-CoV-2 nsp3 macrodomain more potently than remdesivir metabolite GS-441524: computational approach. Turkish J Biol 45:503–517. https://doi.org/10.3906/biy-2106-64
Kumar A, Mishra DC, Angadi UB et al (2021) Inhibition potencies of phytochemicals derived from sesame against SARS-CoV-2 main protease: a molecular docking and simulation study. Front Chem 9:1–16. https://doi.org/10.3389/fchem.2021.744376
Wu Y, Lou L, Xie ZR (2020) A pilot study of all-computational drug design protocol-from structure prediction to interaction analysis. Front Chem 8:1–9. https://doi.org/10.3389/fchem.2020.00081
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenb DJ (2009) Gaussian 09 Revision D.1
Halim SA, Ibrahim MA (2022) Synthesis, spectral analysis, quantum studies, NLO, and thermodynamic properties of the novel 5-(6-hydroxy-4-methoxy-1-benzofuran-5-ylcarbonyl)-6-amino-3-methyl-1H-pyrazolo[3,4-b] pyridine (HMBPP). RSC Adv 12:13135–13153. https://doi.org/10.1039/d2ra01469f
Dennington R, Keith TA, Millam JM (2016) GaussView Version 6
O’Boyle NM, Tenderholt AL, Langner KM (2007) cclib: a library for package-independent computational chemistry algorithms. J Comput Chem 29:839–845. https://doi.org/10.1002/jcc.20823
Bouzina A, Bouone YO, Sekiou O et al (2023) In vitro antitumor activity, molecular dynamics simulation, DFT study, ADME prediction, and Eg5 binding of enastron analogues. RSC Adv 13:19567–19584. https://doi.org/10.1039/d3ra02904b
Jabin T, Biswas S, Islam S et al (2023) Effects of gamma-radiation on microbial, nutritional, and functional properties of Katimon mango peels: a combined biochemical and in silico studies. Heliyon 9:1–16. https://doi.org/10.1016/j.heliyon.2023.e21556
Murthy TPK, Joshi T, Gunnan S et al (2021) In silico analysis of Phyllanthus amarus phytochemicals as potent drugs against SARS-CoV-2 main protease. Curr Res Green Sustain Chem 4:1–14. https://doi.org/10.1016/j.crgsc.2021.100159
Li N, Sui Z, Liu Y et al (2018) A fast screening model for drug permeability assessment based on native small intestinal extracellular matrix. RSC Adv 8:34514–34524. https://doi.org/10.1039/C8RA05992F
Flores-Holguín N, Frau J, Glossman-Mitnik D (2021) In Silico Pharmacokinetics, ADMET study and conceptual DFT analysis of two plant cyclopeptides isolated from rosaceae as a computational peptidology approach. Front Chem 9:1–13. https://doi.org/10.3389/fchem.2021.708364
Mamizadeh R, Hosseinzadeh Z, Razzaghi-Asl N, Ramazani A (2018) In silico analysis of a few dietary phytochemicals as potential tumor chemo-sensitizers. Struct Chem 29:1139–1151. https://doi.org/10.1007/s11224-018-1098-0
Dechwongya P, Limpisood S, Boonnak N et al (2020) The intestinal efflux transporter inhibition activity of Xanthones from Mangosteen pericarp: an in silico, in vitro and ex vivo approach. Molecules 25:1–14
Vardhan S, Sahoo SK (2020) In silico ADMET and molecular docking study on searching potential inhibitors from limonoids and triterpenoids for COVID-19. Comput Biol Med 124:1–12. https://doi.org/10.1016/j.compbiomed.2020.103936
Sobańska AW, Wanat K, Brzezińska E (2019) Prediction of the blood-brain barrier permeability using RP-18 thin layer chromatography. Open Chem 17:43–56. https://doi.org/10.1515/chem-2019-0005
Hendrickx R, Johansson JG, Lohmann C et al (2013) Identification of novel substrates and structure-activity relationship of cellular uptake mediated by human organic cation transporters 1 and 2. J Med Chem 56:7232–7242. https://doi.org/10.1021/jm400966v
Wu F, Zhou Y, Li L et al (2020) Computational approaches in preclinical studies on drug discovery and development. Front Chem 8:1–32. https://doi.org/10.3389/fchem.2020.00726
Teli DM, Shah MB, Chhabria MT (2021) In silico screening of natural compounds as potential inhibitors of SARS-CoV-2 main protease and spike RBD: targets for COVID-19. Front Mol Biosci 7:1–25. https://doi.org/10.3389/fmolb.2020.599079
Namboodiri HV, Bukhtiyarova M, Ramcharan J et al (2010) Analysis of imatinib and sorafenib binding to p38α Compared with c-Abl and b-Raf provides structural insights for understanding the selectivity of inhibitors targeting the DFG-out form of protein kinases. Biochemistry 49:3611–3618. https://doi.org/10.1021/bi100070r
Liu Y, Gray NS (2006) Rational design of inhibitors that bind to inactive kinase conformations. Nat Chem Biol 2:358–364. https://doi.org/10.1038/nchembio799
Li GR, Wang HB, Qin GW et al (2008) Acacetin, a natural flavone, selectively inhibits human atrial repolarization potassium currents and prevents atrial fibrillation in dogs. Circulation 117:2449–2457. https://doi.org/10.1161/CIRCULATIONAHA.108.769554
Zhao N, Dong Q, Fu XX et al (2014) Acacetin blocks Kv1.3 channels and inhibits human T cell activation. Cell Physiol Biochem 34:1359–1372. https://doi.org/10.1159/000366343
Cai Y, Zheng Q, Sun R et al (2020) Recent progress in the study of Artemisiae Scopariae Herba (Yin Chen), a promising medicinal herb for liver diseases. Biomed Pharmacother 130:1–15. https://doi.org/10.1016/j.biopha.2020.110513
Heimfarth L, da Nascimento LDS, Amazonas da Silva MJ et al (2021) Neuroprotective and anti-inflammatory effect of pectolinarigenin, a flavonoid from Amazonian Aegiphila integrifolia (Jacq.), against lipopolysaccharide-induced inflammation in astrocytes via NFκB and MAPK pathways. Food Chem Toxicol 157:1–11. https://doi.org/10.1016/j.fct.2021.112538
Serino E, Chahardoli A, Badolati N et al (2021) Salvigenin, a trimethoxylated flavone from Achillea Wilhelmsii c. Koch, exerts combined lipid-lowering and mitochondrial stimulatory effects. Antioxidants 10:1–18. https://doi.org/10.3390/antiox10071042
Wang Q, Liao XL, Xiang C, Yang J (2017) A practical synthesis of the flavone, scutellarein. J Chem Res 41:157–159. https://doi.org/10.3184/174751917X14873588907765
Hevener KE, Zhao W, Ball DM et al (2009) Validation of molecular docking programs for virtual screening against dihydropteroate synthase. J Chem Inf Model 49:444–460
Patan A, Aanandhi MV, Gopinath P (2023) Molecular dynamics simulation approach of hybrid chalcone-thiazole complex derivatives for DNA gyrase B inhibition: lead generation. RSC Adv 13:24291–24308. https://doi.org/10.1039/d3ra00732d
Singh M, Haque MA, Tikhomirov AS et al (2022) Computational and biophysical characterization of heterocyclic derivatives of anthraquinone against human aurora kinase A. ACS Omega 7:39603–39618. https://doi.org/10.1021/acsomega.2c00740
Uhomoibhi JO, Idowu KA, Shode FO, Sabiu S (2022) Molecular modeling identification of potential drug candidates from selected African plants against SARS-CoV-2 key druggable proteins. Sci African 17:1–11. https://doi.org/10.1016/j.sciaf.2022.e01279
Aziz M, Ejaz SA, Zargar S et al (2022) Deep learning and structure-based virtual screening for drug discovery against NEK7: a novel target for the treatment of cancer. Molecules 27. https://doi.org/10.3390/molecules27134098
Samad A, Ajmal A, Mahmood A et al (2023) Identification of novel inhibitors for SARS-CoV-2 as therapeutic options using machine learning-based virtual screening, molecular docking and MD simulation. Front Mol Biosci 10:1–17. https://doi.org/10.3389/fmolb.2023.1060076
Kumar S, Abbas F, Ali I et al (2023) Integrated network pharmacology and in-silico approaches to decipher the pharmacological mechanism of Selaginella tamariscina in the treatment of non-small cell lung cancer. Phytomed Plus 3:1–35. https://doi.org/10.1016/j.phyplu.2023.100419
Ahmed A, Saeed A, Ejaz SA et al (2022) Novel adamantyl clubbed iminothiazolidinones as promising elastase inhibitors: design, synthesis, molecular docking, ADMET and DFT studies. RSC Adv 12:11974–11991. https://doi.org/10.1039/d1ra09318e
Dennington R, Keith TA, Millam JM (2019) GaussView 6.1.1 | Gaussian.com. 1–16
El-Saady AA, Roushdy N, Farag AAM et al (2023) Exploring the molecular spectroscopic and electronic characterization of nanocrystalline metal-free phthalocyanine: a DFT investigation. Springer US
Abid S, Khalid M, Sagir M et al (2023) Exploration of nonlinear optical enhancement in acceptor-π-donor indacenodithiophene based derivatives via structural variations: a DFT approach. RSC Adv 13:28076–28088. https://doi.org/10.1039/d3ra04858f
Marinho MM, Almeida-Neto FWQ, Marinho EM et al (2021) Quantum computational investigations and molecular docking studies on amentoflavone. Heliyon 7:1–13. https://doi.org/10.1016/j.heliyon.2021.e06079
Srivastava R (2021) Theoretical studies on the molecular properties, toxicity, and biological efficacy of 21 new chemical entities. ACS Omega 6:24891–24901. https://doi.org/10.1021/acsomega.1c03736
Puthanveedu V, Muraleedharan K (2022) Phytochemicals as potential inhibitors for COVID – 19 revealed by molecular docking, molecular dynamic simulation and DFT studies. Struct Chem 33:1423–1443. https://doi.org/10.1007/s11224-022-01982-4
Kumar S, Abbas F, Ali I et al (2023) Integrated network pharmacology and in-silico approaches to decipher the pharmacological mechanism of Selaginella tamariscina in the treatment of non-small cell lung cancer. Phytomed Plus 3(1–35):100419. https://doi.org/10.1016/j.phyplu.2023.100419
Acknowledgements
The authors would like to acknowledge Amity Institute of Biotechnology & Amity Institute of Pharmacy, Amity University Rajasthan, Jaipur for providing the needed facilities. The authors extend their appreciation to the Deanship of Scientific Research & Innovation, Ministry of Education in Saudi Arabia for the financial support through the project number IFP-IMSUI-2023107. The authors also appreciate the Deanship of Scientific Research at Imam Mohammad Ibn Saud Islamic University (IMSIU) for supporting this project.
Author information
Authors and Affiliations
Contributions
NY & VK conceptualized and designed this study, methodology, and was instrumental in investigation, analysis, drafting the entire manuscript with visualizations. AAC, SK, PVA, SSL, & PKS revised and edited the manuscript in its final form. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethics approval
Not applicable.
Consent to participate
Yes. All authors agreed to participate in this research.
Consent for publication
Yes. All authors have approved the last version of the manuscript for its submission.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yasmeen, N., Chaudhary, A.A., Khan, S. et al. Antiangiogenic potential of phytochemicals from Clerodendrum inerme (L.) Gaertn investigated through in silico and quantum computational methods. Mol Divers (2024). https://doi.org/10.1007/s11030-024-10846-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11030-024-10846-4