Abstract
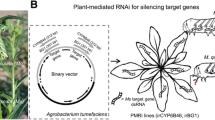
Host-induced gene silencing (HIGS) of insect growth and development is a promising measure for pest control in practice; the feasibility of disarming insect detoxification of toxic phytochemicals via HIGS of insect detoxification enzymes remains largely unexplored. In this study, a HIGS system was applied to interfere with glutathione S-transferase (GST) gene expression in brown planthopper (BPH, Nilaparvata lugens), a key insect pest in rice of Asia, and regulate rice resistance to the insect. The recombination vector in transgenic plants was expressed in almost all organs, especially in vascular bundle tissues, mesophyll, epidermal cells and leaf sheath sap, but not in xylem. Target BPH NlGST1-1 was successfully integrated into rice genome, the resultant transgenic rice lines harbored a single-copy of BPH NlGST1-1 dsRNA, and expression of NlGST1-1 dsRNA was detected in leaves and leaf sheaths of transgenic lines. However, the content of phytochemical gramine, an inherent alkaloid in gramineous plants, did not change in these transgenic plants. BPH nymphs reared on the transgenic rice expressing NlGST1-1 dsRNA displayed significant reductions in NlGST1-1 transcript expression and GST activity. The transgenic plants significantly retarded relative growth rate of the nymphs and the insect female fecundity, the plants also showed enhanced resistance when attacked by BPH. Our study demonstrated the feasibility of reinforcing gramine-mediated plant defense via HIGS of GSTs in herbivorous insects. HIGS-mediated suppression of the herbivore adaptive mechanism has significant implications for efficiently managing BPH and other pests in rice.






Similar content being viewed by others
Availability of data and materials
The datasets used and analyzed during the current study are available from the corresponding authors on reasonable request.
Abbreviations
- ANOVA:
-
The analysis of variance
- ATMT:
-
Agrobacterium tumefaciens-mediated transformation
- BPH:
-
Brown planthopper
- Bt:
-
Bacillus thuringiensis
- DIG:
-
Digoxigenin
- dsRNA:
-
Double-stranded RNA
- EDTA:
-
Ethylenediaminetetraacetic acid
- GFP:
-
Green fluorescent protein
- GST:
-
Glutathione S-transferase
- GE:
-
Genetically engineered
- GUS:
-
β-Glucuronidase
- HIGS:
-
Host-induced gene silencing
- LSD:
-
The least significant difference
- MS medium:
-
Murashige and Skoog medium
- NCBI:
-
National Center for Biotechnology Information
- NR:
-
RNA mixed
- nt:
-
Nucleotide
- NTO:
-
Non-target organism
- P450:
-
Cytochrome P450 monooxygenase
- PCR:
-
Polymerase chain reaction
- qRT-PCR:
-
Quantitative real-time polymerase chain reaction
- RISC:
-
RNA-induced silencing complex
- RNAi:
-
RNA interference
- SDS:
-
Sodium dodecyl sulfate
- SSC:
-
Mixed buffer of sodium chloride and sodium citrate
- TN1:
-
Taichung Native 1 (a rice cultivar)
- WBPH:
-
White-backed planthopper
- WT:
-
Wild-type
References
Abdollahi MR, Memari HR, van Wijnen AJ (2011) Factor affecting the endogenous beta-glucuronidase activity in rapeseed haploid cells: how to avoid interference with the Gus transgene in transformation studies. Gene 487:96–102
Alam SN, Cohen MB (1998) Detection and analysis of QTLs for resistance to the brown planthopper, Nilaparvata lugens, in a doubled-haploid rice population. Theor Appl Genet 97:1370–1379
Alvarez MA (2014) Plant secondary metabolism. In: Alvarez MA (ed) Plant biotechnology for health: from secondary metabolites to molecular farming. Springer, Cham, pp 15–31
Bachman PM, Bolognesi R, Moar WJ, Mueller GM, Paradise MS, Ramaseshadri P, Tan J, Uffman JP, Warren J, Wiggins BE, Levine SL (2013) Characterization of the spectrum of insecticidal activity of a double-stranded RNA with targeted activity against Western Corn Rootworm (Diabrotica virgifera virgifera LeConte). Trans Res 22:1207–1222
Baum JA, Bogaert T, Clinton W, Heck GR, Feldmann P, Ilagan O, Johnson S, Plaetinck G, Munyikwa T, Pleau M, Vaughn T, Roberts J (2007) Control of coleopteran insect pests through RNA interference. Nat Biotechnol 25:1322–1326
Bautista MAM, Miyata T, Miura K, Tanaka T (2009) RNA interference-mediated knockdown of a cytochrome P450, CYP6BG1, from the diamondback moth, Plutella xylostella, reduces larval resistance to permethrin. Insect Biochem Mol Biol 39:38–46
Board PG, Menon D (2013) Glutathione transferases, regulators of cellular metabolism and physiology. Biochem Biophys Acta 1830:3267–3288
Cai QN, Zhang QW, Cheo M (2004) Contribution of indole alkaloids to Sitobion avenae (F.) resistance in wheat. J Appl Entomol 128:517–521
Cai QN, Han Y, Cao YZ, Hu Y, Zhao X, Bi JL (2009) Detoxification of gramine by the cereal aphid Sitobion avenae. J Chem Ecol 35:320–325
Christiaens O, Dzhambazova T, Kostov K, Arpaia S, Joga MR, Urru I, Sweet J, Smagghe G (2018) Literature review of baseline information on RNAi to support the environmental risk assessment of RNAi-based GM plants. EFSA Support Publ 15:1424E
Damásio J, Guilhermino L, Soares AM, Riva MC, Barata C (2007) Biochemical mechanisms of resistance in Daphnia magna exposed to the insecticide fenitrothion. Chemosphere 70:74–82
Elsayed GM (2011) Plant secondary substances and insects behaviour. Arch Phytopathol Plant Protect 44:1534–1549
Enayati AA, Ranson H, Hemingway J (2005) Insect glutathione transferases and insecticide resistance. Insect Mol Biol 14:3–8
Epa US (2014) RNAi technology as a pesticide: problem formulation for human health and ecological risk assessment. US EPA, Washington
Fairbairn DJ, Cavallaro AS, Bernard M, Mahalinga-Iyer J, Graham MW, Botella JR (2007) Host-delivered RNAi: an effective strategy to silence genes in plant parasitic nematodes. Planta 226:1525–1533
Ghaffar MBA, Pritchard J, Ford-Lloyd B (2011) Brown planthopper (Nilaparvata lugens Stal) feeding behaviour on rice germplasm as an indicator of resistance. PLoS One 6:e22137
Gloss AD, Vassão DG, Hailey AL, Nelson DAC, Schramm K, Reichelt M, Rast TJ, Weichsel A, Cravens MG, Gershenzon J, Montfort WR, Whiteman NK (2014) Evolution in an ancient detoxification pathway is coupled with a transition to herbivory in the Drosophilidae. Mol Biol Evol 31:2441–2456
Head GP, Carroll MW, Evans SP, Rule DM, Willse AR, Clark TL, Storer NP, Flannagan RD, Samuel LW, Meinke LJ (2017) Evaluation of SmartStax and SmartStaxPRO maize against western corn rootworm and northern corn rootworm: efficacy and resistance management. Pest Manag Sci 73:1883–1899
Heidel-Fischer HM, Vogel H (2015) Molecular mechanisms of insect adaptation to plant secondary compounds. Curr Opin Insect Sci 8:8–14
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Jena KK, Jeung JU, Lee JH, Choi HC, Brar DS (2006) High-resolution mapping of a new brown planthopper (BPH) resistance gene, Bph18(t), and marker-assisted selection for BPH resistance in rice (Oryza sativa L.). Theor Appl Genet 112:288–297
Knapp E, Flores R, Scheiblin D, Modla S, Czymmek K, Yusibov V (2012) A cryohistological protocol for preparation of large plant tissue sections for screening intracellular fluorescent protein expression. Biotechniques 52:31–37
Li X, Schuler MA, Berenbaum MR (2007) Molecular mechanisms of metabolic resistance to synthetic and natural xenobiotics. Ann Rev Entomol 52:231–253
Liu Z, Han Z (2006) Fitness costs of laboratory-selected imidacloprid resistance in the brown planthopper, Nilaparvata lugens Stål. Pest Manag Sci 62:279–282
Liu Y, Wu H, Chen H, Liu Y, He J, Kang H, Sun Z, Pan G, Wang Q, Hu J, Zhou K, Zheng X, Ren Y, Chen L, Wang Y, Zhao Z, Lin Q, Wu F, Zhang X, Guo X, Cheng X, Jiang L, Wu C, Wang H, Wan J (2014) A gene cluster encoding lectin receptor kinases confers broad-spectrum and durable insect resistance in rice. Nat Biotechnol 33:301–305
Lumjuan N, Stevenson BJ, Prapanthadara LA, Somboon P, Brophy PM, Loftus BJ, Severson DW, Ranson H (2007) The Aedes aegypti glutathione transferase family. Insect Biochem Mol Biol 37:1026–1035
Luo J, Liang SJ, Li JY, Xu ZP, Li L, Zhu BQ, Li Z, Lei CL, Lindsey K, Chen LZ, Jin SX, Zhang XL (2017) A transgenic strategy for controlling plant bugs (Adelphocoris suturalis) through expression of double-stranded RNA homologous to fatty acyl-coenzyme A reductase in cotton. New Phytol 215:1173–1185
Mao J, Zeng F (2014) Plant-mediated RNAi of a gap gene-enhanced tobacco tolerance against the Myzus persicae. Trans Res 23:145–152
Mao YB, Cai WJ, Wang JW, Hong GJ, Tao XY, Wang LJ, Huang YP, Chen XY (2007) Silencing a cotton bollworm P450 monooxygenase gene by plant-mediated RNAi impairs larval tolerance of gossypol. Nat Biotechnol 25:1307–1313
Noda H, Ishikawa K, Hibino H, Omura T (1991) A reovirus in the brown planthopper, Nilaparvata lugens. J Gen Virol 72(Pt 10):2425–2430
Ortelli F, Rossiter LC, Vontas J, Ranson H, Hemingway J (2003) Heterologous expression of four glutathione transferase genes genetically linked to a major insecticide-resistance locus from the malaria vector Anopheles gambiae. Biochem J 373:957–963
Pan H, Xu L, Noland JE, Li H, Siegfried BD, Zhou X (2016) Assessment of potential risks of dietary RNAi to a soil micro-arthropod, Sinella curviseta Brook (Collembola: Entomobryidae). Front Plant Sci 7:1028
Pan H, Yang X, Romeis J, Siegfried BD, Zhou X (2020) Dietary RNAi toxicity assay exhibits differential responses to ingested dsRNAs among lady beetles. Pest Manag Sci 76:3606–3614
Pitino M, Coleman AD, Maffei ME, Ridout CJ, Hogenhout SA (2011) Silencing of aphid genes by dsRNA feeding from plants. PLoS One 6:e25709
Roberts AF, Devos Y, Lemgo GNY, Zhou X (2015) Biosafety research for non-target organism risk assessment of RNAi-based GE plants. Front Plant Sci 6:958
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Saha P, Majumder P, Dutta I, Ray T, Roy SC, Das S (2006) Transgenic rice expressing Allium sativum leaf lectin with enhanced resistance against sap-sucking insect pests. Planta 223:1329–1343
Singh SP, Coronella JA, Benes H, Cochrane BJ, Zimniak P (2001) Catalytic function of Drosophila melanogaster glutathione S-transferase DmGSTS1-1 (GST-2) in conjugation of lipid peroxidation end products. Eur J Biochem 268:2912–2923
Singh A, Jaswal A, Sarkar S (2019) Incidence of brown planthopper in the rice field with the use of different doses of fertilizers. Agric Sci Digest 39:108–113
Stevenson PC, Kimmins FM, Grayer RJ, Raveendranath S (1996) Schaftosides from rice phloem as feeding inhibitors and resistance factors to brown planthoppers, Nilaparvata lugens. In: Proceedings of the 9th international symposium on insect–plant relationships, vol 53, pp 246–249
Sun XQ, Zhang MX, Yu JY, Jin Y, Ling B, Du JP, Li GH, Qin QM, Cai QN (2013) Glutathione S-transferase of brown planthoppers (Nilaparvata lugens) is essential for their adaptation to gramine-containing host plants. PLoS One 8:e64026
Sun Y, Sparks C, Jones H, Riley M, Francis F, Du W, Xia L (2019) Silencing an essential gene involved in infestation and digestion in grain aphid through plant-mediated RNA interference generates aphid-resistant wheat plants. Plant Biotechnol J 17:852–854
Tabashnik BE, Gassmann AJ, Crowder DW, Carriére Y (2008) Insect resistance to Bt crops: evidence versus theory. Nat Biotechnol 26:199–202
US EPA (2017) Human health and ecological risk assessments for SmartStax PRO (MON 89034 x TC1507 x MON 87411 x DAS-59122-7), a plant-incorporated protectant intended to control corn rootworm through ribonucleic acid (RNA) interference
Vontas JG, Small GJ, Hemingway J (2001) Glutathione S-transferases as antioxidant defence agents confer pyrethroid resistance in Nilaparvata lugens. Biochem J 357:65–72
Vontas JG, Small GJ, Nikou DC, Ranson H, Hemingway J (2002) Purification, molecular cloning and heterologous expression of a glutathione S-transferase involved in insecticide resistance from the rice brown planthopper, Nilaparvata lugens. Biochem J 362:329–337
Walshe DP, Lehane SM, Lehane MJ, Haines LR (2009) Prolonged gene knockdown in the tsetse fly Glossina by feeding double stranded RNA. Insect Mol Biol 18:11–19
Wang Y, Cai QN, Zhang QW, Han Y (2006) Effect of the secondary substances from wheat on the growth and digestive physiology of cotton bollworm Helicoverpa armigera (Lepidoptera: Noctuidae). Eur J Entomol 103:255–258
Whyard S, Singh AD, Wong S (2009) Ingested double-stranded RNAs can act as species-specific insecticides. Insect Biochem Mol Biol 39:824–832
Wi SG, Choi IS, Kim KH, Kim HM, Bae HJ (2013) Bioethanol production from rice straw by popping pretreatment. Biotechnol Biofuels 6:166
Xiong Y, Zeng H, Zhang Y, Xu D, Qiu D (2013) Silencing the HaHR3 gene by transgenic plant-mediated RNAi to disrupt Helicoverpa armigera development. Int J Biol Sci 9:370–381
Yang J, Sun XQ, Yan SY, Pan WJ, Zhang MX, Cai QN (2017) Interaction of ferulic acid with glutathione S-transferase and carboxylesterase genes in the brown planthopper, Nilaparvata lugens. J Chem Ecol 43:693–702
Zha W, Peng X, Chen R, Du B, Zhu L, He G (2011) Knockdown of midgut genes by dsRNA-transgenic plant-mediated RNA interference in the hemipteran insect Nilaparvata lugens. PLoS One 6:e20504
Zhang HW, Qu ZC, Xu XY, Zhang XN, Bai FW, Wan YZ,Shao MH, Ye MM, Shen DL (2002) Selection and expression of peptides which can change the conformation of P20 protein of rice stripe virus. Acta Virol 46:11–17
Zhang M, Fang T, Pu G, Sun X, Zhou X, Cai Q (2013) Xenobiotic metabolism of plant secondary compounds in the English grain aphid, Sitobion avenae (F.) (Hemiptera: Aphididae). Pest Biochem Physiol 107:44–49
Zhou Y, Han Z, Zhang X, Pang C (2003) Influence of rice varieties on the metabolizing enzymes of Nilaparvata lugens Stål. J Nanjing Agric Univ 26:24–28
Zhou WW, Liang QM, Xu Y, Gurr GM, Bao YY, Zhou XP, Zhang CX, Cheng J, Zhu ZR (2013) Genomic insights into the glutathione S-transferase gene family of two rice planthoppers, Nilaparvata lugens (Stal) and Sogatella furcifera (Horvath) (Hemiptera: Delphacidae). PLoS One 8:e56604
Zhu-Salzman K, Bi JL, Liu TX (2005) Molecular strategies of plant defense and insect counter-defense. Insect Sci 12:3–15
Zúñiga GE, Corcuera LJ (1985) Role of an indole alkaloid in the resistance of barley seedlings to aphids. Phytochemistry 24:945–947
Acknowledgements
We are grateful to Professors Chengui Han and Zejian Guo (China Agricultural University) for providing pMCG161 RNAi vector and pCAMBIA1301 transformation vector, respectively. We also thank Professor Hongxia Hua (Huazhong Agricultural University) for providing the brown planthopper (biotype II) and anonymous Reviewers for excellent suggestions.
Funding
This work was supported by the “the National Key Research and Development Program of China” (2016YFD0300203) and the Chinese Universities Scientific Fund (2015NX001).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Communicated by G. Smagghe.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10340_2020_1296_MOESM1_ESM.pdf
Figure S1. Construction of rice RNAi vector. Rice RNAi vector pCAMBIA1301-NlGST1-1 was constructed on the basis of the pCAMBIA1301 plasmid vector and pMCG161 RNAi vector. The two-step cloning procedure for generation of the constructs is illustrated as the followings. The region between the two restriction enzyme cutting sites BamH I and Hind III derived from the T-DNA region of pMCG161 RNAi vector contains the CaMV 35S promoter (CaMV35S) upstream of rice waxy-A1 introns and an octopine synthase 3′ end (OSC3′). Two multiple cloning sites are located on both sides of the rice waxy-A intron (Intron), providing sites for cloning target gene in an inverted orientation (open arrows). The fragment between the left (LB) and right (RB) boundaries of the pCAMBIA1301 plasmid vector contains CaMV 35S polyA (PolyA), hygromycin resistance gene (hpt), CaMV 35S promoter (CaMV35S), multiple cloning sites, and a GUS marker gene (GUS). The pCAMBIA1301 vector was linearized with restriction enzymes BamH I and Hind III and ligated to the region between the two restriction sites BamH I and Hind III of pMCG161 RNAi vector to generate the recombinant vector. The NlGST1-1 sequences (full-length or fragment) were inserted two multiple cloning sites flanked the rice waxy-A intron in the recombinant vector, and finally, the construction of the plant RNAi vector was completed. The restriction enzyme cutting sites used for vector construction are shown in this map. Figure S2. Transgenic rice plants suppress BPH NlGST1-1 expression, GST activity, relative growth rate and female fecundity. When BPH fed on gst651 or gst426 lines, the reduced percentages of BPH NlGST1-1 expression level (a), GST activity (b), relative growth rate (c) and female fecundity (d) were statistically analyzed. Data represent the mean ± SD from four replicates via Paired Samples t Test. * and ** indicate significance at P < 0.05 and 0.01, respectively (PDF 353 kb)
10340_2020_1296_MOESM2_ESM.xlsx
Table S1. Used primers for detection of target gene in RNAi vector and identification of transgenic rice plants. Table S2. BPH NlGST1-1 21-nucleotide (nt). Table S3. The 21-nt matches with other species (XLSX 34 kb)
Rights and permissions
About this article
Cite this article
Yang, J., Sun, XQ., Zhu-Salzman, K. et al. Host-induced gene silencing of brown planthopper glutathione S-transferase gene enhances rice resistance to sap-sucking insect pests. J Pest Sci 94, 769–781 (2021). https://doi.org/10.1007/s10340-020-01296-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10340-020-01296-6