Abstract
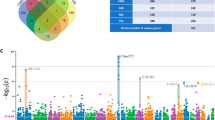
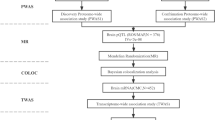
Amyotrophic lateral sclerosis (ALS) is a neurodegenerative disease with strong genetic components. To identity novel risk variants for ALS, utilizing the latest genome-wide association studies (GWAS) and eQTL study data, we conducted a genome-wide expression association analysis by summary data-based Mendelian randomization (SMR) method. Summary data were derived from a large-scale GWAS of ALS, involving 12577 cases and 23475 controls. The eQTL annotation dataset included 923,021 cis-eQTL for 14,329 genes and 4732 trans-eQTL for 2612 genes. Genome-wide single gene expression association analysis was conducted by SMR software. To identify ALS-associated biological pathways, the SMR analysis results were further subjected to gene set enrichment analysis (GSEA). SMR single gene analysis identified one significant and four suggestive genes associated with ALS, including C9ORF72 (P value = 7.08 × 10−6), NT5C3L (P value = 1.33 × 10−5), GGNBP2 (P value = 1.81 × 10−5), ZNHIT3(P value = 2.94 × 10−5), and KIAA1600(P value = 9.97 × 10−5). GSEA identified 7 significant biological pathways, such as PEROXISOME (empirical P value = 0.006), GLYCOLYSIS_GLUCONEOGENESIS (empirical P value = 0.043), and ARACHIDONIC_ACID_ METABOLISM (empirical P value = 0.040). Our study provides novel clues for the genetic mechanism studies of ALS.
Similar content being viewed by others
References
Bonifacino T et al (2016) Altered mechanisms underlying the abnormal glutamate release in amyotrophic lateral sclerosis at a pre-symptomatic stage of the disease. Neurobiol Dis 95:122–133. doi:10.1016/j.nbd.2016.07.011
Calabrese V et al (2005) Oxidative stress, mitochondrial dysfunction and cellular stress response in Friedreich’s ataxia. J Neurol Sci 233:145–162. doi:10.1016/j.jns.2005.03.012
DeJesus-Hernandez M et al (2011) Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72:245–256. doi:10.1016/j.neuron.2011.09.011
Eisen A (2009) Amyotrophic lateral sclerosis: a 40-year personal perspective. J Clin Neurosci 16:505–512. doi:10.1016/j.jocn.2008.07.072
Emerit J, Edeas M, Bricaire F (2004) Neurodegenerative diseases and oxidative stress. Biomed Pharmacother 58:39–46. doi:10.1016/j.biopha.2003.11.004
Ertekin-Taner N (2011) Gene expression endophenotypes: a novel approach for gene discovery in Alzheimer’s disease. Mol Neurodegener 6:31. doi:10.1186/1750-1326-6-31
Farg MA et al (2014) C9ORF72, implicated in amytrophic lateral sclerosis and frontotemporal dementia, regulates endosomal trafficking. Hum Mol Genet 23:3579–3595. doi:10.1093/hmg/ddu068
Federico A, Cardaioli E, Da Pozzo P, Formichi P, Gallus GN, Radi E (2012) Mitochondria, oxidative stress and neurodegeneration. J Neurol Sci 322:254–262. doi:10.1016/j.jns.2012.05.030
Gibson G et al (2012) Brain expression genome-wide association study (eGWAS) identifies human disease-associated variants. PLoS Genet 8:e1002707. doi:10.1371/journal.pgen.1002707
Gijselinck I et al (2012) A C9orf72 promoter repeat expansion in a Flanders-Belgian cohort with disorders of the frontotemporal lobar degeneration-amyotrophic lateral sclerosis spectrum: a gene identification study. Lancet Neurol 11:54–65. doi:10.1016/s1474-4422(11)70261-7
Gordon PH, Mehal JM, Holman RC, Rowland LP, Rowland AS, Cheek JE (2013) Incidence of amyotrophic lateral sclerosis among American Indians and Alaska natives. JAMA Neurol 70:476–480. doi:10.1001/jamaneurol.2013.929
Gu HF, Li HZ, Xie XJ, Tang YL, Tang XQ, Nie YX, Liao DF (2017) Oxidized low-density lipoprotein induced mouse hippocampal HT-22 cell damage via promoting the shift from autophagy to apoptosis. CNS Neurosci Ther. doi:10.1111/cns.12680
Johnson JO et al (2010) Exome sequencing reveals VCP mutations as a cause of familial ALS. Neuron 68:857–864. doi:10.1016/j.neuron.2010.11.036
Kiaei M, Kipiani K, Petri S, Choi DK, Chen J, Calingasan NY, Beal MF (2005) Integrative role of cPLA with COX-2 and the effect of non-steriodal anti-inflammatory drugs in a transgenic mouse model of amyotrophic lateral sclerosis. J Neurochem 93:403–411. doi:10.1111/j.1471-4159.2005.03024.x
Li Y, Willer CJ, Ding J, Scheet P, Abecasis GR (2010) MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet Epidemiol 34:816–834. doi:10.1002/gepi.20533
Majounie E et al (2012) Frequency of the C9orf72 hexanucleotide repeat expansion in patients with amyotrophic lateral sclerosis and frontotemporal dementia: a cross-sectional study. Lancet Neurol 11:323–330. doi:10.1016/S1474-4422(12)70043-1
Marchini J, Howie B, Myers S, McVean G, Donnelly P (2007) A new multipoint method for genome-wide association studies by imputation of genotypes. Nat Genet 39:906–913. doi:10.1038/ng2088
Maruyama H et al (2010) Mutations of optineurin in amyotrophic lateral sclerosis. Nature 465:223–226. doi:10.1038/nature08971
Mizielinska S et al (2014) C9orf72 repeat expansions cause neurodegeneration in Drosophila through arginine-rich proteins. Science 345:1192–1194. doi:10.1126/science.1256800
Murphy A et al (2010) Mapping of numerous disease-associated expression polymorphisms in primary peripheral blood CD4 + lymphocytes. Hum Mol Genet 19:4745–4757. doi:10.1093/hmg/ddq392
Nicolae DL, Gamazon E, Zhang W, Duan S, Dolan ME, Cox NJ (2010) Trait-associated SNPs are more likely to be eQTLs: annotation to enhance discovery from GWAS. PLoS Genet 6:e1000888. doi:10.1371/journal.pgen.1000888
Orr Harry T (2011) FTD and ALS: genetic ties that bind. Neuron 72:189–190. doi:10.1016/j.neuron.2011.10.001
Renton Alan E et al (2011) A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72:257–268. doi:10.1016/j.neuron.2011.09.010
Rosen DR (1993) Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 364:362. doi:10.1038/364362c0
Valbuena GN, Rizzardini M, Cimini S, Siskos AP, Bendotti C, Cantoni L, Keun HC (2016) Metabolomic analysis reveals increased aerobic glycolysis and amino acid deficit in a cellular model of amyotrophic lateral sclerosis. Mol Neurobiol 53:2222–2240. doi:10.1007/s12035-015-9165-7
van Rheenen W et al (2016) Genome-wide association analyses identify new risk variants and the genetic architecture of amyotrophic lateral sclerosis. Nat Genet 48:1043. doi:10.1038/ng.3622
Vance C et al (2009) Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science 323:1208–1211. doi:10.1126/science.1165942
Walford GA (2003) Hypoxia potentiates nitric oxide-mediated apoptosis in endothelial cells via peroxynitrite-induced activation of mitochondria-dependent and -independent pathways. J Biol Chem 279:4425–4432. doi:10.1074/jbc.M310582200
Wang K, Li M, Bucan M (2007) Pathway-based approaches for analysis of genomewide association studies. Am J Hum Genet 81:1278–1283. doi:10.1086/522374
Wang K, Li M, Hakonarson H (2010) Analysing biological pathways in genome-wide association studies. Nat Rev Genet 11:843–854. doi:10.1038/nrg2884
Westra HJ et al (2013) Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat Genet 45:1238–1243. doi:10.1038/ng.2756
Xiao S et al (2016) C9orf72 isoforms in amyotrophic lateral sclerosis and frontotemporal lobar degeneration. Brain Res 1647:43–49. doi:10.1016/j.brainres.2016.04.062
Zarkovic K (2003) 4-Hydroxynonenal and neurodegenerative diseases. Mol Aspects Med 24:293–303. doi:10.1016/s0098-2997(03)00024-4
Zhang F et al (2014) Trans-omics pathway analysis suggests that eQTLs contribute to chondrocyte apoptosis of Kashin-Beck disease through regulating apoptosis pathway expression. Gene 553:166–169. doi:10.1016/j.gene.2014.10.018
Zhu Z et al (2016) Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat Genet 48:481–487. doi:10.1038/ng.3538
Acknowledgements
This study is supported by the National Natural Scientific Foundation of China (81472925, 81673112), and the Technology Research and Development Program of in Shaanxi Province of China (2013KJXX-51), and the Fundamental Research Funds for the Central Universities.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Du, Y., Wen, Y., Guo, X. et al. A Genome-wide Expression Association Analysis Identifies Genes and Pathways Associated with Amyotrophic Lateral Sclerosis. Cell Mol Neurobiol 38, 635–639 (2018). https://doi.org/10.1007/s10571-017-0512-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10571-017-0512-2