Abstract
Objectives
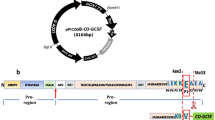
To evaluate different codon optimization parameters on the Saccharomyces cerevisiae-derived mating factor α prepro-leader sequence (MFLS) to improve Candida antarctica lipase B (CAL-B) secretory production in Pichia pastoris.
Results
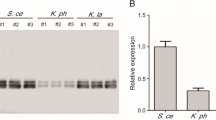
Codon optimization based on the individual codon usage (ICU) and codon context (CC) design parameters enhanced secretory production of CAL-B to 7 U/ml and 12 U/ml, respectively. Only 3 U/ml was obtained with the wild type sequence while the sequence optimized using both ICU and CC objectives showed intermediate performance of 10 U/ml. These results clearly show that CC is the most relevant parameter for the codon optimization of MFLS in P. pastoris, and there is no synergistic effect achieved by considering both ICU and CC together.
Conclusion
The CC optimized MFLS increased secretory protein production of CAL-B in P. pastoris by fourfold.


Similar content being viewed by others
References
Ahn J, Hong J, Lee H, Park M, Lee E, Kim C, Choi E, Jung J, Lee H (2007) Translation elongation factor 1-a gene from Pichia pastoris: Molecular cloning, sequence, and use of its promoter. Appl Microbiol Biotechnol 74:601–608
Anderson EM, Larsson KM, Kirk O (1998) One biocatalyst—many applications: the use of Candida antarctica B-lipase in organic synthesis. Biocatal Biotransform 16:181–204
Chung BK, Lee DY (2012) Computational codon optimization of synthetic gene for protein expression. BMC Syst Biol 6:134
Chung BK, Yusufi FN, Mariati Yang Y, Lee DY (2013) Enhanced expression of codon optimized interferon gamma in CHO cells. J Biotechnol 167:326–333
Dikicioglu D, Wood V, Rutherford KM, McDowall MD, Oliver SG (2014) Improving functional annotation for industrial microbes: a case study with Pichia pastoris. Trends Biotechnol 32:396–399
Fuglsang A (2003) Codon optimizer: a freeware tool for codon optimization. Protein Expr and Purif 31:247–249
Grote A, Hiller K, Scheer M, Munch R, Nortemann B, Hempel DC, Jahn D (2005) JCat: a novel tool to adapt codon usage of a target gene to its potential expression host. Nucleic Acid Res 33:W526–W531
Gustafsson C, Govindarajan S, Minshull J (2004) Codon bias and heterologous protein expression. Trends Biotechnol 22:346–353
Karlin S, Mrázek J, Campbell A, Kaiser D (2001) Characterizations of highly expressed genes of four fast-growing bacteria. J Bacteriol 183:5025–5040
Kim SY, Sohn JH, Pyun YR, Yang IS, Kim KH, Choi ES (2007) In vitro evolution of lipase B from Candida antarctica using surface display in Hansenula polymorpha. J Microbiol Biotechnol 17:1308–1315
Lanza AM, Curran KA, Rey LG, Alper HS (2014) A condition-specific codon optimization approach for improved heterologous gene expression in Saccharomyces cerevisiae. BMC Syst Biol 8:33
Lin-Cereghino GP, Stark CM, Kim D et al (2013) The effect of α-mating factor secretion signal mutations on recombinant protein expression in Pichia pastoris. Gene 519:311–317
Pek HB, Klement M, Ang KS, Chung BK, Ow DS, Lee DY (2015) Exploring codon context bias for synthetic gene design of a thermostable invertase in Escherichia coli. Enz Microb Technol 75–76:57–63
Tuller T, Waldman YY, Kupiec M, Ruppin E (2010) Translation efficiency is determined by both codon bias and folding energy. Proc Natl Acad Sci USA 107:3645–3650
Whang J, Ahn J, Chun CS, Son YJ, Lee H, Choi ES (2009) Efficient, galactose-free production of Candida antarctica lipase B by GAL10 promoter in ∆gal80 mutant of Saccharomyces cerevisiae. Proc Biochem 44:1190–1192
Xiong AS, Yao QH, Peng RH, Han PL, Cheng ZM, Li Y (2005) High level expression of a recombinant acid phytase gene in Pichia pastoris. J Appl Microbiol 98:418–428
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acid Res 31:3406–3415
Acknowledgments
This work was supported by the National University of Singapore, Biomedical Research Council of A*STAR (Agency for Science, Technology and Research), Singapore and Grants from the Global R&D project program (N011500017), Ministry of Trade, Industry and Energy (MOTIE), Republic of Korea, the Next-Generation BioGreen 21 Program (SSAC, No. PJ01109405), Rural Development Administration, Republic of Korea and a grant from KRIBB Research Initiative Program. The authors would like to thank Dr. Chung for his help in the in silico design of sequences.
Supporting information
Additional information: Calculation of ICU and CC fitness. Supplementary Table 1: Primers used in this study. Supplementary Figure 1: 1st repeated comparison of the secretory CAL-B production using different codon optimized MFLS. Supplementary Figure 2: 2nd repeated comparison of the secretory CAL-B production using different codon optimized MFLS with lipase activity measured at different temperatures.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ahn, J., Jang, MJ., Ang, K.S. et al. Codon optimization of Saccharomyces cerevisiae mating factor alpha prepro-leader to improve recombinant protein production in Pichia pastoris . Biotechnol Lett 38, 2137–2143 (2016). https://doi.org/10.1007/s10529-016-2203-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-016-2203-3