Abstract
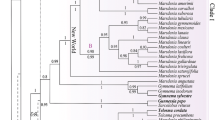
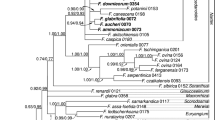
The phylogeny of subgenus Cyathophora and representatives of its closely related taxa within Allium were reconstructed based on nrDNA ITS and two plastid fragments (trnL-F and rpl32-trnL). The constructed phylogenies indicated that subgenus Cyathophora was not monophyletic and to be split in three parts positioned in different clusters. Allium kingdonii was unequivocally placed within subgenus Amerallium and formed an immediate sister relationship with New World Amerallium clade, suggesting an unexpected intercontinental disjunct distribution. For another, Allium trifurcatum was firmly nested within subgenus Butomissa next to A. tuberosum and A. ramosum, but it is distinctly different morphologically from the latter by thinly leathery bulb tunics, uniovulate locule and obviously 3-cleft stigma. Based on the geographic features, morphological and molecular evidences, two new sections, Kingdonia X.J.He et D.Q.Huang for A. kingdonii and Trifurcatum X.J.He et D.Q.Huang for A. trifurcatum, were proposed. The remaining three species of subgenus Cyathophora formed a well-defined clade, and the phylogenetic relationships among them recovered were consistent with previous findings. In addition, A. weschniakowii and A. subtilissimum were proven to be a member of subgenera Rhizirideum sensu stricto (s. str.) and Cepa, respectively, rather than subgenera Cepa and Polyprason previously proposed. Section Rhizomatosa represented by A. caespitosum should be subsumed within section Caespitosoprason of subgenus Rhizirideum s. str.


Similar content being viewed by others
References
Barker FK, Lutzoni FM (2002) The utility of the incongruence length difference test. Syst Biol 51:625–637
Berry PE, Hahn WJ, Sytsma KJ, Hall JC, Mast A (2004) Phylogenetic relationships and biogeography of Fuchsia (Onagraceae) based on noncoding nuclear and chloroplast DNA data. Am J Bot 91:601–614
Burland TG (2000) DNASTAR’s Lasergene sequence analysis software. Methods Mol Biol 132:71–91
Choi HJ, Giussani LM, Jang CG, Oh BU, Cota-Sánchez JH (2012) Systematics of disjunct northeastern Asian and northern North American Allium (Amaryllidaceae). Botany 90:491–508
Diels L (1912) Plantae Chinenses Forrestianae: new and imperfectly known species. Notes Roy Bot Gard Edinburgh 25:302
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Dubouzet JG, Shinoda K (1999) Relationships among old and new world Alliums according to ITS DNA sequence analysis. Theor Appl Genet 98:422–433
Ekberg L (1969) Studies in the genus Allium. II. A new subgenus and new sections from Asia. Bot Notiser 122:57–68
Friesen N, Fritsch RM, Pollner S, Blattner FR (2000) Molecular and morphological evidence for an origin of the aberrant genus Milula within Himalayan species of Allium (Alliaceae). Mol Phylogenet Evol 17:209–218
Friesen N, Fritsch RM, Blattner FR (2006) Phylogeny and new intrageneric classification of Allium (Alliaceae) based on nuclear ribosomal DNA ITS sequences. Aliso 22:372–395
Fritsch RM (1988) Anatomische Untersuchungen an der Blattspreite bei Allium L. (Alliaceae) I. Arten mit einer einfachen Leitbündelreihe. Flora Morphol Geobot Oekophysiol 181:83–100
Fritsch RM (2001) Taxonomy of the genus Allium: contribution from IPK Gatersleben. Herbertia 56:19–50
Fritsch RM, Friesen N (2002) Evolution, domestication and taxonomy. In: Rabinowitch HD, Currah L (eds) Allium crop science: recent advances. CABI Publishing, Wallingford, pp 5–30
Fritsch RM, Blattner FR, Gurushidze M (2010) New classification of Allium L. subg. Melanocrommyum (Webb & Berthel) Rouy (Alliaceae) based on molecular and morphological characters. Phyton 49:145–220
Gurushidze M (2009) Phylogenetic relationships and diversification processes in Allium subgenus Melanocrommyum. Ph.D thesis, Martin-Luther-Universität Halle-Wittenberg, Halle
Hanelt P (1992) Ovule number and seed weight in the genus Allium L. In: Hanelt P, Hammer K, Knúpffer H (eds) The genus Allium: taxonomic problems and genetic resources. Gatersleben, Germany, pp 99–105
Hanelt P, Schulze-Motel J, Fritsch RM, Kruse J, Maab HI, Ohle H, Pistrick K (1992) Infrageneric grouping of Allium-the Gatersleben approach. In: Hanelt P, Hammer K, Knúpffer H (eds) The genus Allium: taxonomic problems and genetic resources. Gatersleben, Germany, pp 107–123
Huang RF, Xu JM, Yu H (1995) A study on karyotypes and their evolutionary trends in Allium sect. Bromatorrhiza Ekberg (Liliaceae). Cathaya 7:133–145
Klaas M, Friesen N (2002) Molecular markers in Allium. In: Rabinowitch HD, Currah L (eds) Allium crop science: resent advances. CABI Publishing, Wallingford, pp 159–185
Li QQ, Zhou SD, He XJ, Yu Y, Zhang YC, Wei XQ (2010) Phylogeny and biogeography of Allium (Amaryllidaceae: Allieae) based on nuclear ribosomal internal transcribed spacer and chloroplast rps16 sequences, focusing on the inclusion of species endemic to China. Ann Bot 106:709–733
Meerow AW, Fay MF, Guy CL, Li QB, Zaman FQ, Chase MW (1999) Systematics of Amaryllidaceae based on cladistic analysis of plastid sequence data. Am J Bot 86:1325–1345
Nguyen NH, Driscoll HE, Specht CD (2008) A molecular phylogeny of the wild onions (Allium; Alliaceae) with a focus on the western North American center of diversity. Mol Phylogenet Evol 47:1157–1172
Ni NC (2002) Molecular phylogenetic studies on sect. Bromatorrhiza (Allium) from China. Master thesis, Sichuan University, Chengdu
Nylander JAA (2004) MrModeltest 2.2. Computer program and documentation distributed by the author. Evolutionary Biology Centre, Uppsala University, Uppsala
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Shaw J, Lickey EB, Schilling EE, Small RL (2007) Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III. Am J Bot 94:275–288
Souza LGR, Crosa O, Speranza P, Guerra M (2012) Cytogenetic and molecular evidence suggest multiple origins and geographical parthenogenesis in Nothoscordum gracile (Alliaceae). Ann Bot 109:987–999
Stearn WT (1960) Allium and Milula in the Central and Eastern Nepal Himalaya. Bull Br Mus (Nat Hist) Bot 2:159–191
Swofford DL (2003) PAUP*. Phylogenetic analysis using parsimony (*and other methods), version 4. Sinauer Associates, Sunderland
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17:1105–1109
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Wheeler EJ, Mashayekhi S, McNeal DW, Columbus JT, Pires JC (2013) Molecular systematics of Allium subgenus Amerallium (Amaryllidaceae) in North America. Am J Bot 100:701–711
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic, San Diego, pp 315–322
Xie L, Wagner WL, Ree RH, Berry PE, Wen J (2009) Molecular phylogeny divergence time estimates and historical biogeography of Circaea (Onagraceae) in the Northern Hemisphere. Mol Phylogenet Evol 53:995–1009
Xu JM (1980) Allium L. In: Wang FZ, Tang J (eds) Flora Reipublicae Popularis Sinicae, vol 14. Science Press, Beijing, pp 202–213
Xu JM (1991) Allium L. In: Xu JM (ed) Flora Sichuanica, vol 7. Sichuan Ethnic Publishing House, Chengdu, pp 132–176
Xu JM, Kamelin RV (2000) Allium L. In: Wu ZY, Raven PH (eds) Flora of China, vol 24. Science Press/Missouri Botanical Garden Press, Beijing/St. Louis, pp 165–202
Xue PF (1994) Study on karyotypes and their relationships in Allium sect. Bromatorrhiza from China. Master thesis, Sichuan University, Chengdu
Yang L (1995) Systematic relationships of Allium sect. Bromatorrhiza from China. Master thesis, Sichuan University, Chengdu
Acknowledgments
We are very grateful to Prof. Jie-Mei Xu for Allium species identification and to Xiang-Guang MA and Yun-Dong GAO for assistance in gathering plant material and valuable discussions on our manuscript. This work was supported by the National Natural Science Foundation of China (Grant No. 31270241, 31070166, 31100161), and the specimen platform of China, teaching specimen’s sub-platform (http://mnh.scu.edu.cn/).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Huang, DQ., Yang, JT., Zhou, CJ. et al. Phylogenetic reappraisal of Allium subgenus Cyathophora (Amaryllidaceae) and related taxa, with a proposal of two new sections. J Plant Res 127, 275–286 (2014). https://doi.org/10.1007/s10265-013-0617-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-013-0617-8