Abstract
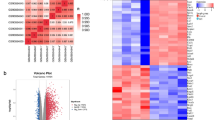
Spinal cord injury (SCI) is a devastating neurological disease with no cure that usually results in irreversible loss of sensory and voluntary motor functions below the injury site. We conducted an in-depth bioinformatics analysis combining the gene expression omnibus spinal cord injury database and the autophagy database and found that the expression of the autophagy gene CCL2 was significantly upregulated and the PI3K/Akt/mTOR signaling pathway was activated after SCI. The results of the bioinformatics analysis were verified by constructing animal and cellular models of SCI. We then used small interfering RNA to inhibit the expression of CCL2 and PI3K to inhibit and activate the PI3K/Akt/mTOR signaling pathway; western blot, immunofluorescence, monodansylcadaverine, and cell flow techniques were used to detect the expression of key proteins involved in downstream autophagy and apoptosis. We found that when PI3K inhibitors were activated, apoptosis decreased, the levels of autophagy-positive proteins LC3-I/LC3-II and Bcl-1 increased, the levels of autophagy-negative protein P62 decreased, the levels of pro-apoptotic proteins Bax and caspase-3 decreased, the levels of the apoptosis-inhibiting protein Bcl-2 increased. In contrast, when a PI3K activator was used, autophagy was inhibited, and apoptosis was increased. This study revealed the effect of CCL2 on autophagy and apoptosis after SCI through the PI3K/Akt/mTOR signaling pathway. By blocking the expression of the autophagy-related gene CCL2, the autophagic protective response can be activated, and apoptosis can be inhibited, which may be a promising strategy for the treatment of SCI.







Similar content being viewed by others
Data Availability
All data is real and guarantee the validity of results.
References
Abbaszadeh F, Fakhri S, Khan H (2020) Targeting apoptosis and autophagy following spinal cord injury: therapeutic approaches to polyphenols and candidate phytochemicals. Pharmacol Res 160:105069
Alizadeh A, Dyck SM, Karimi-Abdolrezaee S (2019) Traumatic spinal cord Injury: an overview of pathophysiology, Models and Acute Injury Mechanisms. Front Neurol 10::282
Allen A (1991) Surgery of experimental lesion of spinal cord equivalent to crush injury of fracture dislocation of spinal columm. JAMA 57
Badhiwala JH, Ahuja CS, Fehlings MG (2018) Time is spine: a review of translational advances in spinal cord injury. J Neurosurg Spine 30:1–18
Brown H, Chung M, Uffing A, Batistatou N, Tsang T, Doskocil S, Mao W, Willbold D, Bast RC Jr, Lu Z, Weiergraber OH, Kritzer JA (2022) Structure-based design of stapled peptides that bind GABARAP and inhibit autophagy. J Am Chem Soc 144:14687–14697
Chamankhah M, Eftekharpour E, Karimi-Abdolrezaee S, Boutros PC, San-Marina S, Fehlings MG (2013) Genome-wide gene expression profiling of stress response in a spinal cord clip compression injury model. BMC Genomics 14:583
Chen C, Chen Q, Mao Y, Xu S, Xia C, Shi X, Zhang JH, Yuan H, Sun X (2010) Hydrogen-rich saline protects against spinal cord injury in rats. Neurochem Res 35:1111–1118
Chittaranjan S, Bortnik S, Dragowska WH, Xu J, Abeysundara N, Leung A, Go NE, DeVorkin L, Weppler SA, Gelmon K, Yapp DT, Bally MB, Gorski SM (2014) Autophagy inhibition augments the anticancer effects of epirubicin treatment in anthracycline-sensitive and -resistant triple-negative breast cancer. Clin Cancer Res 20:3159–3173
Deng Y, Zhu L, Cai H, Wang G, Liu B (2018) Autophagic compound database: a resource connecting autophagy-modulating compounds, their potential targets and relevant diseases. Cell Prolif 51:e12403
Di Giovanni S, Knoblach SM, Brandoli C, Aden SA, Hoffman EP, Faden AI (2003) Gene profiling in spinal cord injury shows role of cell cycle in neuronal death. Ann Neurol 53:454–468
Donnelly DJ, Longbrake EE, Shawler TM, Kigerl KA, Lai W, Tovar CA, Ransohoff RM, Popovich PG (2011) Deficient CX3CR1 signaling promotes recovery after mouse spinal cord injury by limiting the recruitment and activation of Ly6Clo/iNOS + macrophages. J Neurosci 31:9910–9922
Fan B, Wei Z, Yao X, Shi G, Cheng X, Zhou X, Zhou H, Ning G, Kong X, Feng S (2018) Microenvironment Imbalance of spinal cord Injury. Cell Transpl 27:853–866
Fang WB, Yao M, Jokar I, Alhakamy N, Berkland C, Chen J, Brantley-Sieders D, Cheng N (2015) The CCL2 chemokine is a negative regulator of autophagy and necrosis in luminal B breast cancer cells. Breast Cancer Res Treat 150:309–320
Fang S, Zhong L, Wang AQ, Zhang H, Yin ZS (2021) Identification of regeneration and hub genes and pathways at different time points after spinal cord Injury. Mol Neurobiol 58:2643–2662
Figueroa JD, Serrano-Illan M, Licero J, Cordero K, Miranda JD, De Leon M (2016) Fatty acid binding protein 5 modulates Docosahexaenoic Acid-Induced recovery in rats undergoing spinal cord Injury. J Neurotrauma 33:1436–1449
Fraiberg M, Elazar Z (2020) Genetic defects of autophagy linked to disease. Prog Mol Biol Transl Sci 172:293–323
Galluzzi L, Bravo-San Pedro JM, Blomgren K, Kroemer G (2016) Autophagy in acute brain injury. Nat Rev Neurosci 17:467–484
Gao G, Duan Y, Chang F, Zhang T, Huang X, Yu C (2022) METTL14 promotes apoptosis of spinal cord neurons by inducing EEF1A2 m6A methylation in spinal cord injury. Cell Death Discov 8:15
Greene LA, Tischler AS (1976) Establishment of a noradrenergic clonal line of rat adrenal pheochromocytoma cells which respond to nerve growth factor. Proc Natl Acad Sci U S A 73:2424–2428
Handa K, Kanno H, Matsuda M, Sugaya T, Murakami T, Prudnikova M, Ozawa H, Itoi E (2020) Chaperone-mediated autophagy after spinal cord Injury. J Neurotrauma 37:1687–1695
Hänzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14:7
He M, Ding Y, Chu C, Tang J, Xiao Q, Luo ZG (2016) Autophagy induction stabilizes microtubules and promotes axon regeneration after spinal cord injury. Proc Natl Acad Sci U S A 113:11324–11329
Heras-Sandoval D, Pérez-Rojas JM, Hernández-Damián J, Pedraza-Chaverri J (2014) The role of PI3K/AKT/mTOR pathway in the modulation of autophagy and the clearance of protein aggregates in neurodegeneration. Cell Signal 26:2694–2701
Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T, Kominami E, Ohsumi Y, Yoshimori T (2000) LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. Embo j 19:5720–5728
Kanno H, Ozawa H, Sekiguchi A, Itoi E (2009) Spinal cord injury induces upregulation of beclin 1 and promotes autophagic cell death. Neurobiol Dis 33:143–148
Lee YL, Shih K, Bao P, Ghirnikar RS, Eng LF (2000) Cytokine chemokine expression in contused rat spinal cord. Neurochem Int 36:417–425
Levine B, Kroemer G (2019) Biological Functions of Autophagy genes: a Disease Perspective. Cell 176:11–42
Li XG, Du JH, Lu Y, Lin XJ (2019a) Neuroprotective effects of rapamycin on spinal cord injury in rats by increasing autophagy and akt signaling. Neural Regen Res 14:721–727
Li Z, Chen T, Cao Y, Jiang X, Lin H, Zhang J, Chen Z (2019b) Pros and cons: Autophagy in Acute spinal cord Injury. Neurosci Bull 35:941–945
Li Q, Li B, Tao B, Zhao C, Fan B, Wang Q, Sun C, Duan H, Pang Y, Fu X, Feng S (2021) Identification of four genes and biological characteristics associated with acute spinal cord injury in rats integrated bioinformatics analysis. Ann Transl Med 9:570
Liu S, Sarkar C, Dinizo M, Faden AI, Koh EY, Lipinski MM, Wu J (2015) Disrupted autophagy after spinal cord injury is associated with ER stress and neuronal cell death. Cell Death Dis 6:e1582
McDonald JW, Sadowsky C (2002) Spinal-cord injury. The Lancet 359:417–425
Mizushima N, Levine B (2020) Autophagy in Human Diseases. N Engl J Med 383:1564–1576
Nikoletopoulou V, Papandreou ME, Tavernarakis N (2015) Autophagy in the physiology and pathology of the central nervous system. Cell Death Differ 22:398–407
Oyinbo CA (2011) Secondary injury mechanisms in traumatic spinal cord injury: a nugget of this multiply cascade. Acta Neurobiol Exp (Wars) 71:281–99
Ravanan P, Srikumar IF, Talwar P (2017) Autophagy: the spotlight for cellular stress responses. Life Sci 188:53–67
Roca H, Varsos ZS, Mizutani K, Pienta KJ (2008) CCL2, survivin and autophagy: new links with implications in human cancer. Autophagy 4:969–971
Roca H, Varsos ZS, Sud S, Craig MJ, Ying C, Pienta KJ (2009) CCL2 and interleukin-6 promote survival of human CD11b + peripheral blood mononuclear cells and induce M2-type macrophage polarization. J Biol Chem 284:34342–34354
Saraswat Ohri S, Bankston AN, Mullins SA, Liu Y, Andres KR, Beare JE, Howard RM, Burke DA, Riegler AS, Smith AE, Hetman M, Whittemore SR (2018) Blocking autophagy in oligodendrocytes limits functional recovery after spinal cord Injury. J Neurosci 38:5900–5912
Stavoe AKH, Holzbaur ELF (2019) Autophagy in neurons. Annu Rev Cell Dev Biol 35:477–500
Tang P, Hou H, Zhang L, Lan X, Mao Z, Liu D, He C, Du H, Zhang L (2014) Autophagy reduces neuronal damage and promotes locomotor recovery via inhibition of apoptosis after spinal cord injury in rats. Mol Neurobiol 49:276–287
van Beek N, Klionsky DJ, Reggiori F (2018) Genetic aberrations in macroautophagy genes leading to diseases. Biochim Biophys Acta Mol Cell Res 1865:803–816
Wang YC, Zhang S, Du TY, Wang B, Sun XQ (2010) Hyperbaric oxygen preconditioning reduces ischemia-reperfusion injury by stimulating autophagy in neurocyte. Brain Res 1323:149–151
Wang J, Wu GS (2014) Role of autophagy in cisplatin resistance in ovarian cancer cells. J Biol Chem 289:17163–17173
Wang Z, Zhou L, Zheng X, Chen G, Pan R, Li J, Liu W (2017) Autophagy protects against PI3K/Akt/mTOR-mediated apoptosis of spinal cord neurons after mechanical injury. Neurosci Lett 656:158–164
Wang C, Wang Q, Lou Y, Xu J, Feng Z, Chen Y, Tang Q, Zheng G, Zhang Z, Wu Y, Tian N, Zhou Y, Xu H, Zhang X (2018) Salidroside attenuates neuroinflammation and improves functional recovery after spinal cord injury through microglia polarization regulation. J Cell Mol Med 22:1148–1166
Wang Z, Zheng S, Gu Y, Zhou L, Lin B, Liu W (2020) 4-PBA enhances autophagy by inhibiting endoplasmic reticulum stress in recombinant human Beta nerve growth Factor-Induced PC12 cells after mechanical Injury via PI3K/AKT/mTOR signaling pathway. World Neurosurg 138:e659–e664
Wiatrak B, Kubis-Kubiak A, Piwowar A, Barg E (2020) PC12 Cell Line: Cell Types, Coating of Culture Vessels, Differentiation and Other Culture Conditions. Cells 9
Xu W, Wei Q, Han M, Zhou B, Wang H, Zhang J, Wang Q, Sun J, Feng L, Wang S, Ye Y, Wang X, Zhou J, Jin H (2018) CCL2-SQSTM1 positive feedback loop suppresses autophagy to promote chemoresistance in gastric cancer. Int J Biol Sci 14:1054–1066
Xu P, Zhang F, Chang MM, Zhong C, Sun CH, Zhu HR, Yao JC, Li ZZ, Li ST, Zhang WC, Sun GD (2021) Recruitment of gammadelta T cells to the lesion via the CCL2/CCR2 signaling after spinal cord injury. J Neuroinflammation 18:64
Yu D, Li M, Ni B, Kong J, Zhang Z (2013) Induction of neuronal mitophagy in acute spinal cord injury in rats. Neurotox Res 24:512–522
Zhang HY, Wang ZG, Wu FZ, Kong XX, Yang J, Lin BB, Zhu SP, Lin L, Gan CS, Fu XB, Li XK, Xu HZ, Xiao J (2013) Regulation of autophagy and ubiquitinated protein accumulation by bFGF promotes functional recovery and neural protection in a rat model of spinal cord injury. Mol Neurobiol 48:452–464
Zhang LH, Yang AJ, Wang M, Liu W, Wang CY, Xie XF, Chen X, Dong JF, Li M (2016) Enhanced autophagy reveals vulnerability of P-gp mediated epirubicin resistance in triple negative breast cancer cells. Apoptosis 21:473–488
Zhang Q, Zhu C, Li X, Shi Y, Zhang Z (2021) CCR2 downregulation attenuates spinal cord injury by suppressing inflammatory monocytes. Synapse 75:e22191
Zhao YG, Zhang H (2019) Core autophagy genes and human diseases. Curr Opin Cell Biol 61:117–125
Zhou K, Chen H, Xu H, Jia X (2021) Trehalose Augments Neuron Survival and Improves Recovery from Spinal Cord Injury via mTOR-Independent Activation of Autophagy. Oxid Med Cell Longev 2021:8898996
Zhou KL, Zhou YF, Wu K, Tian NF, Wu YS, Wang YL, Chen DH, Zhou B, Wang XY, Xu HZ, Zhang XL (2015) Stimulation of autophagy promotes functional recovery in diabetic rats with spinal cord injury. Sci Rep 5:17130
Funding
This study was supported by the National Natural Science Foundation of China, No. 81871785.
And Hefei Second People’s Hospital Doctoral Special Fund Project Grant No. 2022bszx08.
Author information
Authors and Affiliations
Contributions
Sheng Fang: Conceived the original idea and designed the outlines of the study. Hao Tang: Helped do some experiments, Ming-zhi Li, Jian-Jun Chu Helped collect, organize the data. Zong-Sheng Yin, Qi-yu Jia: Aided in revising the manuscript. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no conflict of interest with the Gene Expression Omnibus (GEO) database used in this study.
Ethics approval
This study was approved by the Ethics Committee of Anhui Medical University (20180402).
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fang, S., Tang, H., Li, MZ. et al. Identification of the CCL2 PI3K/Akt axis involved in autophagy and apoptosis after spinal cord injury. Metab Brain Dis 38, 1335–1349 (2023). https://doi.org/10.1007/s11011-023-01181-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11011-023-01181-y