Abstract
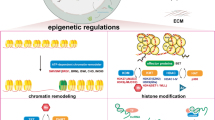
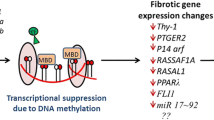
Cardiac fibrosis is defined as excess deposition of extracellular matrix (ECM), resulting in tissue scarring and organ dysfunction. In recent years, despite the underlying mechanisms of cardiac fibrosis are still unknown, numerous studies suggest that epigenetic regulation of cardiac fibrosis. Cardiac fibrosis is regulated by a myriad of factors that converge on the transcription of genes encoding extracellular matrix protein, a process the epigenetic machinery plays a pivotal role. Epigenetic modifications contain three main processes: DNA methylation, histone modifications, and noncoding RNAs. Here, we review recent studies that have illustrated key roles for epigenetic events in the control of pro-fibrotic gene expression, and highlight the potential of molecule mechanisms that target epigenetic regulators as a means of treating cardiac fibrosis.





Similar content being viewed by others
Abbreviations
- miRs:
-
MicroRNAs
- LncRNA:
-
Long noncoding RNA
- ncRNAs:
-
Noncoding RNAs
- α-SMA:
-
α-smooth muscle actin
- ECM:
-
Extracellular matrix
- RNAi:
-
RNA interference
- DNMTs:
-
DNA methyltransferases
- HAT:
-
Histone acetyl transferase
- HDAC:
-
Histone deacetylase
- HCM:
-
Hypertrophic cardiomyopathy
- DBcAMP:
-
cAMP analog N(6),2’-O-dibutyryladenosine 3′,5′-cyclic monophosphate
- MeCP2:
-
Methyl CpG binding protein 2
- CpG:
-
CpG island
- RASAL1:
-
RAS protein activator like-1
- Ras-GTP:
-
RAS GTPase activating protein
- MI:
-
Myocardial infarction
- RAAS:
-
Renin-angiotensin-aldosterone system
- MMP:
-
Maladjustment of matrix metalloproteinases
- EMT:
-
Epithelial mesenchymal transition
- TGF-β:
-
Transforming growth factor beta
- DUSP5:
-
Dual-specificity phosphatase 5
- SHR:
-
Spontaneously hypertensive rat
- EndMT:
-
Endothelial mesenchymal transition
- RASSF1A:
-
Ras association domain family 1 isoform A
- TET:
-
Ten eleven translocation
- TSA:
-
Trichostatin A
- VPA:
-
Valproic acid
- MR:
-
Mineralocorticoid receptor
- MRTF-A:
-
Myocardin-related transcription factor A
- Ang II:
-
Angiotensin II
- EZH1:
-
Enhancer of zeste homolog 1
- TFIID:
-
Transcription factor II D
References
Shimada-Sugimoto M, Otowa T, Miyagawa T, Umekage T, Kawamura Y, Bundo M, Iwamoto K, Tochigi M, Kasai K, Kaiya H, Tanii H, Okazaki Y, Tokunaga K, Sasaki T (2017) Epigenome-wide association study of DNA methylation in panic disorder. Clin Epigenetics 9:6. https://doi.org/10.1186/s13148-016-0307-1
Zam W, Khadour A (2017) Impact of phytochemicals and dietary patterns on epigenome and cancer. Nutr Cancer 69(2):184–200. https://doi.org/10.1080/01635581.2017.1263746
Zhang Y, Ren J (2016) Epigenetics and obesity cardiomyopathy: from pathophysiology to prevention and management. Pharmacol Ther 161:52–66. https://doi.org/10.1016/j.pharmthera.2016.03.005
Rodriguez-Rodero S, Delgado-Alvarez E, Diaz-Naya L, Martin Nieto A, Menendez Torre E (2017) Epigenetic modulators of thyroid cancer. Endocrinol Diabetes Nutr 64(1):44–56. https://doi.org/10.1016/j.endinu.2016.09.006
Satoh A, Niwano S, Niwano H, Kishihara J, Aoyama Y, Oikawa J, Fukaya H, Tamaki H, Ako J (2017) Aliskiren suppresses atrial electrical and structural remodeling in a canine model of atrial fibrillation. Heart Vessel 32(1):90–100. https://doi.org/10.1007/s00380-016-0874-2
Stratton MS, McKinsey TA (2016) Epigenetic regulation of cardiac fibrosis. J Mol Cell Cardiol 92:206–213. https://doi.org/10.1016/j.yjmcc.2016.02.011
Tao H, Yang JJ, Shi KH (2015) Non-coding RNAs as direct and indirect modulators of epigenetic mechanism regulation of cardiac fibrosis. Expert Opin Ther Targets 19(5):707–716. https://doi.org/10.1517/14728222.2014.1001740
Yu LM, Xu Y (2015) Epigenetic regulation in cardiac fibrosis. World J Cardiol 7(11):784–791. https://doi.org/10.4330/wjc.v7.i11.784
Feng B, Cao Y, Chen S, Chu X, Chu Y, Chakrabarti S (2016) miR-200b mediates endothelial-to-mesenchymal transition in diabetic cardiomyopathy. Diabetes 65(3):768–779. https://doi.org/10.2337/db15-1033
Gyongyosi M, Winkler J, Ramos I, Do QT, Firat H, McDonald K, Gonzalez A, Thum T, Diez J, Jaisser F, Pizard A, Zannad F (2017) Myocardial fibrosis: biomedical research from bench to bedside. Eur J Heart Fail 19(2):177–191. https://doi.org/10.1002/ejhf.696
Riaz S, Zeidan A, Mraiche F (2017) Myocardial proteases and cardiac remodeling. J Cell Physiol 232:3244–3250. https://doi.org/10.1002/jcp.25884
Mewhort HE, Lipon BD, Svystonyuk DA, Teng G, Guzzardi DG, Silva C, Yong VW, Fedak PW (2016) Monocytes increase human cardiac myofibroblast-mediated extracellular matrix remodeling through TGF-beta1. Am J Phys Heart Circ Phys 310(6):H716–H724. https://doi.org/10.1152/ajpheart.00309.2015
Tallquist MD, Molkentin JD (2017) Redefining the identity of cardiac fibroblasts. Nat Rev Cardiol 14:484–491. https://doi.org/10.1038/nrcardio.2017.57
Takawale A, Zhang P, Patel VB, Wang X, Oudit G, Kassiri Z (2017) Tissue inhibitor of matrix metalloproteinase-1 promotes myocardial fibrosis by mediating CD63-integrin beta1 interaction. Hypertension 69:1092–1103. https://doi.org/10.1161/HYPERTENSIONAHA.117.09045
Bollong MJ, Yang B, Vergani N, Beyer BA, Chin EN, Zambaldo C, Wang D, Chatterjee AK, Lairson LL, Schultz PG (2017) Small molecule-mediated inhibition of myofibroblast transdifferentiation for the treatment of fibrosis. Proc Natl Acad Sci U S A 114:4679–4684. https://doi.org/10.1073/pnas.1702750114
Prabhu SD, Frangogiannis NG (2016) The biological basis for cardiac repair after myocardial infarction: from inflammation to fibrosis. Circ Res 119(1):91–112. https://doi.org/10.1161/CIRCRESAHA.116.303577
Sag CM, Schnelle M, Zhang J, Murdoch CE, Kossmann S, Protti A, Santos CX, Sawyer GJ, Zhang X, Mongue-Din H, Richards DA, Brewer AC, Prysyazhna O, Maier LS, Wenzel P, Eaton PJ, Shah AM (2017) Distinct regulatory effects of myeloid cell and endothelial cell Nox2 on blood pressure. Circulation 135:2163–2177. https://doi.org/10.1161/CIRCULATIONAHA.116.023877
Seeger T, Xu QF, Muhly-Reinholz M, Fischer A, Kremp EM, Zeiher AM, Dimmeler S (2016) Inhibition of let-7 augments the recruitment of epicardial cells and improves cardiac function after myocardial infarction. J Mol Cell Cardiol 94:145–152. https://doi.org/10.1016/j.yjmcc.2016.04.002
De Cecco CN, Muscogiuri G, Varga-Szemes A, Schoepf UJ (2017) Cutting edge clinical applications in cardiovascular magnetic resonance. World J Radiol 9(1):1–4. https://doi.org/10.4329/wjr.v9.i1.1
Ambrosi C, Manzo M, Baubec T (2017) Dynamics and context-dependent roles of DNA methylation. J Mol Biol 429:1459–1475. https://doi.org/10.1016/j.jmb.2017.02.008
Parrilla-Doblas JT, Ariza RR, Roldan-Arjona T (2017) Targeted DNA demethylation in human cells by fusion of a plant 5-methylcytosine DNA glycosylase to a sequence-specific DNA binding domain. Epigenetics 12(4):296–303. https://doi.org/10.1080/15592294.2017.1294306
Amort T, Lusser A (2017) Detection of 5-methylcytosine in specific poly(A) RNAs by bisulfite sequencing. Methods Mol Biol 1562:107–121. https://doi.org/10.1007/978-1-4939-6807-7_8
Singh VB, Sribenja S, Wilson KE, Attwood KM, Hillman JC, Pathak S, Higgins MJ (2017) Blocked transcription through KvDMR1 results in absence of methylation and gene silencing resembling Beckwith-Wiedemann syndrome. Development 144:1820–1830. https://doi.org/10.1242/dev.145136
Agorio A, Durand S, Fiume E, Brousse C, Gy I, Simon M, Anava S, Rechavi O, Loudet O, Camilleri C, Bouche N (2017) An arabidopsis natural epiallele maintained by a feed-forward silencing loop between histone and DNA. PLoS Genet 13(1):e1006551. https://doi.org/10.1371/journal.pgen.1006551
Blevins T, Wang J, Pflieger D, Pontvianne F, Pikaard CS (2017) Hybrid incompatibility caused by an epiallele. Proc Natl Acad Sci U S A 114(14):3702–3707. https://doi.org/10.1073/pnas.1700368114
Chen C, Wang L, Chen S, Wu X, Gu M, Chen X, Jiang S, Wang Y, Deng Z, Dedon PC, Chen S (2017) Convergence of DNA methylation and phosphorothioation epigenetics in bacterial genomes. Proc Natl Acad Sci U S A 114:4501–4506. https://doi.org/10.1073/pnas.1702450114
Scott H, Smith AE, Barker GR, Uney JB, Warburton EC (2017) Contrasting roles for DNA methyltransferases and histone deacetylases in single-item and associative recognition memory. Neuroepigenetics 9:1–9. https://doi.org/10.1016/j.nepig.2017.02.001
Mendonca A, Sanchez OF, Liu W, Li Z, Yuan C (2017) CpG dinucleotide positioning patterns determine the binding affinity of methyl-binding domain to nucleosomes. Biochim Biophys Acta 1860:713–720. https://doi.org/10.1016/j.bbagrm.2017.03.006
Sassa A, Kanemaru Y, Kamoshita N, Honma M, Yasui M (2016) Mutagenic consequences of cytosine alterations site-specifically embedded in the human genome. Genes Environ 38(1):17. https://doi.org/10.1186/s41021-016-0045-9
Yet I, Tsai PC, Castillo-Fernandez JE, Carnero-Montoro E, Bell JT (2016) Genetic and environmental impacts on DNA methylation levels in twins. Epigenomics 8(1):105–117. https://doi.org/10.2217/epi.15.90
Benesova M, Trejbalova K, Kucerova D, Vernerova Z, Hron T, Szabo A, Amouroux R, Klezl P, Hajkova P, Hejnar J (2017) Overexpression of TET dioxygenases in seminomas associates with low levels of DNA methylation and hydroxymethylation. Mol Carcinog 56:1837–1850. https://doi.org/10.1002/mc.22638
Chen C, Li R, Ross RS, Manso AM (2016) Integrins and integrin-related proteins in cardiac fibrosis. J Mol Cell Cardiol 93:162–174. https://doi.org/10.1016/j.yjmcc.2015.11.010
Grimaldi V, De Pascale MR, Zullo A, Soricelli A, Infante T, Mancini FP, Napoli C (2017) Evidence of epigenetic tags in cardiac fibrosis. J Cardiol 69(2):401–408. https://doi.org/10.1016/j.jjcc.2016.10.004
Jeong HY, Kang WS, Hong MH, Jeong HC, Shin MG, Jeong MH, Kim YS, Ahn Y (2015) 5-Azacytidine modulates interferon regulatory factor 1 in macrophages to exert a cardioprotective effect. Sci Rep 5:15768. https://doi.org/10.1038/srep15768
Salim T, Sershen CL, May EE (2016) Investigating the role of TNF-alpha and IFN-gamma activation on the dynamics of iNOS gene expression in LPS stimulated macrophages. PLoS One 11(6):e0153289. https://doi.org/10.1371/journal.pone.0153289
Lv T, Du Y, Cao N, Zhang S, Gong Y, Bai Y, Wang W, Liu H (2016) Proliferation in cardiac fibroblasts induced by beta1-adrenoceptor autoantibody and the underlying mechanisms. Sci Rep 6:32430. https://doi.org/10.1038/srep32430
Fang X, Robinson J, Wang-Hu J, Jiang L, Freeman DA, Rivkees SA, Wendler CC (2015) cAMP induces hypertrophy and alters DNA methylation in HL-1 cardiomyocytes. Am J Physiol Cell Physiol 309(6):C425–C436. https://doi.org/10.1152/ajpcell.00058.2015
Spitler KM, Ponce JM, Oudit GY, Hall DD, Grueter CE (2017) Cardiac Med1 deletion promotes early lethality, cardiac remodeling, and transcriptional reprogramming. Am J Phys Heart Circ Phys 312(4):H768–H780. https://doi.org/10.1152/ajpheart.00728.2016
Mayer SC, Gilsbach R, Preissl S, Monroy Ordonez EB, Schnick T, Beetz N, Lother A, Rommel C, Ihle H, Bugger H, Ruhle F, Schrepper A, Schwarzer M, Heilmann C, Bonisch U, Gupta SK, Wilpert J, Kretz O, von Elverfeldt D, Orth J, Aktories K, Beyersdorf F, Bode C, Stiller B, Kruger M, Thum T, Doenst T, Stoll M, Hein L (2015) Adrenergic repression of the epigenetic reader MeCP2 facilitates cardiac adaptation in chronic heart failure. Circ Res 117(7):622–633. https://doi.org/10.1161/CIRCRESAHA.115.306721
Tao H, Yang JJ, Hu W, Shi KH, Deng ZY, Li J (2016) MeCP2 regulation of cardiac fibroblast proliferation and fibrosis by down-regulation of DUSP5. Int J Biol Macromol 82:68–75. https://doi.org/10.1016/j.ijbiomac.2015.10.076
Davis JM 3rd, Lin G, Oh JK, Crowson CS, Achenbach SJ, Therneau TM, Matteson EL, Rodeheffer RJ, Gabriel SE (2017) Five-year changes in cardiac structure and function in patients with rheumatoid arthritis compared with the general population. Int J Cardiol 240:379–385. https://doi.org/10.1016/j.ijcard.2017.03.108
Stenzig J, Hirt MN, Loser A, Bartholdt LM, Hensel JT, Werner TR, Riemenschneider M, Indenbirken D, Guenther T, Muller C, Hubner N, Stoll M, Eschenhagen T (2016) DNA methylation in an engineered heart tissue model of cardiac hypertrophy: common signatures and effects of DNA methylation inhibitors. Basic Res Cardiol 111(1):9. https://doi.org/10.1007/s00395-015-0528-z
Watson CJ, Horgan S, Neary R, Glezeva N, Tea I, Corrigan N, McDonald K, Ledwidge M, Baugh J (2016) Epigenetic therapy for the treatment of hypertension-induced cardiac hypertrophy and fibrosis. J Cardiovasc Pharmacol Ther 21(1):127–137. https://doi.org/10.1177/1074248415591698
Singh P, O'Connell M, Shubhashish S (2016) Epigenetic regulation of human DCLK-1 gene during colon-carcinogenesis: clinical and mechanistic implications. Stem cell Invest 3:51. https://doi.org/10.21037/sci.2016.09.07
Xu X, Tan X, Tampe B, Nyamsuren G, Liu X, Maier LS, Sossalla S, Kalluri R, Zeisberg M, Hasenfuss G, Zeisberg EM (2015) Epigenetic balance of aberrant Rasal1 promoter methylation and hydroxymethylation regulates cardiac fibrosis. Cardiovasc Res 105(3):279–291. https://doi.org/10.1093/cvr/cvv015
Tao H, Yang JJ, Chen ZW, Xu SS, Zhou X, Zhan HY, Shi KH (2014) DNMT3A silencing RASSF1A promotes cardiac fibrosis through upregulation of ERK1/2. Toxicology 323:42–50. https://doi.org/10.1016/j.tox.2014.06.006
Takamura M, Kurokawa K, Ootsuji H, Inoue O, Okada H, Nomura A, Kaneko S, Usui S (2017) Long-term administration of eicosapentaenoic acid improves post-myocardial infarction cardiac remodeling in mice by regulating macrophage polarization. J Am Heart Assoc 6(2):e004560. https://doi.org/10.1161/JAHA.116.004560
Watson CJ, Collier P, Tea I, Neary R, Watson JA, Robinson C, Phelan D, Ledwidge MT, McDonald KM, McCann A, Sharaf O, Baugh JA (2014) Hypoxia-induced epigenetic modifications are associated with cardiac tissue fibrosis and the development of a myofibroblast-like phenotype. Hum Mol Genet 23(8):2176–2188. https://doi.org/10.1093/hmg/ddt614
Metes-Kosik N, Luptak I, Dibello PM, Handy DE, Tang SS, Zhi H, Qin F, Jacobsen DW, Loscalzo J, Joseph J (2012) Both selenium deficiency and modest selenium supplementation lead to myocardial fibrosis in mice via effects on redox-methylation balance. Mol Nutr Food Res 56(12):1812–1824. https://doi.org/10.1002/mnfr.201200386
Nagai S, Davis RE, Mattei PJ, Eagen KP, Kornberg RD (2017) Chromatin potentiates transcription. Proc Natl Acad Sci U S A 114(7):1536–1541. https://doi.org/10.1073/pnas.1620312114
Wilson MD, Benlekbir S, Fradet-Turcotte A, Sherker A, Julien JP, McEwan A, Noordermeer SM, Sicheri F, Rubinstein JL, Durocher D (2016) The structural basis of modified nucleosome recognition by 53BP1. Nature 536(7614):100–103. https://doi.org/10.1038/nature18951
Leung A, Trac C, Du J, Natarajan R, Schones DE (2016) Persistent chromatin modifications induced by high fat diet. J Biol Chem 291(20):10446–10455. https://doi.org/10.1074/jbc.M115.711028
Taberlay PC, Achinger-Kawecka J, Lun AT, Buske FA, Sabir K, Gould CM, Zotenko E, Bert SA, Giles KA, Bauer DC, Smyth GK, Stirzaker C, O'Donoghue SI, Clark SJ (2016) Three-dimensional disorganization of the cancer genome occurs coincident with long-range genetic and epigenetic alterations. Genome Res 26(6):719–731. https://doi.org/10.1101/gr.201517.115
Bauer AJ, Martin KA (2017) Coordinating regulation of gene expression in cardiovascular disease: interactions between chromatin modifiers and transcription factors. Front Cardiovasc Med 4:19. https://doi.org/10.3389/fcvm.2017.00019
Filgueiras LR, Brandt SL, Ramalho TR, Jancar S, Serezani CH (2017) Imbalance between HDAC and HAT activities drives aberrant STAT1/MyD88 expression in macrophages from type 1 diabetic mice. J Diabetes Complicat 31(2):334–339. https://doi.org/10.1016/j.jdiacomp.2016.08.001
Ji S, Zhu L, Gao Y, Zhang X, Yan Y, Cen J, Li R, Zeng R, Liao L, Hou C, Gao Y, Gao S, Wei G, Hui L (2017) Baf60b-mediated ATM-p53 activation blocks cell identity conversion by sensing chromatin opening. Cell Res 27:642–656. https://doi.org/10.1038/cr.2017.36
Meyners C, Mertens M, Wessig P, Meyer-Almes FJ (2017) A fluorescence-lifetime-based binding assay for class IIa histone deacetylases. Chemistry 23(13):3107–3116. https://doi.org/10.1002/chem.201605140
Di Giorgio E, Franforte E, Cefalu S, Rossi S, Dei Tos AP, Brenca M, Polano M, Maestro R, Paluvai H, Picco R, Brancolini C (2017) The co-existence of transcriptional activator and transcriptional repressor MEF2 complexes influences tumor aggressiveness. PLoS Genet 13(4):e1006752. https://doi.org/10.1371/journal.pgen.1006752
Mahendrarajah N, Paulus R, Kramer OH (2016) Histone deacetylase inhibitors induce proteolysis of activated CDC42-associated kinase-1 in leukemic cells. J Cancer Res Clin Oncol 142(11):2263–2273. https://doi.org/10.1007/s00432-016-2229-x
Li RF, Cao SS, Fang WJ, Song Y, Luo XT, Wang HY, Wang JG (2017) Roles of HDAC2 and HDAC8 in cardiac remodeling in renovascular hypertensive rats and the effects of valproic acid sodium. Pharmacology 99(1–2):27–39. https://doi.org/10.1159/000449467
Yu L, Yang G, Weng X, Liang P, Li L, Li J, Fan Z, Tian W, Wu X, Xu H, Fang M, Ji Y, Li Y, Chen Q, Xu Y (2015) Histone methyltransferase SET1 mediates angiotensin II-induced endothelin-1 transcription and cardiac hypertrophy in mice. Arterioscler Thromb Vasc Biol 35(5):1207–1217. https://doi.org/10.1161/ATVBAHA.115.305230
Barreiro E, Tajbakhsh S (2017) Epigenetic regulation of muscle development. J Muscle Res Cell Motil 38:31–35. https://doi.org/10.1007/s10974-017-9469-5
Zhang M, Liu H, Gao Y, Zhu Z, Chen Z, Zheng P, Xue L, Li J, Teng M, Niu L (2016) Structural insights into the association of Hif1 with histones H2A-H2B dimer and H3-H4 tetramer. Structure 24(10):1810–1820. https://doi.org/10.1016/j.str.2016.08.001
Wu Z, Connolly J, Biggar KK (2017) Beyond histones: the expanding roles of protein lysine methylation. FEBS J 284:2732–2744. https://doi.org/10.1111/febs.14056
Harr JC, Gonzalez-Sandoval A, Gasser SM (2016) Histones and histone modifications in perinuclear chromatin anchoring: from yeast to man. EMBO Rep 17(2):139–155. https://doi.org/10.15252/embr.201541809
Kim H, Ramirez CN, Su ZY, Kong AN (2016) Epigenetic modifications of triterpenoid ursolic acid in activating Nrf2 and blocking cellular transformation of mouse epidermal cells. J Nutr Biochem 33:54–62. https://doi.org/10.1016/j.jnutbio.2015.09.014
Ahmadi M, Gharibi T, Dolati S, Rostamzadeh D, Aslani S, Baradaran B, Younesi V, Yousefi M (2017) Epigenetic modifications and epigenetic based medication implementations of autoimmune diseases. Biomed Pharmacother 87:596–608. https://doi.org/10.1016/j.biopha.2016.12.072
Luo T, Chen B, Wang X (2015) 4-PBA prevents pressure overload-induced myocardial hypertrophy and interstitial fibrosis by attenuating endoplasmic reticulum stress. Chem Biol Interact 242:99–106. https://doi.org/10.1016/j.cbi.2015.09.025
Pinkerneil M, Hoffmann MJ, Schulz WA, Niegisch G (2017) HDACs and HDAC inhibitors in urothelial carcinoma—perspectives for an antineoplastic treatment. Curr Med Chem (in press)
Schuetze KB, Stratton MS, Blakeslee WW, Wempe MF, Wagner FF, Holson EB, Kuo YM, Andrews AJ, Gilbert TM, Hooker JM, McKinsey TA (2017) Overlapping and divergent actions of structurally distinct histone deacetylase inhibitors in cardiac fibroblasts. J Pharmacol Exp Ther 361(1):140–150. https://doi.org/10.1124/jpet.116.237701
Wang WW, Han JH, Wang L, Bao TH (2017) Scutellarin may alleviate cognitive deficits in a mouse model of hypoxia by promoting proliferation and neuronal differentiation of neural stem cells. Iran J Basic Med Sci 20(3):272–279. https://doi.org/10.22038/IJBMS.2017.8355
Tao H, Yang JJ, Shi KH, Li J (2016) Epigenetic factors MeCP2 and HDAC6 control alpha-tubulin acetylation in cardiac fibroblast proliferation and fibrosis. Inflamm Res 65(5):415–426. https://doi.org/10.1007/s00011-016-0925-2
Chen Y, Du J, Zhao YT, Zhang L, Lv G, Zhuang S, Qin G, Zhao TC (2015) Histone deacetylase (HDAC) inhibition improves myocardial function and prevents cardiac remodeling in diabetic mice. Cardiovasc Diabetol 14:99. https://doi.org/10.1186/s12933-015-0262-8
Kang SH, Seok YM, Song MJ, Lee HA, Kurz T, Kim I (2015) Histone deacetylase inhibition attenuates cardiac hypertrophy and fibrosis through acetylation of mineralocorticoid receptor in spontaneously hypertensive rats. Mol Pharmacol 87(5):782–791. https://doi.org/10.1124/mol.114.096974
Seok YM, Lee HA, Park KM, Hwangbo MH, Kim IK (2016) Lysine deacetylase inhibition attenuates hypertension and is accompanied by acetylation of mineralocorticoid receptor instead of histone acetylation in spontaneously hypertensive rats. Naunyn Schmiedeberg's Arch Pharmacol 389(7):799–808. https://doi.org/10.1007/s00210-016-1246-2
Weng X, Yu L, Liang P, Li L, Dai X, Zhou B, Wu X, Xu H, Fang M, Chen Q, Xu Y (2015) A crosstalk between chromatin remodeling and histone H3K4 methyltransferase complexes in endothelial cells regulates angiotensin II-induced cardiac hypertrophy. J Mol Cell Cardiol 82:48–58. https://doi.org/10.1016/j.yjmcc.2015.02.010
Weng X, Yu L, Liang P, Chen D, Cheng X, Yang Y, Li L, Zhang T, Zhou B, Wu X, Xu H, Fang M, Gao Y, Chen Q, Xu Y (2015) Endothelial MRTF-A mediates angiotensin II induced cardiac hypertrophy. J Mol Cell Cardiol 80:23–33. https://doi.org/10.1016/j.yjmcc.2014.11.009
Leus NG, van den Bosch T, van der Wouden PE, Krist K, Ourailidou ME, Eleftheriadis N, Kistemaker LE, Bos S, Gjaltema RA, Mekonnen SA, Bischoff R, Gosens R, Haisma HJ, Dekker FJ (2017) HDAC1-3 inhibitor MS-275 enhances IL10 expression in RAW264.7 macrophages and reduces cigarette smoke-induced airway inflammation in mice. Sci Rep 7:45047. https://doi.org/10.1038/srep45047
Iyer A, Fenning A, Lim J, Le GT, Reid RC, Halili MA, Fairlie DP, Brown L (2010) Antifibrotic activity of an inhibitor of histone deacetylases in DOCA-salt hypertensive rats. Br J Pharmacol 159(7):1408–1417. https://doi.org/10.1111/j.1476-5381.2010.00637.x
Zhang J, Wang P, Wan L, Xu S, Pang D (2017) The emergence of noncoding RNAs as Heracles in autophagy. Autophagy 13:1004–1024. https://doi.org/10.1080/15548627.2017.1312041
da Rocha ST, Heard E (2017) Novel players in X inactivation: insights into Xist-mediated gene silencing and chromosome conformation. Nat Struct Mol Biol 24(3):197–204. https://doi.org/10.1038/nsmb.3370
Zapata JC, Campilongo F, Barclay RA, DeMarino C, Iglesias-Ussel MD, Kashanchi F, Romerio F (2017) The human immunodeficiency virus 1 ASP RNA promotes viral latency by recruiting the polycomb repressor complex 2 and promoting nucleosome assembly. Virology 506:34–44. https://doi.org/10.1016/j.virol.2017.03.002
Liu X, Liu S (2017) Role of microRNAs in the pathogenesis of diabetic cardiomyopathy. Biomed Rep 6(2):140–145. https://doi.org/10.3892/br.2017.841
Wang A, Kwee LC, Grass E, Neely ML, Gregory SG, Fox KAA, Armstrong PW, White HD, Ohman EM, Roe MT, Shah SH, Chan MY (2017) Whole blood sequencing reveals circulating microRNA associations with high-risk traits in non-ST-segment elevation acute coronary syndrome. Atherosclerosis 261:19–25. https://doi.org/10.1016/j.atherosclerosis.2017.03.041
Pitchiaya S, Heinicke LA, Park JI, Cameron EL, Walter NG (2017) Resolving subcellular miRNA trafficking and turnover at single-molecule resolution. Cell Rep 19(3):630–642. https://doi.org/10.1016/j.celrep.2017.03.075
Heery R, Finn SP, Cuffe S, Gray SG (2017) Long non-coding RNAs: key regulators of epithelial-mesenchymal transition, tumour drug resistance and cancer stem cells. Cancers 9(4). https://doi.org/10.3390/cancers9040038
Han D, Gao Q, Cao F (2017) Long noncoding RNAs (LncRNAs)—the dawning of a new treatment for cardiac hypertrophy and heart failure. Biochim Biophys Acta 1863:2078–2084. https://doi.org/10.1016/j.bbadis.2017.02.024
Liu H, Shang X, Zhu H (2017) LncRNA/DNA binding analysis reveals losses and gains and lineage specificity of genomic imprinting in mammals. Bioinformatics 33(10):1431–1436. https://doi.org/10.1093/bioinformatics/btw818
Hirschi A, Martin WJ, Luka Z, Loukachevitch LV, Reiter NJ (2016) G-quadruplex RNA binding and recognition by the lysine-specific histone demethylase-1 enzyme. RNA 22(8):1250–1260. https://doi.org/10.1261/rna.057265.116
Mazidi M, Penson P, Gluba-Brzozka A, Rysz J, Banach M (2017) Relationship between long noncoding RNAs and physiological risk factors of cardiovascular disease. J Clinical Lipidol 11:617–623. https://doi.org/10.1016/j.jacl.2017.03.009
Qu X, Song X, Yuan W, Shu Y, Wang Y, Zhao X, Gao M, Lu R, Luo S, Zhao W, Zhang Y, Sun L, Lu Y (2016) Expression signature of lncRNAs and their potential roles in cardiac fibrosis of post-infarct mice. Biosci Rep 36(3). https://doi.org/10.1042/BSR20150278
Tao H, Cao W, Yang JJ, Shi KH, Zhou X, Liu LP, Li J (2016) Long noncoding RNA H19 controls DUSP5/ERK1/2 axis in cardiac fibroblast proliferation and fibrosis. Cardiovascular Pathol 25(5):381–389. https://doi.org/10.1016/j.carpath.2016.05.005
Peters T, Hermans-Beijnsberger S, Beqqali A, Bitsch N, Nakagawa S, Prasanth KV, de Windt LJ, van Oort RJ, Heymans S, Schroen B (2016) Long non-coding RNA Malat-1 is dispensable during pressure overload-induced cardiac remodeling and failure in mice. PLoS One 11(2):e0150236. https://doi.org/10.1371/journal.pone.0150236
Jiang X, Zhang F, Ning Q (2015) Losartan reverses the down-expression of long noncoding RNA-NR024118 and Cdkn1c induced by angiotensin II in adult rat cardiac fibroblasts. Pathol Biol 63(3):122–125. https://doi.org/10.1016/j.patbio.2015.04.001
Jiang XY, Ning QL (2014) Expression profiling of long noncoding RNAs and the dynamic changes of lncRNA-NR024118 and Cdkn1c in angiotensin II-treated cardiac fibroblasts. Int J Clin Exp Pathol 7(4):1325–1336
Tao L, Bei Y, Chen P, Lei Z, Fu S, Zhang H, Xu J, Che L, Chen X, Sluijter JP, Das S, Cretoiu D, Xu B, Zhong J, Xiao J, Li X (2016) Crucial role of miR-433 in regulating cardiac fibrosis. Theranostics 6(12):2068–2083. https://doi.org/10.7150/thno.15007
Du W, Liang H, Gao X, Li X, Zhang Y, Pan Z, Li C, Wang Y, Liu Y, Yuan W, Ma N, Chu W, Shan H, Lu Y (2016) MicroRNA-328, a potential anti-fibrotic target in cardiac interstitial fibrosis. Cell Physiol Biochem 39(3):827–836. https://doi.org/10.1159/000447793
Piletic K, Kunej T (2016) MicroRNA epigenetic signatures in human disease. Arch Toxicol 90(10):2405–2419. https://doi.org/10.1007/s00204-016-1815-7
Belleville-Rolland T, Sassi Y, Decouture B, Dreano E, Hulot JS, Gaussem P, Bachelot-Loza C (2016) MRP4 (ABCC4) as a potential pharmacologic target for cardiovascular disease. Pharmacol Res 107:381–389. https://doi.org/10.1016/j.phrs.2016.04.002
Xu Z, Sun J, Tong Q, Lin Q, Qian L, Park Y, Zheng Y (2016) The role of ERK1/2 in the development of diabetic cardiomyopathy. Int J Mol Sci 17(12). https://doi.org/10.3390/ijms17122001
Renaud L, Harris LG, Mani SK, Kasiganesan H, Chou JC, Baicu CF, Van Laer A, Akerman AW, Stroud RE, Jones JA, Zile MR, Menick DR (2015) HDACs regulate miR-133a expression in pressure overload-induced cardiac fibrosis. Circ Heart Fail 8(6):1094–1104. https://doi.org/10.1161/CIRCHEARTFAILURE.114.001781
Zhu WS, Tang CM, Xiao Z, Zhu JN, Lin QX, Fu YH, Hu ZQ, Zhang Z, Yang M, Zheng XL, Wu SL, Shan ZX (2016) Targeting EZH1 and EZH2 contributes to the suppression of fibrosis-associated genes by miR-214-3p in cardiac myofibroblasts. Oncotarget 7(48):78331–78342. https://doi.org/10.18632/oncotarget.13048
Takawale A, Sakamuri SS, Kassiri Z (2015) Extracellular matrix communication and turnover in cardiac physiology and pathology. Compr Physiol 5(2):687–719. https://doi.org/10.1002/cphy.c140045
Jia G, Jia Y, Sowers JR (2016) Role of mineralocorticoid receptor activation in cardiac diastolic dysfunction. Biochim Biophys Acta 1863:2012–2018. https://doi.org/10.1016/j.bbadis.2016.10.025
Acknowledgements
This project was supported by the National Natural Science Foundation of China (81700212, 81570295) and Natural Science Foundation of Anhui Provincial Education Department (KJ2017A168).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Tao, H., Song, ZY., Ding, XS. et al. Epigenetic signatures in cardiac fibrosis, special emphasis on DNA methylation and histone modification. Heart Fail Rev 23, 789–799 (2018). https://doi.org/10.1007/s10741-018-9694-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10741-018-9694-z