Abstract
Analogous to the recently introduced ARTSY method for measurement of one-bond 1H–15N residual dipolar couplings (RDCs) in large perdeuterated proteins, we introduce methods for measurement of base 13C–1H and 15N–1H RDCs in protonated nucleic acids. Measurements are based on quantitative analysis of intensities in 1H–15N and 13C–1H TROSY-HSQC spectra, and are illustrated for a 71-nucleotide adenine riboswitch. Results compare favorably with those of conventional frequency-based measurements in terms of completeness and convenience of use. The ARTSY method derives the size of the coupling from the ratio of intensities observed in two TROSY-HSQC spectra recorded with different dephasing delays, thereby minimizing potential resonance overlap problems. Precision of the RDC measurements is limited by the signal-to-noise ratio, S/N, achievable in the 2D TROSY-HSQC reference spectrum, and is approximately given by 30/(S/N) Hz for 15N–1H and 65/(S/N) Hz for 13C–1H. The signal-to-noise ratio of both 1H–15N and 1H–13C spectra greatly benefits when water magnetization during the experiments is not perturbed, such that rapid magnetization transfer from bulk water to the nucleic acid, mediated by rapid amino and hydroxyl hydrogen exchange coupled with 1H–1H NOE transfer, allows for fast repetition of the experiment. RDCs in the mutated helix 1 of the riboswitch are compatible with nucleotide-specifically modeled, idealized A-form geometry and a static orientation relative to the helix 2/3 pair, which differs by ca 6° relative to the X-ray structure of the native riboswitch.
Similar content being viewed by others
Introduction
It is well recognized that residual dipolar couplings, as measured by solution state NMR, carry important structural information regarding internuclear vector orientation relative to the principal axis system of the molecule’s alignment tensor. This “global” information complements local structural restraints derived from NOE, chemical shift, and J-coupling measurements and has proven useful in biomolecular structure determination (Clore 2000; Prestegard et al. 2000; Bax and Grishaev 2005; Tolman and Ruan 2006). Moreover, RDCs can provide important information on dynamic processes that can remain elusive by conventional relaxation measurements, both in proteins (Peti et al. 2002; Tolman 2002; Bouvignies et al. 2005; Yao et al. 2008) and in nucleic acids (Zhang et al. 2006, 2007). A wide range of different types of RDCs has been measured, including 1H–15N, 1H–13C, 1H–1H, 15N–13C, and 13C–13C (Tjandra and Bax 1997a, b; Yang et al. 1999; Permi et al. 2000; Skrynnikov et al. 2000; Skrynnikov and Kay 2000; Evenas et al. 2001; Boisbouvier et al. 2003; Vijayan and Zweckstetter 2005), and many of these interactions can be measured even in lowly populated transient conformers, not directly visible by conventional NMR (Hansen et al. 2008a, b; Vallurupalli et al. 2008; van Ingen et al. 2009). However, in larger slowly tumbling systems, measurements are often restricted to the backbone amide 1H–15N RDC for proteins, and the imino 1H–15N RDC for base-paired nucleic acids.
A recently introduced method, ARTSY, yields 1H–15N amide RDCs in perdeuterated proteins from analysis of the intensity ratio in two TROSY-HSQC spectra, recorded with different dephasing delays (Fitzkee and Bax 2010), and provides facile measurement of 1DNH RDCs in larger proteins. Here, we present methods based on the ARTSY concept for measuring 1H–15N and 1H–13C base RDCs in nucleic acids, where TROSY-HSQC spectra also yield considerably improved spectral appearance relative to regular 1H–15N and 1H–13C HSQC spectra. The ability to extract RDCs from spectra that have optimized characteristics in terms of spectral resolution makes the ARTSY method particularly attractive for larger oligonucleotides, where resonance overlap associated with spectral crowding and rapid transverse relaxation invariably present a challenge. Because imino and base protons of nucleic acids resonate in reasonably well isolated regions, the effects of 1H–1H dephasing during the amplitude-encoding magnetization transfer step of the TROSY experiment can be minimized by the use of band-selective pulses, obviating the need for perdeuteration, and thereby making the ARTSY amplitude encoding method applicable to the commonly used uniformly 13C/15N-enriched nucleic acids.
A second, important benefit of the use of band-selective pulses throughout the pulse sequence lies in the fact that they leave the water magnetization unperturbed. It was recently shown that band-selective BEST methods, previously developed for increasing the rate at which protein experiments can be repeated for enhanced signal-to-noise per unit of measuring time (Schanda et al. 2006; Lescop et al. 2007), are also applicable to enhancing sensitivity of imino 1H–15N correlation experiments in nucleic acids (Farjon et al. 2009). Similarly, we here show that a comparable enhancement can be obtained for 1H–13C correlation experiments involving the nucleic acid base protons. An added benefit of recording the 1H–13C correlation experiments in H2O rather than D2O relates to the nearly 20% lower viscosity (Cho et al. 1999), which results in longer transverse relaxation times and improved spectral resolution, a detail particularly relevant for the application to larger systems.
Measurement of one-bond 1H–15N and 1H–13C RDCs in nucleic acids to date has been carried out almost exclusively in the frequency domain, by measurement of 15N–{1H} (imino), 13C–{1H} (sugar) or 1H–{13C} (base) doublet splittings, which represent the sum of the one-bond J coupling and the corresponding RDC. Filtering of the downfield and upfield doublet components into separate spectra, either via the inphase-antiphase (IPAP) (Ottiger et al. 1998) or individual multiplet component (IMC) selection methods (Andersson et al. 1998; Tolbert et al. 2010), largely eliminates the increased crowding associated with the doubling of the number of resonances when recording non-decoupled 1H–15N or 1H–13C correlation spectra. However, for both 15N and aromatic base 13C, the upfield doublet component has a strongly enhanced transverse relaxation rate (anti-TROSY effect). The anti-TROSY effect can be mitigated by measuring the position of a resonance which evolves as anti-TROSY magnetization for only a fraction of its evolution time (Yang et al. 1999; Kontaxis et al. 2000; Bhattacharya et al. 2010; Mantylahti et al. 2010), but this also proportionately scales down the size of the measured RDC. For 1H–13C correlation spectra, where the IMC method separates the two 1H–{13C} doublet components, line widths of the two components are comparable because the TROSY effect in the 1H dimension for aromatic systems tends to be very small (Pervushin et al. 1998a). However, measurement of the relative displacement for each pair of peaks is hampered by the presence of substantial broadening in the 1H dimension from unresolved 1H–1H RDCs and difficulties in getting uniformly phased 1H spectra when the 1H–13C RDCs are substantial, such that selection of optimal delays in the TROSY pulse scheme cannot be achieved for all nucleotides simultaneously. Even small phase errors, which can be difficult to recognize visually in crowded spectra, can result in substantial shifts of the apparent peak position, and thereby of the corresponding RDC (Kontaxis et al. 2000).
We demonstrate the measurement of 1H–15N and 1H–13C RDCs from ARTSY spectra for a uniformly 13C/15N-enriched 71-nt riboswitch, RiboA, previously used as a test system for development of methods that determine the structure of such molecules from SAXS and imino 1H–15N RDC data (Wang et al. 2009). The RDCs are fully consistent with the packed pair of stem 2 and 3 helices seen in the X-ray structure and with a static idealized A-form geometry for mutated stem 1, which differs by ca 6° relative to the X-ray structure of the native riboswitch.
Experimental Section
RiboA used in the present study contains three A-form helices, and differs from the molecule used for determining its X-ray structure (Serganov and Patel 2007) by alterations in four base pairs of helix 1, introduced to increase the stability of the molecule: G14–U82 → G14–C82, C15–G81 → G15–C81, U16–A80 → A16–U80, and U17–A79 → A17–U79.
Spectra presented here were recorded on three samples: an isotropic sample containing 0.7 mM RiboA in 95% H2O/5% D2O and a sample containing 0.8 mM RiboA in the same solvent but also including 10 mg/mL Pf1, yielding a solvent 2H quadrupole splitting of 10.1 Hz, as well as a 0.8-mM RiboA sample in the presence of 7 mg/mL Pf1 in D2O, with a solvent 2H quadrupole splitting of 6.9 Hz. All samples additionally contained 2 mM MgCl2, 5 mM adenine, 30 mM KCl, and 10 mM potassium phosphate buffer (pH 6.8). Sample volumes were 330 μL each, contained in Shigemi microcells.
Experimental details regarding the recording of the NMR spectra are listed in the figure legends. Unless stated otherwise, all spectra were recorded at 25°C. Spectra were processed and analyzed using NMRPipe (Delaglio et al. 1995).
Parameters used for building idealized nucleotide-specific A-form RNA were derived from a database of high resolution crystal structures, and details regarding the protocol are included as Supporting Information (SI).
Results and discussion
Measurement of imino 1H–15N RDCs
The ARTSY pulse scheme for measurement of imino 1H–15N RDCs is shown in Fig. 1. With the imino proton resonances mostly very well separated from the sugar and base protons, the pulse scheme is easily executed with pulses that are band-selective for the imino region. For the remainder, the experiment is very similar to the regular TROSY-HSQC experiment (Pervushin et al. 1997, 1998b), and principally differs in two aspects. First, as the J couplings and RDC values in ARTSY experiments are extracted from the transfer efficiency of magnetization from 1H to 15N during the first INEPT step, contributions from 15N Boltzmann magnetization which normally add to the transferred magnetization must be eliminated. This is accomplished both by saturating the 15N magnetization prior to the start of the pulse scheme, and additionally by phase cycling of the 1H 90°ϕ1 pulse of the INEPT transfer element. Second, the duration of the first INEPT period is doubled to 1/1JNH compared to the regular TROSY-HSQC experiment, but the experiment is carried out in two modes: an attenuated mode, where the 1H–{15N} dephasing is active during the full 1/1JNH duration, and a reference mode where the dephasing is active for only half this duration. With the exception of homonuclear RDCs to other imino protons, dephasing due to 1H–1H couplings during the period T is largely eliminated by the band-selective REBURP shape of the refocusing pulse (Geen and Freeman 1991), used at its mid-point. A limited degree of dephasing may remain for couplings to 1H that are partially affected by this REBURP pulse, including protons resonating in the 8–10 ppm region, but the fractional attenuation resulting from such dephasing to a very good approximation is the same in the reference and attenuated experiment and therefore does not impact the intensity ratio, used for extracting the coupling.
ARTSY pulse sequence for measurement of imino 1DNH in nucleic acids. Narrow and wide filled bars represent 90° and 180° pulses, respectively, while wide vertically hatched bars correspond to composite 180° pulses (90x–220y–90x) (Freeman et al. 1980). Composite 15N pulse marked a is only applied for recording the reference spectrum, pulse b only for the attenuated spectrum. Filled half-ellipsoids denote shaped 1H pulses, with e, e*, and r indicating EBURP2, time-reversed EBURP2, and ReBURP pulses, respectively (Geen and Freeman 1991). 1H shaped pulses each have a duration of 1.0 ms (at 900.27 MHz 1H frequency; 1.5 ms at 600 MHz) and are centered at 12.2 ppm. The 15N and 13C pulses are applied at 153 and 160 ppm, respectively. All pulses have phase x unless otherwise indicated. Delays: T = 1/JNH; δ = 1.0 ms (isotropic) and 0.78 ms (Pf1) at 900 MHz (δ = 0.45 ms at 600 MHz for the isotropic sample), empirically optimized to minimize the residual 15N anti-TROSY component for the isotropic and the aligned sample, respectively; τ = 2.8 ms. Note that the last 180° ReBURP pulse is shifted toward the acquisition by δ′ = 90.5 μs relative to the 180° 15N pulses such that 1H chemical shift evolution occurring during the G8 decoding gradient pulse (81 μs) and its recovery delay (100 μs) is refocused at the start of t2. Phase cycling: ϕ1 = y, −y; ϕ2 = y, y, x, x, −y, −y, −x, −x; ϕ3 = y, ϕ4 = y; and ϕrec = y, −y, −x, x, −y, y, x, −x. Gradient pulses: G1,2,3,4,5,6,7,8 = 21.7, 1.19, 28.7, −39.9, 39.9, 19.6, 25.9, and 39.9 G/cm with durations of 650, 2500, 650, 400, 400, 570, 977, and 81 μs, respectively. All gradient pulses are sine bell shaped except for G2 which is rectangular. Quadrature detection is achieved using the echo-antiecho method with the G4 and G5 gradient pulses as well as the ϕ3 and ϕ4 phases inverted and ϕ2 changed to y, y, −x, −x, −y, −y, x, x for every second FID (Pervushin et al., 1998b). Fast repetition of the experiment (interscan delay of 0.6 s) can be used because H2O magnetization remains along z and causes fast recovery of imino 1H z magnetization. Bruker pulse sequence code included as SI
Besides elimination of most 1H–1H RDC dephasing, a secondary benefit of using imino-selective 1H pulses throughout the pulse scheme stems from the rapid recovery of imino magnetization through indirect magnetization transfer from the H2O resonance. Amino NH2 protons, in close spatial proximity to imino protons, rapidly exchange with solvent, while the use of band-selective pulses ensures that the H2O magnetization remains close to its equilibrium Boltzmann value. Therefore, when evaluating the intensity of the ARTSY reference spectrum as a function of the interscan delay (SI Fig. S1), a very short “apparent T1” value of 0.5–0.7 s reflects the rapid magnetization transfer from water to the imino protons. Assuming zero z magnetization for the imino protons at the start of data acquisition, a delay till the start of the next scan of ca 1.3×T1 is optimal for S/N purposes, and a high repetition rate of the experiment can therefore be used (Bax et al. 1983; Farjon et al. 2009). The gain achieved by using band-selective imino proton pulses throughout, rather than non-selective pulses and a separate water suppression scheme such as WATERGATE (Piotto et al. 1992), strongly depends on the spectrometer and parameterization of the pulse sequence. Even when using non-selective pulses, at high magnetic field strengths radiation damping of the water signal will return it back to the z axis much faster than through its relatively slow longitudinal relaxation process. Therefore, depending on the instrument, the strength of gradients used, and the rate at which radiation damping takes place, a relatively short interscan delay may also be applicable for experiments that do not avoid water excitation.
As described previously (Fitzkee and Bax 2010), the magnitude of the sum of the 1JNH and 1H–15N RDC is extracted from the intensity ratio, R, observed for a given imino group in the attenuated (IA) and reference (IR) spectra:
which can be rewritten as:
Assuming identical noise levels, N, in the attenuated and reference spectra, the uncertainty σ in the total (1JNH + 1DNH) splitting extracted from the pair of spectra is given by (Fitzkee and Bax 2010):
which for the common situation where R ≪ 1 is approximately given by
As seen from Eq. 3b, the uncertainty in the derived coupling correlates inversely with the signal to noise ratio, IR/N, in the reference spectrum and samples local minima for T durations where R ≈ 0. The first local minimum is reached when the dephasing period, T, is ~1/1JNH. In principle, use of a longer T value, e.g. 2/1JNH, would reduce σ two-fold for the same IR/N ratio. However, in practice, transverse relaxation will decrease considerably the attainable IR/N ratio. Moreover, for larger RDC values \( \left( {\left| {^{ 1} {\text{D}}_{{{\text{NH}} }} } \right| \ge \sim 10\,{\text{Hz}}} \right) \) the R ≪1 approximation no longer applies for all resonances simultaneously if the 2/1JNH dephasing delay were used, thereby increasing σ (c.f. Eq. 3a), and T ≈ 1/1JNH is therefore used for all experiments.
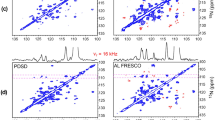
Figure 2 compares the imino region of the reference and attenuated spectra of RiboA, aligned in Pf1. A similar set of spectra was also recorded for a sample in the absence of Pf1, where the molecule is weakly aligned due to its magnetic susceptibility anisotropy (MSA). The three helices of RiboA are packed with their axes in roughly parallel orientations (Serganov and Patel 2007; Wang et al. 2009), causing the long axis of the molecule to align parallel to the magnetic field in the Pf1 suspension, but orthogonal to the magnetic field in the isotropic sample where MSA of the nucleic acid bases dominates the alignment. As a result, in Pf1 suspension |1JNH + 1DNH| values for the Watson–Crick base-paired imino groups are all increased relative to |1JNH|, resulting in an optimal dephasing time T ≈ 1/|1JNH + 1DNH| ≈ 10.1 ms.
Reference (left) and attenuated (right) ARTSY spectra of the imino groups in RiboA, recorded at 900 MHz 1H frequency in the presence of 10 mg/mL Pf1. Spectra were recorded with 16 scans per FID, and the total matrix for the interleaved set of spectra comprised 2 × 200* × 1024* data points, with N* referring to N complex data points, for acquisition times of 48 ms (t1) and 84 ms (t2). Contour levels for the attenuated spectrum are 4 times lower than shown for the reference spectrum. The delay T (Fig. 1) was set to 10.1 ms for the Pf1-aligned sample (11.8 ms for the isotropic sample), such that in the attenuated spectrum most peak intensities are close to the null condition for RiboA. The rate at which imino proton z magnetization recovered in RiboA when the water remains along z was ca 2 s−1, permitting a fast repetition of the experiment (interscan delay of 0.6 s). The total measuring time for the two interleaved spectra was ca 3 h
Measurement of base C6–H6, C8–H8 and C2–H2 RDCs
Except for pyrimidine H5 protons, carbon-attached nucleic acid base protons mostly resonate in a relatively narrow band from 6.5 to 8.5 ppm, again permitting the recording of HSQC-TROSY spectra by using band-selective 1H pulses (Fig. 3). These experiments then are best carried out in H2O rather than in D2O solution, such that magnetization is rapidly transferred from water via the exchangeable amino and hydroxyl protons to the non-exchangeable protons via NOE. At 900 MHz, RiboA base proton signal intensity as observed in the first t1 increment of the C–H ARTSY experiment of Fig. 3 for different interscan delay durations shows rapid recovery of magnetization during this delay (Fig. 4), indicative of apparent base 1H T1 values of ca 1.1 s. In contrast, when the experiment is carried out in D2O, recovery of 1H magnetization is nearly four-fold slower (SI Fig. S2). Moreover, the higher viscosity of D2O relative to H2O increases natural line widths by nearly 20%. The disadvantage of the use of band-selective pulses, used in the C–H ARTSY experiment of Fig. 3, is that protons resonating outside the selected bandwidth will escape detection, as applies to RiboA A21–H2 which resonates at ~5.4 ppm.
ARTSY pulse sequence optimized for simultaneous measurement of base 1DC2–H2, 1DC6–H6 and 1DC8–H8 in H2O. Narrow and wide filled bars represent 90° and 180° pulses, respectively, while wide vertically hatched bars correspond to composite 180° pulses (90x–220y–90x) (Freeman et al. 1980). Composite 13C pulse marked a is only applied for recording the reference spectrum, pulse b only for the attenuated spectrum. Filled half-ellipsoids denote shaped 1H pulses, with e, e*, and r indicating their profiles of EBURP2, time-reverse EBURP2, and ReBURP, respectively. All the 1H shaped pulses are H2/H6/H8 selective and have a duration of 1.35 ms (at 900 MHz), centered at 8.1 and 8.6 ppm for the EBURP2 (and its time-reversed pulse) and ReBURP pulses, respectively. All 13C pulses are centered at 145.9 ppm. The C6 selective ReBURP pulses have a duration of 700 μs (at 226.37 MHz 13C frequency; 50% longer at 150.9 MHz)) and are applied to refocus the chemical shift evolution of C2/C8 of purines and C6 of pyrimidines and to remove the 1 J C6C5 coupling. The 15N carrier is set to 190 ppm. All pulses have phase x unless otherwise indicated. Delays T = 1/1JCH (see also footnote to SI Table S1); δ = 295 μs (empirically optimized for minimizing the anti-TROSY 13C component; note that 1 J CH evolution also occurs for ca 60% of the total durations of the time-reversed EBURP2 and the EBURP2 pulses; the 1H and 13C REBURP pulses in the middle of 2δ delays are left-aligned to reduce the 1 J CH evolution during these two pulses; accordingly, the timing for application of the 90°ϕ4 13C pulse is adjusted such that the 13C chemical shift evolution is refocused during this spin-state-selective coherence transfer); τ = 485 μs (additional 1 J CH evolution for ~80% of the 1H ReBURP pulse duration also occurs). Phase cycling: ϕ1 = y, −y; ϕ2 = x, x, −x, −x; ϕ3 = y; ϕ4 = 4(y), 4(−y); and ϕrec = x, −x, −x, x, −x, x, x, −x. Gradient pulses: G1,2,3,4,5,6,7,8 = 14.7, 1.19, 37.1, −35, 35, 4.9, 7, and 42 G/cm with durations of 1000, 1100, 1000, 500, 500, 79, 251, and 251 μs, respectively. All gradients pulses have a sine bell profile, except for G2 which is rectangular. Quadrature detection is achieved using the echo-antiecho method, inverting the G4 and G5 gradient pulses as well as ϕ3 for every second FID (Kay et al. 1992). Bruker pulse sequence code available as SI
Recovery of H2, H6 and H8 magnetization as a function of interscan delay in the C–H ARTSY experiment of Fig. 3. The 1D spectra shown (left) correspond to the first FID of the 900 MHz ARTSY reference experiment, and have been recorded for interscan delays ranging from 0.3 to 7 s. Recovery of the overlapping 1H resonances is not fully exponential, indicating a range of recovery rates but can be fitted with I(t) = Ieq[1 − 0.82exp(−t/1.08s)] (right panel)
Besides the usual amplitude encoding of the signal during the first INEPT transfer from 1H → 13C, optimized for an average isotropic \( ^{1} {\text{J}}_{{{\text{CH}} }} = 196\,{\text{Hz}} \) by selecting T = 5.1 ms (the impact of different dephasing delays on the measured \( ^{ 1} {\text{J}}_{{{\text{CH}} }} \) value was also evaluated, see below and SI Table S1), the pulse scheme of Fig. 3 employs 13C–13C homonuclear decoupling during the TROSY 13C t1 evolution period. This homonuclear decoupling is accomplished by a combination of a band-selective (C2, C6, C8) REBURP-shaped pulse (Geen and Freeman 1991) followed by a broad-band composite inversion pulse, applied at the mid-point of t1 evolution. The composite and REBURP pulses are separated by a short pulsed field gradient (G4), needed together with G5 for gradient-encoding of the NMR signal (Kay et al. 1992). A second composite inversion pulse is applied at the start of t1 evolution, and serves to invert the phase error incurred by the composite pulse immediately preceding gradient G4 (Levitt and Freeman 1981; Hwang et al. 1997). Instead of the standard non-selective 180° pulse, a 13C REBURP pulse at the midpoint of the 2δ period of the TROSY transfer back to 1H is used to eliminate 13C–13C J dephasing during this period. With no significant TROSY effect achievable for base protons, composite pulse decoupling is used during data acquisition, thereby eliminating small “anti-TROSY” artifacts that result from the fact that the refocusing delay 2τ cannot be simultaneously optimized for the wide range of 1JCH + 1DCH values.
Figure 5 shows a small region of the reference and attenuated ARTSY spectra for the A–C2, A/G–C8 and U/C–C6 carbons in RiboA in the presence of Pf1 (See SI Fig. S3 for the isotropic spectra). Despite the substantial size of RiboA, the vast majority of 1H–13C correlations are sufficiently resolved for accurate quantitative intensity measurement. 13C TROSY line-narrowing for the nucleic acid base carbons is slightly less effective at 900 MHz 1H frequency than at 600 MHz (Boisbouvier et al. 1999; Ying et al. 2006), and spectra of nearly equivalent 13C resolution (but decreased 1H resolution) can be obtained at lower field strengths too (SI Fig. S4).
Small region of the reference (left) and attenuated (right) ARTSY spectra of the 13C6,8–1H moieties in RiboA, recorded at 900 MHz 1H frequency in the presence of 10 mg/mL Pf1. Spectra were recorded with 24 scans per FID, and the total matrix for the interleaved set of spectra comprised 2 × 256* × 1,024* data points, with N* referring to N complex data points, for acquisition times of 49 ms (t1) and 63 ms (t2). Contour levels for the attenuated spectrum are at a level 5 times lower than shown for the reference spectrum. The delay T (Fig. 3) was set to 4.5 ms. Due to the substantial differences in isotropic 1JCH couplings (U/C 1JC6H6 ≈ 180 ± 1 Hz; A/G 1JC8H8 ≈ 215 ± 1 Hz; A 1JC2H2 ≈ 200 ± 1 Hz, all values in the absence of MSA alignment) a close-to-null condition cannot be established for all types of carbons simultaneously and A/G C8–H8 are all negative (red contours). Assignments marked with asterisks correspond to nucleotides outside the helical stem regions. The total measuring time for the two interleaved spectra was ca 5.3 h
Measurement of pyrimidine C5–H5 RDCs
Pyrimidine H5 protons resonate in the 5–6 ppm region, very close to the water resonance. Consequently, the use of pulses that band-selectively excite these protons but leave water magnetization unaffected is not feasible. Instead, we explored two alternate solutions to record ARTSY data for obtaining the C5–H5 RDCs. The first solution (Fig. 6) uses non-selective pulses, with the exception of the band-selective 1H5 pulse at the mid-point of the initial INEPT transfer of magnetization from 1H to 13C, which constitutes the amplitude encoding for the ARTSY experiment. While resulting in nearly full inversion of the H2O and H1′ resonances too, JHH and DHH modulation due to coupling between H5 and H6/H2/H8 protons is effectively eliminated by the band-selective nature of the H5 pulse. An S3E element (Meissner et al. 1997) is used to selectively convert the downfield TROSY 13C5 component to z magnetization, followed by a short delay, Δrd, which serves for return of H2O magnetization to the z axis by means of radiation damping. A subsequent 13C5 evolution period, employing the same combination of a pair of composite inversion pulses, combined with a band-selective 13C5 REBURP pulse as was used for C2/C6/C8 detection (Fig. 3), is followed by a non-selective gradient-enhanced “Rance-Kay” readout scheme (Palmer et al. 1991; Kay et al. 1992). In order to suppress the strong water signal, it is essential to use relatively strong pulsed field gradients, and only on room temperature probeheads equipped with three-axis pulsed field gradients can we suppress the water signal to levels comparable to the thermal noise (Fig. 7a, b). As pointed out above, the strong advantage of recording nucleic acid spectra in the presence of fully relaxed H2O magnetization lies in the rapid recovery of nucleic acid z magnetization, through hydrogen exchange of the C2′ hydroxyl and base NH2 protons, and subsequent NOE to non-exchangeable protons. This allows again for short delays between scans (Fig. 8a), thereby improving signal-to-noise, in addition to the narrower line width obtained in H2O compared to the more viscous D2O alternative. Note that, due to the reliance of the experiment on radiation damping, the water magnetization will attenuate strongly when the repetition rate is too fast, decreasing the z magnetization of the nucleic acid protons (Fig. 8a). Moreover, in particular U–H5 tends to have few or no nearby exchangeable protons and recovery of its magnetization by transfer from H2O typically will involve multiple NOE steps.
C5H5 ARTSY pulse sequence optimized for measurement of pyrimidine 1DC5–H5 in H2O. Narrow and wide filled bars represent 90° and 180° pulses, respectively, while wide vertically hatched bars correspond to composite 180° pulses (90x–220y–90x) (Freeman et al. 1980). Composite 13C pulse marked a is only applied for recording the reference spectrum, pulse b only for the attenuated spectrum. Filled half-ellipsoids denote shaped pulses, all of the ReBURP type. The H5 ReBURP pulse is 1.6 ms (at 747 MHz 1H frequency) and is centered at 4.1 ppm, while all other 1H pulses are applied on resonance with water at 4.77 ppm. The C5 ReBURP pulse has a duration of 800 μs (at 188 MHz 13C frequency). All the 13C pulses are centered at 101.5 ppm, except for the two composite 180° pulses immediately before and after the first t1/2 period, which are applied at 130.2 ppm to invert all the C4, C5, and C6 resonances for optimal 1 J CC decoupling during t1. The 15N carrier is set to 155 ppm. All pulses have phase x unless otherwise indicated. Delays used in the experiments: T = 5.7 and 5.1 ms for the isotropic and Pf1-aligned samples, respectively; ε = 600 μs; Δrd = 10–15 ms, empirically optimized to let H2O magnetization fully return to the +z axis through radiation damping; τ = 1.2 ms; δ = 353 μs. Phase cycling: ϕ1 = y, −y; ϕ2 = 4(45°), 4(225°); ϕ3 = x, x, −x, −x; ϕ4 = x; ϕrec = x, −x, −x, x, −x, x, x, −x. Gradient pulses: G1,2,3,4,5,6,7,8,9,10 in the x/y/z directions with strengths 7.8/7.8/0, 0/0/2.1, 0/0/9.1, 18.6/24.6/14.7, 45/45/52.5, −45/−45/−52.5, 0/0/6.3, 0/0/4.9, −3/−3/−3.5, and 42/42/49 G/cm and durations of 1700, 1216, 503, 1700, 500, 500, 779, 631, 251, 251 μs, respectively. The G1,4,7,8 gradient pulses are sine bell shaped, while the G2,3 gradients are rectangular and G5,6,9,10 are smoothed rectangular. Quadrature detection is achieved using the echo-antiecho approach with the G5 and G6 gradient pulses as well as the ϕ4 pulse inverted for every other FID (Kay et al. 1992). Bruker pulse sequence code included as SI
Sections of the reference (left) and attenuated (right) ARTSY spectra of the U 13C5–1H5 moieties in RiboA, recorded in the presence of 10 mg/mL Pf1. Contour levels for the attenuated spectra are drawn at levels that are 5 times lower than shown for the corresponding reference spectrum. a, b Spectra recorded at 747 MHz with the pulse scheme of Fig. 6, using a dephasing interval of T = 5.1 ms, employing a room temperature 3-axis gradient probehead, with 8 scans per FID, and an interscan delay of 1.5 s, for a total measurement time of ca 9 h for the interleaved 2 × 600* × 512* data matrix (with t1,max = 72 ms, t2,max = 43 ms). c, d Spectra recorded at 600 MHz with the C5(C6)H6-ARTSY pulse scheme of Fig. 9, using a dephasing interval of T = 5.2 ms, employing a cryogenic probehead equipped with a z gradient, using 128 scans per FID, and an interscan delay of 0.7 s, for a total measurement time of ca 32 h for the interleaved 2 × 150* × 512* data matrix (with t1,max = 75 ms, t2,max = 47 ms)
Recovery of H5 magnetization as a function of interscan delay in a the C5–H5 ARTSY experiment of Fig. 6 (747 MHz), recorded in H2O, and b the C5(C6)H6 ARTSY experiment of Fig. 9 (600 MHz). The 1D spectra (left) correspond to the first FID of the ARTSY reference experiment, and have been recorded for interscan delays ranging from 0.5 to 15 s in a and from 0.3 to 7 s in b. Recovery of the overlapping 1H resonances is not fully exponential, indicating a range of recovery rates, which is adequately fitted with a I(t) = Ieq[1 − 1.07exp(−t/1.21s)] (red boxed 1H resonance) or I(t) = Ieq[1 − 1.07exp(−t/1.07s)] (blue boxed resonance), and (B) I(t) = Ieq[1 − 0.95exp(−t/1.05s)]
An alternate ARTSY way to measure C5–H5 RDCs (Fig. 9) encodes the size of the coupling again in the transfer of magnetization from H5 to C5, followed by selection of the downfield 13C–{1H} doublet component by means of an S3E element, and a mixed-time 13C evolution period (Ying et al. 2007a). However, instead of transferring this C5 magnetization to its directly attached H5, the antiphase 13C5–{13C6} magnetization is converted to antiphase 13C6–{13C5} magnetization, followed by a (2JC5C6)−1 rephasing delay and a reverse INEPT transfer of 13C6 magnetization back to 1H6 for detection. Use of H6 band-selective pulses during this last transfer step, combined with pulsed field gradients, permits effective suppression of the H2O signal. The final spectrum (Fig. 7c, d) correlates H6 protons with C5 carbons. The experiment again allows for short interscan delays (Fig. 8b), with water radiation damping taking place during the mixed-time 13C5 evolution period (of minimum duration 2Δ) and additionally the subsequent 13C6 rephasing delay, 2Δ (Fig. 9).
Pulse scheme of the C5(C6)H6 ARTSY experiment for measurement of pyrimidine 1DC5–H5 in H2O by detection of H6. Narrow and wide filled bars represent 90° and 180° pulses, respectively, while wide vertically hatched bars are composite 180° pulses (90x–220y–90x) (Freeman et al. 1980). Composite 13C pulse marked a is only applied for recording the reference spectrum, pulse b only for the attenuated spectrum. Filled half-ellipsoids denote shaped pulses, with the type of nuclei excited, inverted or refocused by each selective pulse (r = ReBURP; e = EBURP2; h = hyperbolic secant; i = IBURP2; i* = time-reversed IBURP2) labeled below the shaped pulse. The adiabatic hyperbolic secant pulses are broad-band and invert C4, C5, and C6 of pyrimidines. All the 1H selective pulses have a duration of 2 ms (at 1H frequency of 600.13MHz) and are centered at 4.1 ppm for the H5 selective ReBURP pulse, at 8.1 ppm for the H6 selective EBURP2 pulse, and at 8.6 ppm for the H6 selective time-reversed IBURP2 and ReBURP pulses. The C5-selective ReBURP and EBURP2 pulses have durations of 1.0 and 1.245 ms, respectively, and are applied at 101.5 ppm. The 600 μs hyperbolic secant pulses is centered at 133 ppm; the 525 μs C4 selective IBURP2 pulses at 189 ppm, and the 525 μs C6/C5 selective ReBURP pulse at 121 ppm, all at 151 MHz 13C (600 MHz 1H) frequency. At 900 MHz 1H frequency, all selective pulses are 33.3% shorter. All other 1H, 13C, and 15N pulses are centered at 4.77, 101.5, and 155 ppm, respectively. All the pulses have phase x unless otherwise indicated. Delays: T = 5.7 ms for the isotropic sample, and 5.2 and 5.1 ms in the two measurements for the weakly aligned sample in Pf1, adjusted to attain attenuated spectra close to the null condition; ε = 150 μs (additional 1 J C5H5 evolution occurs for ~80% of the C5 selective ReBURP pulse); δ = 700 μs (ca 400 μs 1 J C6H6 evolution takes place during the H6-selective time-reversed IBURP2 pulse where the effective inversion point is at ca 20% of its duration); τ = 600 μs; Δ = 4 ms. The C5 chemical shifts are recorded in t1 using the so-called mixed-time (MT) evolution approach (Ying et al. 2007a, b) as follows: ξ1 = max(0, t1/2 − Δ); ξ2 = t1/2; and ξ3 = min(t1/2, Δ). Phase cycling: ϕ1 = y, −y; ϕ2 = 4(45°), 4(225°); ϕ3 = 8(y), 8(−y); ϕ4 = 8(x), 8(−x); ϕ5 = 16(x), 16(y); ϕ6 = x, x, y, y; and ϕrec = x, −x, −x, x, −x, x, x, −x, −x, x, x, −x, x, −x, −x, x, −x, x, x, −x, x, −x, −x, x, x, −x, −x, x, −x, x, x, −x. All gradient pulses are sine bell shaped with mid-point strengths of G1,2,3 = 32.9, 22.4 and 47.6 G/cm, and durations of 477, 497, and 500 μs, respectively. Quadrature detection in t1 by the States-TPPI method, incrementing ϕ3. Pulse sequence code included as SI
The experiment is also readily modified into a 3D experiment by converting the (2JC5C6)−1 rephasing delay into a constant-time or mixed-time 13C6 evolution period. However, this extension was found to be unnecessary for RiboA, in part due to the excellent resolution in the 13C5 dimension afforded by the superior TROSY property of this nucleus. This C5(C6)H6-ARTSY experiment detects H6 protons that resonate relatively far away from the H2O resonance, and water suppression therefore presents little problem and can be accomplished with single-axis z-gradients, as commonly available on cryogenic probeheads. However, the increased signal-to-noise ratio attainable by the use of such a probehead over the use of a room-temperature probehead in our experiments was more than offset by the increased magnetization transfer losses when comparing this experiment to the C5H5-ARTSY experiment (Fig. 6). RDC values for C5H5 measured with the different pulse schemes and different ARTSY dephasing time T are presented in SI Table S2.
Accuracy of 1DNH RDCs derived from ARTSY spectra
Evaluation of the accuracy of RDCs derived from any given measurement method depends on availability of a “gold standard”, and a good fit of observed RDCs to an experimental structure does not guarantee the absence of systematic errors. For example, if all RDCs measured by any given method were underestimated by 10% this would not impact the fit to an atomic model but simply reduce the fitted alignment tensor by 10%. For proteins we have previously shown that ARTSY-derived RDCs are in excellent agreement with 15N frequency-based measurements using the IPAP-HSQC experiment (Fitzkee and Bax 2010). With a root-mean-square difference (rmsd) of 0.49 Hz, we similarly find excellent agreement between ARTSY- and IPAP-HSQC derived 1DNH RDCs (Fig. 10a; RP = 0.997) for RiboA. This pairwise rmsd is only slightly above the average uncertainty of 0.30 Hz, derived from Eq. 3.
Evaluation of the quality of ARTSY-derived 1DNH couplings for RiboA, measured at 900 MHz. a comparison of RDCs derived from the ARTSY spectra of Fig. 2 with those measured from an IPAP-HSQC spectrum (Yao et al. 2009). b Singular value decomposition (SVD) fit (Losonczi et al. 1999) of ARTSY-derived 1DNH couplings (SI Table S3) to the core (nucleotides 18–78) of the RiboA X-ray structure (PDB entry 1Y26), where 1DNH couplings are derived from the difference in 1JNH+1DNH measured in the presence and absence of 10 mg/mL Pf1. c Same as b, but using isotropic couplings calculated from the empirical equation that predicts 1JNH from the 1H chemical shift (i.e., lacking an MSA-induced 1DNH contribution) (Ying et al. 2007a). Uncertainties of the experimentally derived RDCs in all panels are comparable and smaller than the size of the open circles for the majority of residues (see SI Table S3)
As is often seen for proteins, a fit of the 1DNH RDCs to the atomic coordinates is limited not only by the precision at which the RDCs can be measured, but also by the accuracy of the structural model, and by the assumption that all RDCs are equally scaled by internal motion. RiboA contains three A-form helical stems, separated by regions of irregular structure, and with the helices 1 and 2 arranged in a roughly parallel manner, and helix 3 packed against helix 2 in an antiparallel arrangement (Serganov and Patel 2007; Wang et al. 2009). The orientations of the three helices, all roughly parallel to the z axis of the Pf1 molecular alignment tensor, result in a relatively narrow range of the ARTSY-derived 1DNH RDCs for the helical residues (−12 ± 5 Hz). For the RiboA sequence used in our study, four basepair switches in helix 1 were used in order to improve the stability of the aptamer relative to the sequence used for the X-ray structure (see Experimental Section), and the modeled orientation and atomic structure of helix 1 therefore contain additional uncertainty relative to those of helices 2 and 3.
1DNH RDCs fit well to the coordinates of the nucleotides that are conserved relative to the prior X-ray study (nucleotides 18–78; below referred to as “the core” of RiboA). For the 28 observable imino groups of this core (SI Table S3), an excellent fit of the RDCs to the X-ray structure is observed (Fig. 10b; rmsd = 1.65 Hz; Q = 0.107; N = 28). Very similar quality fits are obtained when using 1DNH RDCs obtained from an IPAP-HSQC spectrum (rmsd = 1.55 Hz; Q = 0.100; N = 28, data not shown). 1DNH RDCs for helix 1 also fit fairly well to a model of RiboA based on the novel G2G method (Wang et al. 2009), previously derived on the basis of SAXS data and 1DNH RDCs, which includes the mutated helix 1 (rmsd = 3.7 Hz; Q = 0.22; N = 32).
Interestingly, when using an empirical equation for obtaining the isotropic 1JNH value from the 1HN chemical shift (Ying et al. 2007b), rather than 1JNH splittings measured in the absence of Pf1, good agreement with the core of the RiboA structure is obtained too (Fig. 10c; rmsd = 1.60 Hz; Q = 0.131; N = 28). However, in this case the fitted alignment tensor magnitude is 22% smaller because the isotropic 1JNH values calculated from 1HN chemical shifts do not include the alignment resulting from magnetic susceptibility anisotropy. The 1JNH splittings measured in the absence of Pf1 include small RDCs which are oppositely signed compared to those induced by Pf1 alignment, thereby increasing the difference between \( ^{ 1} {\text{J}}_{{{\text{NH}}^{ - } }} \) splittings measured in Pf1-free and Pf1-containing samples.
Accuracy of 1DCH RDCs derived for C2, C6 and C8
When comparing 1DCH values derived with the ARTSY and IMC methods, excellent agreement is seen for the A–1DC2H2 values, but substantial systematic differences are seen for 1DC6H6 and 1DC8H8 values, where IMC-derived RDCs are smaller by ca 4 and 2 Hz, respectively (Fig. 11a). Fits of the 1DCH RDCs to the RiboA structure require assignment of the corresponding 1H and 13C spectra, a non-trivial task for a molecule the size of RiboA. Assignments used in this study are based on a novel Global Structure Aided Bayesian method (Wang et al. unpublished results), supplemented by 3D NOESY and through-bond J-connectivity data.
Evaluation of the quality of ARTSY-derived 1DC2H2 (blue), 1DC6H6 (red) and 1DC8H8 (green) values measured for RiboA. Outliers are labeled. a Comparison of ARTSY- and IMC-derived 1DCH values. The values from the ARTSY experiment with T = 4.9 ms at 900 MHz (Table S1) are used in the plot. Nucleotide specific assignments are not yet available for a subset of the values shown, including the three 13C–1H pairs with negative RDCs. b Comparison of ARTSY-derived RDCs (average of all the 5 sets given in Table S1) with 1DCH values predicted for the core (nucleotides 18–78) of the X-ray structure (PDB entry 1Y26), using the \( ^{ 1} {\text{D}}_{{{\text{NH}} }} \) derived alignment tensor (rmsd = 3.5 Hz; Q = 0.11; N = 36). c SVD fit of the same 1DCH values as in b to the X-ray coordinates of the core of RiboA (rmsd = 2.8 Hz; Q = 0.086; N = 36). The highly reproducible, negative RDCs seen in a correspond to nucleotides yielding intense, narrow cross peaks, indicative of large amplitude internal dynamics, and are not compatible with any base C–H vector in the X-ray structure
Figure 11b shows the correlation between the experimental ARTSY-derived 1DCH RDCs for the core of RiboA and values predicted for the X-ray structure on the basis of the \( ^{ 1} {\text{D}}_{{{\text{NH}} }} \) derived alignment tensor. Bond lengths used in these analyses (rNH = 1.043 Å for G and 1.057 Å for U; rCH = 1.104 Å) are adjusted such that they include the effect of zero-point librations and vibrations (Ying et al. 2006; Grishaev et al. 2009).
As noted previously, accurate measurement of one-bond splittings in the 1H dimension using the IMC method can be impacted by cross-correlated relaxation between 1H–1H and 1H–13C/15N dipolar interactions, which gives rise to asymmetry in the unresolved 1H–1H multiplet (Tjandra and Bax 1997b; de Alba and Tjandra 2006a, b; Grishaev et al. 2008). Notably, for A–H2 and A/G–H8 protons in the isotropic sample, which lack 1H–1H splittings, ARTSY- and IMC- derived 1JCH splittings agree closely (pairwise rmsd 1.7 Hz). In contrast, in the aligned sample unresolved RDC between H8 and H2′i−1 combined with cross-correlated relaxation by the 13C8–1H8 and 1H8–1H2′i−1 dipolar mechanisms results in a systematic underestimate of the actual splitting (−1.7 ± 3.2 Hz) when measured in the 1H dimension. The magnitude of this effect scales with the size of the 1H–1H RDC which is proportional to the geometric factor, 3cos2θ − 1, where θ is the angle between the 1H–1H vector and the C–H bond, which varies substantially, even within A-form helical regions (SI Table S4).
The IMC-derived RDC values also show considerably worse agreement with values predicted on the basis of the X-ray structure for the core of RiboA (rmsd = 6.7 Hz; Q = 0.21; N = 36) than 1DNH values, and tend to underestimate the actual RDC values. Comparison of observed and predicted ARTSY-derived 1DCH RDCs also show increased scatter compared to what was observed for 1DNH (Fig. 11b; rmsd = 3.5 Hz; Q = 0.11; N = 36), but to a much lesser degree. In part this could be caused by errors in the alignment tensor which was derived from a limited number of 28 1DNH couplings. Indeed, a better fit is obtained when using an SVD fit between experimental 1DCH RDCs of the core of RiboA and coordinates of the X-ray structure (rmsd = 2.8 Hz; Q = 0.086; N = 36; Fig. 11c). Importantly, there is no systematic under- or over-estimate of the ARTSY 1DCH couplings relative to values predicted on the basis of the \( ^{ 1} {\text{D}}_{{{\text{NH}}^{ - } }} \) derived alignment tensor, confirming validity of the effective rNH and rCH bond lengths used in our analysis.
Considering the substantial range of isotropic 1JCH values, from 180 Hz for 1JC6H6 to 215 Hz for 1JC8H8, no single T duration satisfies the requirement to yield a close-to-zero intensity for the resonances of all base 13C–1H groups in the attenuated ARTSY spectrum. However, as can be seen from Eq. 3, the requirement for a close-to-zero intensity is not particularly stringent. Indeed, repeating the CH-ARTSY measurement on the Pf1 containing sample for four different T durations yielded 1JCH+1DCH values that (with few exceptions, e.g. A17 C8H8 and U20 and U25 C6H6) do not vary by more than the S/N-derived uncertainties (SI Table S1).
Accuracy of 1DC5H5 RDCs
Comparison of the RDCs measured at 750 MHz 1H frequency on a room temperature probehead with the scheme of Fig. 6 and with the ARTSY-C5(C6)H6 experiment of Fig. 9 at 600 MHz shows a fair correlation (RP = 0.87; pairwise rmsd = 7.2 Hz; N = 33), but is dominated by the lower signal to noise ratio of the ARTSY-C5(C6)H6 experiment (Fig. 12a).
Evaluation of 1DC5H5 values measured for RiboA by direct (C5H5) and indirect (C5(C6)H6) ARTSY methods. a Plot of directly measured 1DC5H5 values extracted from the 747-MHz ARTSY spectra of Fig. 7a and b with values derived from the indirect 600 MHz C5(C6)H6) ARTSY spectra of Fig. 7c and d. U and C nucleotides are in red and blue, respectively. b Correlation between predicted 1DC5H5 values using SVD fit parameters obtained by fitting all 1DCH and 1DNH of the core residues (SI Tables S1 and S3), versus the directly (C5H5, blue, 750 MHz) or indirectly (red, C5(C6)H6, 600 MHz) derived 1DC5H5 RDCs. In both panels, the averaged values of two indirectly H6-detected C5–H5 RDCs at 600 MHz are used, while for the directly detected RDCs only one set of values on the H2O sample was collected (SI Table S2)
For the values that differ the most between the two methods, we note the following: The discrepancies between couplings measured for U51, U80 and U71 appear to be caused by partial overlap in the C5(C6)H6 spectrum, whereas correlations for C26 and C82 were weak and therefore resulted in intrinsically large uncertainties in the corresponding RDCs. No clear reason for the discrepancy between the 1DC5H5 values measured by C5H5 and C5(C6)H6 methods for U25 and U47 could be identified, but the fact that couplings measured by the C5H5 scheme fit better to the structure suggests that they are more accurate. Except for the two outliers U25 and U71 in Fig. 12b, RDCs measured by C5H5 and C5(C6)H6 methods fit about equally well to the X-ray structure of helical residues in the RiboA core (nucleotides 18–78). Our finding that the simpler C5H5 pulse scheme appears less prone to generate aberrant RDCs makes this the method of choice for 1DC5H5 measurement, but it requires three-axis pulsed field gradient hardware if this measurement is to be carried out on an H2O sample.
Refinement of RiboA model
The natural sequence of stem 1 in RiboA consists of G13GCUUCAUA21 base paired with C83UGAAGUAU75, but is mutated to G13GGAACAUA21/C83CCUUGUAU75 in the modified riboswitch used in the current study. In the X-ray structure, none of the modified base pairs make intramolecular tertiary interactions and a logical way to model stem 1 therefore is to simply model its first 5 basepairs as an ideal A-form helical extension of the top of the A-form helix 1, seen to contact the linker between helices 2 and 3 in the X-ray structure (Serganov and Patel 2007). Generation of an idealized A-form helical extension was carried out in two ways: In a first method, loose harmonic restraints for all phosphodiester backbone torsion and χ angles, together with loose distance restraints were combined with the potential of mean force previously developed for refining nucleic acid structures (Kuszewski et al. 2001). This yielded a model for helix 1 that fits very well to the 32 RDCs (using all available RDCs for helix 1; see Tables S1, S2, and S3; averaged values were used for sites where multiple measurements were made) of this 18-nt helix (rmsd = 1.9 Hz; Q=0.123). In a second method, optimized for generating A-form RNA helices, the PDB was searched for all such helices solved at high resolution (≤2.5 Å resolution), and nucleotide-specific torsion angles and internucleotide distances were extracted, including their standard deviations, and used as input for building an A-form helix (Supporting Information). The A-form helix generated with this second method yielded an SVD fit to the RDC values that was even closer (rmsd = 1.45 Hz; Q=0.098), and is used for all subsequent evaluations.
With few long range contacts between helix 1 and the remainder of RiboA, the question arises to what extent its relative orientation remains the same as that seen in the X-ray structure. Notably, the alignment strength and asymmetry of the best-fitted alignment tensor for helix 1 are the same as observed for the core (18–78) of RiboA, indicating the absence of large scale dynamics of the isolated helix 1 relative to the remainder of the structure. A systematic search over the three angles that define the orientation of helix 1 to helices 2 and 3 indicates that a small improvement in the fit of the RDCs is obtained when helix 1 is rotated by ~6° relative to the position seen in the X-ray structure (Fig. 13). The drop in rmsd between observed (SI Tables S1, S2, and S3) and predicted RDCs (from 1.86 Hz to 1.62 Hz; N=87) upon inclusion of three additional fitting parameters is statistically significant according to F statistics (p=0.01). Considering the minute differences between the original X-ray structure and the refined model (Fig. 13), it is safe to conclude that the structure of the mutated RiboA studied here in solution resembles very closely that seen in the X-ray structure. Loop nucleotides for which assignments are not yet available may pose a possible exception to this similarity, considering that a subset of these exhibit substantial internal mobility in solution, as indicated by longer transverse relaxation times.
Concluding remarks
Our study demonstrates that the ARTSY method previously introduced for measurement of 1DNH couplings in large perdeuterated proteins is readily adapted to measurement of both 1DNH and base 1DCH RDCs in relatively large, protonated oligonucleotides. As for proteins, an important advantage of the ARTSY-based measurement is that couplings are derived from 2D TROSY spectra that offer optimized spectral resolution and good sensitivity. The measurements can readily be carried out on samples in H2O solvent, benefiting from lower viscosity relative to D2O and making it easier to unambiguously compare 1DNH and 1DCH RDCs.
The use of a band-selective 1H pulse at the mid-point of the ARTSY amplitude-encoding interval to a good approximation eliminates the impact of 1H–1H scalar and residual dipolar coupling, while band-selective 13C pulses result in removal of homonuclear 13C–13C couplings in the 13C dimension of the TROSY spectra.
RDC values obtained for helices 2 and 3 of RiboA fit well to the X-ray structure of this oligonucleotide, and the facile measurement of a large set of base RDCs offers valuable constraints in determining a high quality solution structure of this riboswitch once resonance assignments are completed.
In the absence of resonance overlap, the size limit of systems to which the ARTSY method is applicable is governed by the attainable signal-to-noise ratio in the reference spectrum (cf eq 3b). Considering that the shape of RiboA is highly prolate, its rotational diffusion is quite anisotropic. However, under the conditions studied (at 25°C), transverse 15N relaxation rates and 15N–{1H} dipolar/CSA transverse cross-correlation rates (both measured in the presence of Pf1) correspond to rates expected for isotropic rotational diffusion with correlation times of 17.4 ± 3.7 ns, where the large range reflects the different orientations of the imino groups relative to the long axis of the molecule. Upon lowering the temperature to 10°C, transverse 15N and cross-correlated relaxation rates (measured on the same sample) increase by ca 70%, corresponding to an effective correlation time of ca 30 ns. At this lower temperature, the signal to noise ratio in the reference ARTSY spectrum drops by ca 65% for the base 13C correlations (C2, C6 and C8), and by ca 50% for the imino correlations. For base 13C–1H pairs, 30 ns therefore approaches the intrinsic correlation time limit where useful RDCs can be measured in a reasonable amount of time. Accurate RDC measurement for the C5 carbons is somewhat more difficult than for C2, C6 and C8, and it appears that 20–25 ns is the upper limit for correlation times where these RDCs remain measurable by the methods presented here. The imino 15N–1H RDC measurements are less affected by the lower temperature than the aromatic 13C–1H pairs, owing to the favorable TROSY 1H and 15N line widths, and these RDCs can be measured even for systems tumbling somewhat slower than 30 ns.
References
Andersson P, Annila A, Otting G (1998) An alpha/beta-HSQC-alpha/beta experiment for spin-state selective editing of IS cross peaks. J Magn Reson 133:364–367
Bax A, Grishaev A (2005) Weak alignment NMR: a hawk-eyed view of biomolecular structure. Curr Opin Struct Biol 15:563–570
Bax A, Griffey RH, Hawkins BL (1983) Correlation of proton and nitrogen-15 chemical shifts by multiple quantum NMR. J Magn Reson 55:301–315
Bhattacharya A, Revington M, Zuiderweg ERP (2010) Measurement and interpretation of N-15-H-1 residual dipolar couplings in larger proteins. J Magn Reson 203:11–28
Boisbouvier J, Brutscher B, Simorre JP, Marion D (1999) C-13 spin relaxation measurements in RNA: Sensitivity and resolution improvement using spin-state selective correlation experiments. J Biomol NMR 14:241–252
Boisbouvier J, Delaglio F, Bax A (2003) Direct observation of dipolar couplings between distant protons in weakly aligned nucleic acids. Proc Natl Acad Sci USA 100:11333–11338
Bouvignies G, Bernado P, Meier S, Cho K, Grzesiek S, Bruschweiler R, Blackledge M (2005) Identification of slow correlated motions in proteins using residual dipolar and hydrogen-bond scalar couplings. Proc Natl Acad Sci USA 102:13885–13890
Cho CH, Urquidi J, Singh S, Robinson GW (1999) Thermal offset viscosities of liquid H2O, D2O, and T2O. J Phys Chem B 103:1991–1994
Clore GM (2000) Accurate and rapid docking of protein–protein complexes on the basis of intermolecular nuclear Overhauser enhancement data and dipolar couplings by rigid body minimization. Proc Natl Acad Sci USA 97:9021–9025
de Alba E, Tjandra N (2006a) Interference between cross-correlated relaxation and the measurement of scalar and dipolar couplings by quantitative J. J Biomol NMR 35:1–16
de Alba E, Tjandra N (2006b) On the accurate measurement of amide one-bond N-15-H-1 couplings in proteins: Effects of cross-correlated relaxation, selective pulses and dynamic frequency shifts. J Magn Reson 183:160–165
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRpipe—a multidimensional spectral processing system based on Unix pipes. J Biomol NMR 6:277–293
Evenas J, Mittermaier A, Yang DW, Kay LE (2001) Measurement of C-13(alpha)-C-13(beta) dipolar couplings in N- 15, C-13, H-2-labeled proteins: Application to domain orientation in maltose binding protein. J Am Chem Soc 123:2858–2864
Farjon J, Boisbouvier J, Schanda P, Pardi A, Simorre JP, Brutscher B (2009) Longitudinal-Relaxation-Enhanced NMR Experiments for the Study of Nucleic Acids in Solution. J Am Chem Soc 131:8571–8577
Fitzkee NC, Bax A (2010) Facile measurement of H-1-N-15 residual dipolar couplings in larger perdeuterated proteins. J Biomol NMR 48:65–70
Freeman R, Kempsell SP, Levitt MH (1980) Radiofrequency pulse sequences which compensate their own imperfections. J Magn Reson 38:453–479
Geen H, Freeman R (1991) Band-selective radiofrequency pulses. J Magn Reson 93:93–141
Grishaev A, Ying J, Canny MD, Pardi A, Bax A (2008) Solution structure of tRNA(Val) from refinement of homology model against residual dipolar coupling and SAXS data. J Biomol NMR 42:99–109
Grishaev A, Yao LS, Ying JF, Pardi A, Bax A (2009) Chemical Shift Anisotropy of Imino N-15 Nuclei in Watson-Crick Base Pairs from Magic Angle Spinning Liquid Crystal NMR and Nuclear Spin Relaxation. J Am Chem Soc 131:9490–9492
Hansen DF, Vallurupalli P, Kay LE (2008a) Quantifying two-bond (HN)-H-1-(CO)-C-13 and one-bond H-1(alpha)-C-13(alpha) dipolar couplings of invisible protein states by spin-state selective relaxation dispersion NMR spectroscopy. J Am Chem Soc 130:8397–8405
Hansen DF, Vallurupalli P, Kay LE (2008b) Using relaxation dispersion NMR spectroscopy to determine structures of excited, invisible protein states. J Biomol NMR 41:113–120
Hwang TL, van Zijl PCM, Garwood M (1997) Broadband adiabatic refocusing without phase distortion. J Magn Reson 124:250–254
Kay LE, Keifer P, Saarinen T (1992) Pure absorption gradient enhanced heteronuclear single quantum correlation spectroscopy with improved sensitivity. J Am Chem Soc 114:10663–10665
Kontaxis G, Clore GM, Bax A (2000) Evaluation of cross-correlation effects and measurement of one- bond couplings in proteins with short transverse relaxation times. J Magn Reson 143:184–196
Kuszewski J, Schwieters C, Clore GM (2001) Improving the accuracy of NMR structures of DNA by means of a database potential of mean force describing base–base positional interactions. J Am Chem Soc 123:3903–3918
Lescop E, Schanda P, Brutscher B (2007) A set of BEST triple-resonance experiments for time-optimized protein resonance assignment. J Magn Reson 187:163–169
Levitt MH, Freeman R (1981) Compensation for pulse imperfections in NMR spin-echo experiments. J Magn Reson 43:65–80
Losonczi JA, Andrec M, Fischer MWF, Prestegard JH (1999) Order matrix analysis of residual dipolar couplings using singular value decomposition. J Magn Reson 138:334–342
Mantylahti S, Koskela O, Jiang P, Permi P (2010) MQ-HNCO-TROSY for the measurement of scalar and residual dipolar couplings in larger proteins: application to a 557-residue IgFLNa16–21. J Biomol NMR 47:183–194
Meissner A, Duus JO, Sorensen OW (1997) Spin-state-selective excitation. Application for E.COSY-type measurement of J(HH) coupling constants. J Magn Reson 128:92–97
Ottiger M, Delaglio F, Bax A (1998) Measurement of J and dipolar couplings from simplified two-dimensional NMR spectra. J Magn Reson 131:373–378
Palmer AG, Cavanagh J, Wright PE, Rance M (1991) Sensitivity improvement in proton-detected 2-dimensional heteronuclear correlation Nmr-spectroscopy. J Magn Reson 93:151–170
Permi P, Rosevear PR, Annila A (2000) A set of HNCO-based experiments for measurement of residual dipolar couplings in N-15, C-13, (H-2)-labeled proteins. J Biomol NMR 17:43–54
Pervushin K, Riek R, Wider G, Wuthrich K (1997) Attenuated T-2 relaxation by mutual cancellation of dipole- dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc Natl Acad Sci USA 94:12366–12371
Pervushin K, Riek R, Wider G, Wuthrich K (1998a) Transverse relaxation-optimized spectroscopy (TROSY) for NMR studies of aromatic spin systems in C-13-labeled proteins. J Am Chem Soc 120:6394–6400
Pervushin KV, Wider G, Wuthrich K (1998b) Single transition-to-single transition polarization transfer (ST2-PT) in [N15, H1]-TROSY. J Biomol NMR 12:345–348
Peti W, Meiler J, Bruschweiler R, Griesinger C (2002) Model-free analysis of protein backbone motion from residual dipolar couplings. J Am Chem Soc 124:5822–5833
Piotto M, Saudek V, Sklenár V (1992) Gradient-tailored excitation for single-quantum NMR spectroscopy of aqueous sloutions. J Biomol NMR 2:661–665
Prestegard JH, Al-Hashimi HM, Tolman JR (2000) NMR structures of biomolecules using field oriented media and residual dipolar couplings. Q Rev Biophys 33:371–424
Schanda P, Van Melckebeke H, Brutscher B (2006) Speeding up three-dimensional protein NMR experiments to a few minutes. J Am Chem Soc 128:9042–9043
Serganov A, Patel DJ (2007) Ribozymes, riboswitches and beyond: regulation of gene expression without proteins. Nat Rev Genet 8:776–790
Skrynnikov NR, Kay LE (2000) Assessment of molecular structure using frame-independent orientational restraints derived from residual dipolar couplings. J Biomol NMR 18:239–252
Skrynnikov NR, Goto NK, Yang DW, Choy WY, Tolman JR, Mueller GA, Kay LE (2000) Orienting domains in proteins using dipolar couplings measured by liquid-state NMR: Differences in solution and crystal forms of maltodextrin binding protein loaded with beta-cyclodextrin. J Mol Biol 295:1265–1273
Tjandra N, Bax A (1997a) Direct measurement of distances and angles in biomolecules by NMR in a dilute liquid crystalline medium. Science 278:1111–1114
Tjandra N, Bax A (1997b) Measurement of dipolar contributions to (1)J(CH) splittings from magnetic-field dependence of J modulation in two-dimensional NMR spectra. J Magn Reson 124:512–515
Tolbert BS, Miyazaki Y, Barton S, Kinde B, Starck P, Singh R, Bax A, Case DA, Summers MF (2010) Major groove width variations in RNA structures determined by NMR and impact of C-13 residual chemical shift anisotropy and H-1-C-13 residual dipolar coupling on refinement. J Biomol NMR 47:205–219
Tolman JR (2002) A novel approach to the retrieval of structural and dynamic information from residual dipolar couplings using several oriented media in biomolecular NMR spectroscopy. J Am Chem Soc 124:12020–12030
Tolman JR, Ruan K (2006) NMR residual dipolar couplings as probes of biomolecular dynamics. Chem Rev 106:1720–1736
Vallurupalli P, Hansen DF, Kay LE (2008) Structures of invisible, excited protein states by relaxation dispersion NMR spectroscopy. Proc Natl Acad Sci USA 105:11766–11771
van Ingen H, Korzhnev DM, Kay LE (2009) An Analysis of the Effects of H-1(N)-H-1(N) Dipolar Couplings on the Measurement of Amide Bond Vector Orientations in Invisible Protein States by Relaxation Dispersion NMR. J Phys Chem B 113:9968–9977
Vijayan V, Zweckstetter M (2005) Simultaneous measurement of protein one-bond residual dipolar couplings without increased resonance overlap. J Magn Reson 174:245–253
Wang JB, Zuo XB, Yu P, Xu H, Starich MR, Tiede DM, Shapiro BA, Schwieters CD, Wang YX (2009) A Method for Helical RNA Global Structure Determination in Solution Using Small-Angle X-Ray Scattering and NMR Measurements. J Mol Biol 393:717–734
Yang DW, Venters RA, Mueller GA, Choy WY, Kay LE (1999) TROSY-based HNCO pulse sequences for the measurement of (HN)-H- 1-N-15, N-15-(CO)-C-13, (HN)-H-1-(CO)-C-13, (CO)-C-13-C- 13(alpha) and (HN)-H-1-C-13(alpha) dipolar couplings in N-15, C-13, H-2-labeled proteins. J Biomol NMR 14:333–343
Yao L, Vogeli B, Torchia DA, Bax A (2008) Simultaneous NMR study of protein structure and dynamics using conservative mutagenesis. J Phys Chem B 112:6045–6056
Yao LS, Ying JF, Bax A (2009) Improved accuracy of N-15-H-1 scalar and residual dipolar couplings from gradient-enhanced IPAP-HSQC experiments on protonated proteins. J Biomol NMR 43:161–170
Ying JF, Grishaev AE, Bax A (2006) Carbon-13 chemical shift anisotropy in DNA bases from field dependence of solution NMR relaxation rates. Magn Reson Chem 44:302–310
Ying JF, Chill JH, Louis JM, Bax A (2007a) Mixed-time parallel evolution in multiple quantum NMR experiments: sensitivity and resolution enhancement in heteronuclear NMR. J Biomol NMR 37:195–204
Ying JF, Grishaev A, Latham MP, Pardi A, Bax A (2007b) Magnetic field induced residual dipolar couplings of imino groups in nucleic acids from measurements at a single magnetic field. J Biomol NMR 39:91–96
Zhang Q, Sun XY, Watt ED, Al-Hashimi HM (2006) Resolving the motional modes that code for RNA adaptation. Science 311:653–656
Zhang Q, Stelzer AC, Fisher CK, Al-Hashimi HM (2007) Visualizing spatially correlated dynamics that directs RNA conformational transitions. Nature 450:1263–1267
Acknowledgments
This work was supported by the Intramural Research Programs of the NIDDK and NCI, NIH, and by the Intramural AIDS-Targeted Antiviral Program of the Office of the Director, NIH.
Author information
Authors and Affiliations
Corresponding author
Additional information
This manuscript is dedicated to Professor Lewis E. Kay, on the occasion of his 50th birthday.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ying, J., Wang, J., Grishaev, A. et al. Measurement of 1H–15N and 1H–13C residual dipolar couplings in nucleic acids from TROSY intensities. J Biomol NMR 51, 89 (2011). https://doi.org/10.1007/s10858-011-9544-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10858-011-9544-y