Abstract
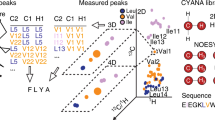
NMR studies of large proteins have gathered much interest in recent years, especially after methyl-transverse relaxation optimized spectroscopy was successfully applied to systems as large as ~1 MDa in molecular weight. However, to fully take advantage of these spectra, there is a need for convenient and robust methods for making resonance assignments rapidly. Here, we present an improved version of our program MAP-XS (methyl assignment prediction from X-ray structure) for the automatic assignment of methyl peaks, based on nuclear Overhauser effects (NOE) correlations and chemical shifts together with available structures. No manual analysis of the NOE data is needed in this new version, which helps to further accelerate the assignment process. A refined algorithm as well as more efficient sampling produces results from single runs of MAP-XSII using unanalyzed NOE data are comparable to those achieved by the old version using manually curated data with every NOE peak correctly attributed to the two related methyl peaks; in addition, checking the results from multiple parallel runs against each other provides an effective mechanism for getting rid of the wrong assignments while keeping the correct ones, which significantly improves the reliability of final assignments. The new program is tested against three different proteins and delivers ~95 % correct assignments; positive results are also achieved for tests using different cut-off distances for NOEs, structures of lower resolutions, and ambiguous residue types.





Similar content being viewed by others
Abbreviations
- TROSY:
-
Transverse relaxation optimized spectroscopy
- NOE:
-
Nuclear overhauser effect
- PRE:
-
Paramagnetic relaxation enhancement
- MBP:
-
Maltose binding protein
- EIN:
-
N-terminal domain of E. coli Enzyme I
- RDC:
-
Residual dipolar coupling
- PCS:
-
Pseudocontact shift
References
Amero C, Dura MA, Noirclerc-Savoye M, Perollier A, Gallet B, Plevin MJ, Vernet T, Franzetti B, Boisbouvier J (2011) A systematic mutagenesis-driven strategy for site-resolved NMR studies of supramolecular assemblies. J Biomol NMR 50(3):229–236. doi:10.1007/s10858-011-9513-5
Chao FA, Shi L, Masterson LR, Veglia G (2012) FLAMEnGO: a fuzzy logic approach for methyl group assignment using NOESY and paramagnetic relaxation enhancement data. J Magn Reson 214:103–110. doi:10.1016/j.jmr.2011.10.008
Fiaux J, Bertelsen EB, Horwich AL, Wuthrich K (2002) NMR analysis of a 900 K GroEL GroES complex. Nature 418(6894):207–211
Gelis I, Bonvin A, Keramisanou D, Koukaki M, Gouridis G, Karamanou S, Economou A, Kalodimos CG (2007) Structural basis for signal-sequence recognition by the translocase motor SecA as determined by NMR. Cell 131(4):756–769. doi:10.1016/j.cell.2007.09.039
Han B, Liu YF, Ginzinger SW, Wishart DS (2011) SHIFTX2: significantly improved protein chemical shift prediction. J Biomol NMR 50(1):43–57. doi:10.1007/s10858-011-9478-4
Hu WD, Namanja AT, Wong S, Chen Y (2012) Selective editing of Val and Leu methyl groups in high molecular weight protein NMR. J Biomol NMR 53(2):113–124. doi:10.1007/s10858-012-9629-2
Pervushin K, Riek R, Wider G, Wuthrich K (1997) Attenuated T2 relaxation by mutual cancellation of dipole–dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc Natl Acad Sci USA 94(23):12366–12371
Ruschak AM, Kay LE (2009) Methyl groups as probes of supra-molecular structure, dynamics and function. J Biomol NMR 46(1):75–87. doi:10.1007/s10858-009-9376-1
Ruschak AM, Religa TL, Breuer S, Witt S, Kay LE (2010) The proteasome antechamber maintains substrates in an unfolded state. Nature 467(7317):868–871. doi:10.1038/nature09444
Salzmann M, Pervushin K, Wider G, Senn H, Wuthrich K (2000) NMR assignment and secondary structure determination of an octameric 110 kDa protein using TROSY in triple resonance experiments. J Am Chem Soc 122(31):7543–7548
Sheppard D, Guo CY, Tugarinov V (2009) 4D H-1-C-13 NMR Spectroscopy for assignments of alanine methyls in large and complex protein structures. J Am Chem Soc 131(4):1364. doi:10.1021/ja808202q
Sprangers R, Kay LE (2007a) Probing supramolecular structure from measurement of methyl (1)H-(13)C residual dipolar couplings. J Am Chem Soc 129(42):12668–12669. doi:10.1021/ja075846i
Sprangers R, Kay LE (2007b) Quantitative dynamics and binding studies of the 20S proteasome by NMR. Nature 445(7128):618–622. doi:10.1038/nature05512
Tugarinov V, Kay LE (2003) Ile, Leu, and Val methyl assignments of the 723-residue malate synthase G using a new labeling strategy and novel NMR methods. J Am Chem Soc 125(45):13868–13878. doi:10.1021/ja030345s
Tugarinov V, Hwang PM, Ollerenshaw JE, Kay LE (2003) Cross-correlated relaxation enhanced 1H[bond]13C NMR spectroscopy of methyl groups in very high molecular weight proteins and protein complexes. J Am Chem Soc 125(34):10420–10428. doi:10.1021/ja030153x
Tugarinov V, Choy WY, Orekhov VY, Kay LE (2005) Solution NMR-derived global fold of a monomeric 82-kDa enzyme. Proc Natl Acad Sci U S A 102(3):622–627. doi:10.1073/pnas.0407792102
Velyvis A, Schachman HK, Kay LE (2009) Assignment of Ile, Leu, and Val Methyl correlations in supra-molecular systems: an application to aspartate transcarbamoylase. J Am Chem Soc 131(45):16534–16543. doi:10.1021/ja906978r
Venditti V, Fawzi NL, Clore GM (2011) Automated sequence- and stereo-specific assignment of methyl-labeled proteins by paramagnetic relaxation and methyl–methyl nuclear overhauser enhancement spectroscopy. J Biomol NMR 51(3):319–328. doi:10.1007/s10858-011-9559-4
Wen J, Zhou P, Wu JH (2012) Efficient acquisition of high-resolution 4-D diagonal-suppressed methyl–methyl NOESY for large proteins. J Magn Reson 218:128–132. doi:10.1016/j.jmr.2012.02.021
Xu YQ, Liu MH, Simpson PJ, Isaacson R, Cota E, Marchant J, Yang DW, Zhang XD, Freemont P, Matthews S (2009) Automated assignment in selectively methyl-labeled proteins. J Am Chem Soc 131(27):9480. doi:10.1021/ja9020233
Acknowledgments
The research is supported by grants from the Wellcome Trust and BBSRC. We are grateful to Dr. Sprangers and Prof. Kay, Dr. Venditti and Prof. Clore, Dr. Wen and Prof. Wu for sharing their data on proteasome, EIN, and MBP respectively. The program can be downloaded from nmr.bc.ic.ac.uk/map-xs/.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xu, Y., Matthews, S. MAP-XSII: an improved program for the automatic assignment of methyl resonances in large proteins. J Biomol NMR 55, 179–187 (2013). https://doi.org/10.1007/s10858-012-9700-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-012-9700-z