Abstract
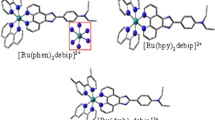
Mononuclear Ru(II)Polypyridyl complexes of type [Ru(A)2BPIIP] (ClO4)2.2H2O, where BPIIP = 2-(3-(4-bromophenyl)isoxazole-5-yl)-1 H-imidazo [4,5-f] [1, 10] phenanthroline and A = bpy = bipyridyl (1), phen = 1,10 Phenanthroline (2), dmb = 4, 4' -dimethyl 2, 2'- bipyridine (3) & dmp = 4,4'-dimethyl-1,10 –Ortho Phenanthroline (4), were synthesized and their antibacterial activity were examined. The synthesized complexes were characterized and their interaction with DNA was studied using Computational and Biophysical methods (Absorption, emission methods, and viscosity). Molecular modelling studies were carried out for molecular geometry and electronic properties (Frontier molecular orbital HOMO—LUMO). The electrostatic potential surface contours for the complexes were analysed to give their nucleophilic level of sensitivity. The study reveals that the Ru(II) Polypyridyl complexes bind to DNA preponderantly by intercalation. The results recommend that the phen and dmp complex have more effective binding ability than the bpy and dmb, indicating the role of the ancillary ligand in determining their specificity for DNA binding. Further molecular docking studies suggested an octahedral geometry and bind to DNA by preferential binding to Guanine. The docking study additionally sustains the binding constant data acquired with the absorption and emission techniques.The results reveal that the nature of the ancillary Ligand plays a considerable role for the intercalation of the Ru(II) polypyridyl complex to DNA, which subsequently influences the antibacterial activity. Biological studies conducted on Gram‐Negative (E.coli and K.pneumonia) and Gram-Positive (S. aureus and E. faecalis) bacteria establish that complex 1 and 2 were considerably active against S. aureus and E. coli.










Similar content being viewed by others
Data Availability
Data generated or analysed during this study are included in this published article [and its supplementary information files].
References
Rasko DA, Sperandio V (2010) Anti-virulence strategies to combat bacteria-mediated disease. Nat Rev Drug Discov 9:117–128. https://doi.org/10.1038/nrd3013
Blair JMA, Webber MA, Baylay AJ et al (2015) Molecular mechanisms of antibiotic resistance. Nat Rev Microbiol 13:42–51. https://doi.org/10.1038/nrmicro3380
Spellberg B, Blaser M, Guidos RJ et al (2011) Combating Antimicrobial Resistance: Policy Recommendations to Save Lives. Clin Infect Dis 52:S397-428. https://doi.org/10.1093/cid/cir153
Boucher HW, Talbot GH, Bradley JS et al (2009) Bad Bugs, No Drugs: No ESKAPE! An Update from the Infectious Diseases Society of America. Clin Infect Dis 48:1–12. https://doi.org/10.1086/595011
Rice LB (2008) Federal Funding for the Study of Antimicrobial Resistance in Nosocomial Pathogens: No ESKAPE. J Infect Dis 197:1079–1081. https://doi.org/10.1086/533452
Walsh CT, Wencewicz TA (2014) Prospects for new antibiotics: a molecule-centered perspective. J Antibiot (Tokyo) 67:7–22. https://doi.org/10.1038/ja.2013.49
World Health Organization. Antimicrobial resistance (2018) October 13, 2019. http://www.who.int/news-room/fact-sheets/detail/antimicrobial-resistance. Accessed 13 Sept 2021
Antimicrobial resistance is rising in India, says ICMR report, Sep 5, 2021. https://timesofindia.indiatimes.com/city/mumbai/antimicrobial-resistance-is-rising-in-india-says-icmr-report/articleshow/85913195.cms. Accessed 14 Sept 2021
Pendleton JN, Gorman SP, Gilmore BF (2013) Clinical relevance of the ESKAPE pathogens. Expert Rev Anti Infect Ther 11:297–308. https://doi.org/10.1586/eri.13.12
De Oliveira DMP, Forde BM, Kidd TJ et al (2020) Antimicrobial resistance in ESKAPE pathogens. Clin Microbiol Rev 33:e00181-e219
Mjos KD, Orvig C (2014) Metallodrugs in Medicinal Inorganic Chemistry. Chem Rev 114:4540–4563. https://doi.org/10.1021/cr400460s
Byrne A, Burke CS, Keyes TE (2016) Precision targeted ruthenium( <scp>ii</scp> ) luminophores; highly effective probes for cell imaging by stimulated emission depletion (STED) microscopy. Chem Sci 7:6551–6562. https://doi.org/10.1039/C6SC02588A
Franco MM, Andrea B, James DH, Marcel R, Stephen BH (2015) Platinum Antitumor Complexes: 50 Years Since Barnett Rosenberg’s Discovery. J Clin Oncol 33(35):4219–4226. https://doi.org/10.1200/JCO.2015.60.7481
Fricker SP (2007) Metal based drugs: from serendipity to design. Dalt Trans 43:4903–4917. https://doi.org/10.1039/b705551j
von AR Katrizky CW Rees (1984) Comprehensive Heterocyclic Chemistry. Herausgegeben Pergamon Press, Oxford, The Structure, Reactions, Synthesis and Uses of Heterocyclic Compounds
Moucheron C (2009) From cisplatin to photoreactive Ru complexes: targeting DNA for biomedical applications. New J Chem 33:235–245. https://doi.org/10.1039/B817016A
Gill MR, Garcia-Lara J, Foster SJ et al (2009) A ruthenium(II) polypyridyl complex for direct imaging of DNA structure in living cells. Nat Chem 1:662–667. https://doi.org/10.1038/nchem.406
Long EC (2009) Metal Complex−DNA Interactions. J Am Chem Soc 131:14124–14125. https://doi.org/10.1021/ja907261x
Thota S, Rodrigues DA, Crans DC, Barreiro EJ (2018) Ru(II) Compounds: Next-Generation Anticancer Metallotherapeutics? J Med Chem 61:5805–5821. https://doi.org/10.1021/acs.jmedchem.7b01689
Gao F, Chao H, Zhou F et al (2006) DNA interactions of a functionalized ruthenium(II) mixed-polypyridyl complex [Ru(bpy)2 ppd]2+. J Inorg Biochem 100:1487–1494. https://doi.org/10.1016/j.jinorgbio.2006.04.008
Vyas NA, Ramteke SN, Kumbhar AS et al (2016) Ruthenium(II) polypyridyl complexes with hydrophobic ancillary ligand as Aβ aggregation inhibitors. Eur J Med Chem 121:793–802. https://doi.org/10.1016/j.ejmech.2016.06.038
Boerner LJ, Zaleski JM (2005) Metal complex–DNA interactions: from transcription inhibition to photoactivated cleavage. Curr Opin Chem Biol 9:135–144. https://doi.org/10.1016/j.cbpa.2005.02.010
Gupta RK, Pandey R, Sharma G et al (2013) DNA Binding and Anti-Cancer Activity of Redox-Active Heteroleptic Piano-Stool Ru(II), Rh(III), and Ir(III) Complexes Containing 4-(2-Methoxypyridyl)phenyldipyrromethene. Inorg Chem 52:3687–3698. https://doi.org/10.1021/ic302196v
Coury JE, Anderson JR, McFail-Isom L et al (1997) Scanning Force Microscopy of Small Ligand−Nucleic Acid Complexes: Tris( o -phenanthroline)ruthenium(II) as a Test for a New Assay. J Am Chem Soc 119:3792–3796. https://doi.org/10.1021/ja9623774
Cook NP, Torres V, Jain D, Martí AA (2011) Sensing Amyloid-β Aggregation Using Luminescent Dipyridophenazine Ruthenium(II) Complexes. J Am Chem Soc 133:11121–11123. https://doi.org/10.1021/ja204656r
Liu J, Mei WJ, Lin LJ et al (2004) Electronic effects on the interactions of complexes [Ru(phen)2(p-L)]2+ (L=MOPIP, HPIP, and NPIP) with DNA. Inorganica Chim Acta 357:285–293. https://doi.org/10.1016/S0020-1693(03)00478-X
Trondl R, Heffeter P, Kowol CR et al (2014) NKP-1339, the first ruthenium-based anticancer drug on the edge to clinical application. Chem Sci 5:2925–2932. https://doi.org/10.1039/C3SC53243G
Blazevic A, Hummer AA, Heffeter P et al (2017) Electronic State of Sodium trans-[Tetrachloridobis(1H-indazole)ruthenate(III)] (NKP-1339) in Tumor, Liver and Kidney Tissue of a SW480-bearing Mouse. Sci Rep 7:40966. https://doi.org/10.1038/srep40966
Pages BJ, Ang DL, Wright EP, Aldrich-Wright JR (2015) Metal complex interactions with DNA. Dalt Trans 44:3505–3526. https://doi.org/10.1039/C4DT02700K
Comba P, Morgen M, Wadepohl H (2013) Tuning of the Properties of Transition-Metal Bispidine Complexes by Variation of the Basicity of the Aromatic Donor Groups. Inorg Chem 52(11):6481–6501. https://doi.org/10.1021/ic4004214
Comba P (2021) 'Modeling of Molecular Properties' In: Comprehensive Coordination Chemistry 3. Comba (ed), Elsevier, Wiley-VCH 107–121
Araya G, Benites J, Reyes JS, Marcoleta AE, Valderrama JA, Lagos R, Monasterio O (2019) Inhibition of Escherichia coli and Bacillus subtilis FtsZ Polymerization and Bacillus subtilis Growth by Dihydroxynaphtyl Aryl Ketones. Front Microbiol 10:1225. https://doi.org/10.3389/fmicb.2019.01225
Salvà-Serra F, Gomila M, Svensson-Stadler L, Busquets A, Jaén-Luchoro D, Karlsson R, Moore ER (2018) A protocol for extraction and purification of high-quality and quantity bacterial DNA applicable for genome sequencing: a modified version of the Marmur procedure. Protocol Exchange. https://doi.org/10.1038/protex.2018.084
Vijayalakshmi R, Kanthimathi M, Subramanian V, Nair B (2000) Interaction of DNA with [Cr(Shiff base)(H2O)2]ClO4. Biochim Biophys Acta 1475:157−162. https://doi.org/10.1016/S0304-4165(00)00063-5
Zheng RH, Guo HC, Jiang HJ, Xu KH, Liu BB, Sun WL, Shen ZQ (2010) A new and convenient synthesis of phendiones oxidated by KBr O3/H2SO4 at room temperature. Chinese Chem Lett 21(11):1270–1272. https://doi.org/10.1016/j.cclet.2010.05.030
Goss CA, Abruna HD (1985) Spectral, electrochemical and electrocatalytic properties of 1,10-phenanthroline-5,6-dione complexes of transition metals. Inorg Chem 24(25):4263–4267. https://doi.org/10.1021/ic00219a012
Mohammad S, Ali R, Syed SR, Priyanka S, Ramesh CG, Sushil KD, Arvind M (2016) Design and Synthesis of Some Imidazolyl Derivatives: Photophysical Studies and Application in the Detection of Anions. Open Chem J 3:35–51. https://doi.org/10.2174/1874842201603010035
Wolfe A, Shimer GH, Meehan T (1987) Polycyclic aromatic hydrocarbons physically intercalate into duplex regions of denatured DNA. Biochemistry 26:6392–6396. https://doi.org/10.1021/bi00394a013
McGhee JD, von Hippel PH (1974) Theoretical aspects of DNA-protein interactions: Co-operative and non-co-operative binding of large ligands to a one-dimensional homogeneous lattice. J Mol Biol 86:469–489. https://doi.org/10.1016/0022-2836(74)90031-X
Satyanarayana S, Dabrowiak JC, Chaires JB (1993) Tris(phenanthroline)ruthenium(II) enantiomer interactions with DNA: Mode and specificity of binding. Biochemistry 32:2573–2584. https://doi.org/10.1021/bi00061a015
Satyanarayana S, Dabrowiak JC, Chaires JB (1992) Neither.DELTA.- nor.LAMBDA.-tris(phenanthroline)ruthenium(II) binds to DNA by classical intercalation. Biochemistry 31:9319–9324. https://doi.org/10.1021/bi00154a001
Long EC, Barton JK (1990) On demonstrating DNA intercalation. Acc Chem Res 23(9):271–273. https://doi.org/10.1021/ar00177a001
Barton JK, Raphael AL (1984) Photoactivated stereospecific cleavage of double-helical DNA by cobalt(III) complexes. J Am Chem Soc 106:2466–2468. https://doi.org/10.1021/ja00320a058
Anupama B, Aruna A, Manga V et al (2017) Synthesis, Spectral Characterization, DNA/ Protein Binding, DNA Cleavage, Cytotoxicity, Antioxidative and Molecular Docking Studies of Cu(II)Complexes Containing Schiff Base-bpy/Phen Ligands. J Fluoresc 27:953–965. https://doi.org/10.1007/s10895-017-2030-5
Comba P, Dovalil N, Hanson GR et al (2014) Insights into the Electronic Structure of Cu II Bound to an Imidazole Analogue of Westiellamide. Inorg Chem 53:12323–12336. https://doi.org/10.1021/ic5014413
Bosch S, Comba P, Gahan LR et al (2015) Selective Coordination of Gallium(III), Zinc(II), and Copper(II) by an Asymmetric Dinucleating Ligand: A Model for Metallophosphatases. Chem - A Eur J 21:18269–18279. https://doi.org/10.1002/chem.201503348
Park JW, Al-Saadon R, MacLeod MK, Shiozaki T, Vlaisavljevich B (2020) Multireference Electron Correlation Methods: Journeys along Potential Energy Surfaces. United States: Chemical Rev 120:13. https://doi.org/10.1021/acs.chemrev.9b00496
Bursch M, Hansen A, Pracht P, Kohn J, Grimme S (2020) Theoretical study of conformational energies of transition metal complexes. Phys Chem Phys 23. https://doi.org/10.1039/D0CP04696E
Computational Chemistry Software (2003) Hyperchem 7.5 Evaluation. Hypercube, Inc
Reddy MR, Reddy PV, Kumar YP et al (2014) Synthesis, Characterization, DNA Binding, Light Switch “On and Off”, Docking Studies and Cytotoxicity, of Ruthenium(II) and Cobalt(III) Polypyridyl Complexes. J Fluoresc 24:803–817. https://doi.org/10.1007/s10895-014-1355-6
Nambigari N, Dulapalli R, Mustyala KK et al (2013) Molecular dynamic simulations of Co(III) and Ru(II) polypyridyl complexes and docking studies with dsDNA. Med Chem Res 22:5557–5565. https://doi.org/10.1007/s00044-013-0540-5
Grippo L, Lucidi S (1997) A globally convergent version of the Polak-Ribière conjugate gradient method. Math Program 78:375–391. https://doi.org/10.1007/BF02614362
Howard A, McIver J (1994) HyperChem Computational Chemistry. Hypercube Inc., Waterloo
El-Ghamaz NA, Diab MA, El-Bindary AA et al (2014) Geometrical structure and optical properties of antipyrine Schiff base derivatives. Mater Sci Semicond Process 27:521–531. https://doi.org/10.1016/j.mssp.2014.07.022
El-Sonbati AZ, Diab MA, El-Bindary AA, Morgan SM (2014) Supramolecular spectroscopic and thermal studies of azodye complexes. Spectrochim Acta Part A Mol Biomol Spectrosc 127:310–328. https://doi.org/10.1016/j.saa.2014.02.037
El-Sonbati AZ, Diab MA, El-Bindary AA et al (2016) Molecular docking, DNA binding, thermal studies and antimicrobial activities of Schiff base complexes. J Mol Liq 218:434–456. https://doi.org/10.1016/j.molliq.2016.02.072
Schneidman-Duhovny D, Inbar Y, Nussinov R, Wolfson HJ (2005) PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res 33:W363–W367. https://doi.org/10.1093/nar/gki481
Drew WL, Barry AL, O’Toole R, Sherris JC (1972) Reliability of the Kirby-Bauer Disc Diffusion Method for Detecting Methicillin-Resistant Strains of Staphylococcus aureus. Appl Microbiol 24:240–247. https://doi.org/10.1128/AEM.24.2.240-247.1972
El-Ajaily MM, Abdlseed FA, Ben-Gweirif S (2007) Preparation, characterization and antibacterial activity of some metal ion complexes. E-Journal Chem 4. https://doi.org/10.1155/2007/636290
Srishailam A, Kumar YP, Venkat Reddy P et al (2014) Cellular uptake, cytotoxicity, apoptosis, DNA-binding, photocleavage and molecular docking studies of ruthenium(II) polypyridyl complexes. J Photochem Photobiol B Biol 132:111–123. https://doi.org/10.1016/j.jphotobiol.2014.02.003
Lepecq J-B, Paoletti C (1967) A fluorescent complex between ethidium bromide and nucleic acids. J Mol Biol 27:87–106. https://doi.org/10.1016/0022-2836(67)90353-1
Vuradi RK, Avudoddi S, Putta VR et al (2017) Synthesis, Characterization and Luminescence Sensitivity with Variance in pH, DNA and BSA Binding Studies of Ru(II) Polypyridyl Complexes. J Fluoresc 27:939–952. https://doi.org/10.1007/s10895-017-2029-y
Vuradi RK, Nambigari N, Pendyala P et al (2020) Study of Anti-Apoptotic mechanism of Ruthenium (II)Polypyridyl Complexes via RT-PCR and DNA binding. Appl Organomet Chem 34:e5332. https://doi.org/10.1002/aoc.5332
Chaires JB (1997) Energetics of drug–DNA interactions. Biopolymers 44:201–215. https://doi.org/10.1002/(SICI)1097-0282(1997)44:3%3c201::AID-BIP2%3e3.0.CO;2-Z
Hay BP (1993) Methods for molecular mechanics modeling of coordination compounds. Coord Chem Rev 126:111–236. https://doi.org/10.1016/0010-8545(93)85036-4
Fukui K (1982) Role of Frontier Orbitals in Chemical Reactions. Science (80- ) 218:747–754. https://doi.org/10.1126/science.218.4574.747
Mashiach E, Schneidman-Duhovny D, Andrusier N et al (2008) FireDock: a web server for fast interaction refinement in molecular docking. Nucleic Acids Res 36:W229–W232. https://doi.org/10.1093/nar/gkn186
Taqui Khan B, Annapoorna K (1990) Mixed ligand complexes of ruthenium(III)edta with pyrimidines. Inorganica Chim Acta 171(2):157–163. https://doi.org/10.1016/S0020-1693(00)80426-0
Acknowledgements
The authors AK and NN are thankful to The Head, Department of Chemistry and the Principal, University College of Science, Saifabad, Osmania University, Hyderabad for the facilities to carry out this work.
Funding
No financial support from any agency.
Author information
Authors and Affiliations
Contributions
Navaneetha Nambigari:"Data curation, Conceptualization; Funding acquisition; Methodology; Project administration; Resources; Software; Supervision; Original draft writing, review & editing" Aruna Kodipaka: "Data curation, Formal analysis, Investigation; Validation; Visualization; "., Ravi Kumar Vuradi: Data Interpretation. Praveen Kumar Airva: Biological data curation. Satyanarayana Sirasani: Roles/Writing – original draft Writing.
Corresponding authors
Ethics declarations
Ethical Approval
Not Applicable.
Consent to Participate
Not applicable.
Conflict of Interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nambigari, N., Kodipaka, A., Vuradi, R.K. et al. A Biophysical Study of Ru(II) Polypyridyl Complex, Properties and its Interaction with DNA. J Fluoresc 32, 1211–1228 (2022). https://doi.org/10.1007/s10895-021-02879-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10895-021-02879-x