Abstract
Transcriptomic and metabolomic profiles were used to unravel drought adaptation mechanisms in wild emmer wheat (Triticum turgidum ssp. dicoccoides), the progenitor of cultivated wheat, by comparing the response to drought stress in roots of genotypes contrasting in drought tolerance. The differences between the drought resistant (R) and drought susceptible (S) genotypes were characterized mainly by shifts in expression of hormone-related genes (e.g., gibberellins, abscisic acid (ABA) and auxin), including biosynthesis, signalling and response; RNA binding; calcium (calmodulin, caleosin and annexin) and phosphatidylinositol signalling, in the R genotype. ABA content in the roots of the R genotype was higher in the well-watered treatment and increased in response to drought, while in the S genotype ABA was invariant. The metabolomic profiling revealed in the R genotype a higher accumulation of tricarboxylic acid cycle intermediates and drought-related metabolites, including glucose, trehalose, proline and glycine. The integration of transcriptomics and metabolomics results indicated that adaptation to drought included efficient regulation and signalling pathways leading to effective bio-energetic processes, carbon metabolism and cell homeostasis. In conclusion, mechanisms of drought tolerance were identified in roots of wild emmer wheat, supporting our previous studies on the potential of this genepool as a valuable source for novel candidate genes to improve drought tolerance in cultivated wheat.
Similar content being viewed by others
Introduction
The ever-increasing human population, concomitantly with loss of agricultural land (due to urbanization processes), diminishing water availability and climate change, pose serious challenges to world agriculture (Mittler and Blumwald 2010). Crop plants are often grown under unfavourable environmental conditions such as water deficit, heat stress and/or stress combination that prevent the full expression of their genetic yield potential (Cattivelli et al. 2008). Plant development and productivity are determined by coordinated activities of roots and shoots; while the rate of photosynthesis in shoots is affected by the size and activity of the root system, root growth and its maintenance depend on assimilates supplied by the shoot (Manschadi et al. 2008). Abiotic stress greatly affects the structure and development of the root systems (Malamy 2005; Nibau et al. 2008).
Plant adaptations to environmental cues such as temperature fluctuations, water and nutrients imbalance and response(s) to pathogens are mediated by phytohormones that can act either at the site of synthesis or following their transport, elsewhere in the plant (reviewed by Peleg and Blumwald 2011). Genomics and physiological data indicate that plant hormones act within a complex network with extensive cross talk (Nemhauser et al. 2006). One of the faster responses of plants to soil water stress is the accumulation of abscisic acid (ABA) in the roots (Thompson et al. 2007), which is transported through the xylem to the shoot (Wilkinson and Davies 2010) causing stomatal closure, reducing water loss via transpiration (Schroeder et al. 2001) and eventually restricting cellular growth. Recent evidence show that other plant hormones, such as gibberellin (GA) and auxin (indole-3-acetic acid, IAA), may also play a role in plant adaptation to stress. GA promotes plant growth through cell elongation via the disappearance of nuclear DELLA proteins (Murase et al. 2008). IAA has an important role in the fine tuning of hormone signalling in the roots (Iglesias et al. 2010) and in hydrotropism, which is the response of roots to a moisture gradient in the soil (Miyazawa et al. 2009). Most information on molecular processes activated in roots comes from studies of the model plant Arabidopsis thaliana, which is characterized by simple anatomy. The major differences in roots anatomy of dicots vs. monocots are also mirrored by gene expression patterns (Jiao et al. 2009). Furthermore, the current available information on the role of phytohormones in roots adaptation to abiotic stress in cereal crops is limited (Brady et al. 2007; Levin et al. 2009; Benfey et al. 2010 and references therein).
Wheat (Triticum spp.), with ~620 million tons produced annually worldwide (FAOstat 2009, http://faostat.fao.org/site/339/default.aspx), provides about one fifth of the calories consumed by humans. Wheat production is seriously affected in many growing areas of the world by reduction of water availability. The genetic diversity of crop species, including wheat, has been severely eroded throughout the processes of domestication and modern breeding. Wild relatives of crop plants offer a promising genetic resource for crop improvement (Tanksley and McCouch 1997). Wild emmer wheat [Triticum turgidum ssp. dicoccoides (Körn.) Thell] is the allo-tetraploid (2n = 4x = 28; genome BBAA) progenitor of both domesticated tetraploid durum wheat [T. turgidum ssp. durum (Desf.) MacKey] and hexaploid (2n = 6x = 42; BBAADD) bread wheat (Triticum aestivum L.) (Feldman 2001). Wild emmer genepool harbour wide allelic repertoire for various agronomically important traits (Nevo et al. 2002 and references therein) including for drought tolerance (Peleg et al. 2005, 2007, 2009). Recently, we have used transcriptome analysis for identification of drought adaptation mechanisms in wild emmer (Krugman et al. 2010).
In the current study, we have investigated drought adaptation mechanisms in the roots of wild emmer wheat using transcriptomic and metabolomic profiling. The combination of gene expression and metabolites profiling revealed differential expression of genes encoding enzymes associated with GA and ABA biosynthesis and signalling and with response to IAA, in addition to hormone-regulated metabolic changes. These results shed new light on drought adaptation mechanism in wheat roots that can serve as a basis for improvement of drought tolerance in cultivated wheat and other cereal crops.
Material and methods
Plant materials
In previous studies, 110 accessions of wild emmer collected across a water availability gradient were characterized for their responses to drought conditions in the field (Peleg et al. 2005, 2007). Based on these multiple-year studies, two genotypes were selected for transcriptome and metabolome analyses of roots: (1) a drought resistant (R) genotype (Y12-3) originated from Yehudiyya (35°42′ N; 32°56′ E) and (2) a drought susceptible (S) genotype (A24-39) originated from Amirim (35°27′ N; 32°55′ E). The R and S genotypes exhibited contrasting productivity under water-limited conditions (e.g. spike and total dry matter) and yield stability across environments (Peleg et al. 2005, 2007; Avneri 2009), as well as considerable differences in leaf transcriptome and its response to drought (Krugman et al. 2010).
Experimental conditions
Seeds were germinated on moist germination paper (Hofman Manufacturing, Inc, Albany). Seedlings were transplanted into 0.5-L pots containing a mixture of pure quartz sand (80%) and peat (20%), supplemented with a slow-release fertilizer (2 g/L, Osmocote® Standard 14–14–14, Scotts-Sierra Horticulture, Marysville, OH, USA) and a weekly application of 100 mL of 0.5× Murashige and Skoog (MS) growth solution (Sigma Chemical Co., St Louis, MO, USA). Plants were grown in a controlled environment greenhouse (23/18°C; 12:12- h day/night cycle). Drought stress was applied 4 weeks after germination when plants were at a five to six leaf stage. Drought stress was applied by withholding water for 7 days until stress symptoms (i.e. leaf rolling and wilting) were visible in leaves of both genotypes. Five biological replicates were grown for each genotype/treatment combination; root tissues of three of them were used for transcriptome analysis and quantitative PCR while five replicates were used for metabolomic analysis. Roots were gently cleaned from sand remains, excised, immediately frozen in liquid nitrogen and stored at −80°C for RNA extraction and for metabolic profiling.
RNA extraction, Affymetrix GeneChip® hybridization and data analyses
RNA extraction, cRNA preparation and Affymetrix GeneChip® hybridization were applied as described in Krugman et al. (2010). Altogether, 12 RNA samples were used for transcriptome analyses, by hybridization to 12 Affymetrix GeneChip® Wheat Genome Arrays (Affymetrix, Santa Clara, CA, USA; i.e. 2 genotypes × 2 environments × 3 biological replicates). The CELL data files of the 12 Affymetrix GeneChip® results were deposited in the public microarray database NCBI GEO (https://www.ncbi.nlm.nih.gov/projects/geo), accession GSE25940.
Statistical analysis
Transformation, normalisation and quality control procedures were performed using JMP Genomics® (ver. 3.2) statistical package (SAS, Cary, NC, USA) as described by Krugman et al. (2010). Briefly, analysis of variance (ANOVA) was employed to test the effects of two factors: genotype (G; A24-39 vs. Y12-3) and environment (E; drought stress vs. well-watered control) and their interactions (G × E) on log2 of expression values. In the ANOVA model, E, G and ‘probe’ were considered as fixed effects and ‘Array’ as random effect. A threshold of significance was based on the Bonferroni correction of P = 0.05 (Hochberg 1988).
Gene filtering
In order to identify transcripts having large differences between the two genotypes in response to drought stress, the full list of differentially expressed transcripts (DETs) obtained by ANOVA, which included thousands of transcripts, was narrowed down by further gene filtering. The filtered DETs in the R or S genotypes (further designated as DETR or DETS) were selected based on the following statistical requirements: significant effect of E (i.e. water regimes) for the resistant or susceptible genotype, effect of G under the drought conditions and G × E interaction effect, a FC > 1.4 (log2 FC > 0.4) was selected as the threshold in the comparisons between R and S genotypes under drought and between treatments in the R or S genotypes.
Bioinformatics
Transcripts were annotated using HarvEST Affymetrix Wheat1 Chip (http://harvest.ucr.edu/; ver. 1.55). Plant MetGenMAP visualization and analysis package (http://bioinfo.bti.cornell.edu/cgi-bin/MetGenMAP/home.cgi) was used to identify ‘significantly changed pathways’ and ‘enriched GO terms’ (described by Joung et al. 2009). The Plant MetGenMAP system does not support wheat; therefore, the Affymetrix probe-set numbers were first translated into rice loci by HarvEST Affymetrix Wheat1 Chip and were then up-loaded to MetGenMAP for further analysis (Krugman et al. 2010).
Quantitative PCR analysis
The RNA samples used for microarray analysis were also served as templates for qPCR (as described in Krugman et al. 2010), with three technical replicates for each biological sample. Ubiquitin was selected as an endogenous gene to normalise well to well within and across plates based on the study of Paolacci et al. (2009) and additional indications of minimal change in expression level between genotypes and treatments in the current microarray analysis. Primer sequences of the selected transcripts used for validation are described in Table S1. Quantification of gene expression was carried out by the relative quantification method (Skern et al. 2005).
Metabolic profiling
Root tissues of five biological replicates of each genotype/treatment were analysed by GC-MS using an established protocol (Lisec et al. 2006). Relative metabolite content was calculated as described in Roessner et al. (2001) following peak identification using TagFinder (Luedemann et al. 2008). Substances were identified by comparison to mass spectral tags library (Schauer et al. 2005; Erban et al. 2006). The data were analysed using Student t test embedded in the R software environment (http://www.R-project.org). The data was subjected to principal component analysis (PCA), performed with the software package TMEV (Saeed et al. 2004, 2006), and graphs were created using Sigma Plot (Sigma Plot 2000, Systat Software Inc., San Jose, CA, USA).
Abscisic acid determination
Determination of ABA was conducted by UPLC-QTOF-MS/MS (Waters Ltd.) in a similar manner to the method described by López-Carbonell and Jáuregui (2005) with modification as pertaining to our tissue. Tissue (100 mg) was extracted using an extraction solution of methanol and water (80:20). During sample running, the mobile phase consisted of 95% water, 5% acetonitrile, 0.1% formic acid (phase A) and 0.1% formic acid in acetonitrile (phase B). The solvent gradient programme was conditioned as follows: 100–60% solvent A over the first 8 min, 60–0% solvent A over 1 min and return to the initial 100% A in 3.5 min and conditioning at 100% A. Masses of the eluted compounds was detected by the QTOF equipped with electrospray ionization in negative ion mode. ABA was identified and quantified by comparing with authentic standard calibration curves.
Results
Alteration in gene regulation in response to drought
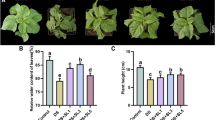
The prolonged drought stress of 7 days resulted in alteration of growth and development in both genotypes, including roots elongation and growth inhibition of the aerial part of the plants. Stress symptoms that were observed in both genotypes after withholding water were more visible in the leaves of the S genotype; however, no significant difference was observed in root length between the two genotypes. The whole transcriptome comparison of drought response in roots of the R and S genotypes revealed 2,962 DETs at least in one genotype, R or S. A total of 745 DETs were up- and 558 were downregulated (altogether 1,303 DETs) in the R genotype, while only 314 DETs were differentially expressed in the S genotype, of which 138 were up- and 176 were downregulated. In addition, 1,345 DETs showed similar expression patterns in response to drought stress in both genotypes, of which 648 were up- and 697 were downregulated. The trend of differential gene regulation in response to drought in the two genotypes is illustrated by a Venn diagram (Fig. 1a). A higher number of DETs in the R genotype was observed previously in the leaves of the same genotype under terminal drought (Krugman et al. 2010). A similar trend of gene expression was obtained also in drought resistant and susceptible genotypes of potato clones (Legay et al. 2011). As described in the M&M, further gene filtering narrowed down the list of DETs to 185 DETRs and DETSs showing the largest difference of gene regulation between the genotypes. Of these 185 transcripts, 125 were highly or uniquely altered in the R genotype (DETRs hereafter), while only 60 transcripts had altered expression in the S genotype (DETSs hereafter). Notably, the majority of these transcripts were upregulated (117 DETRs and 48 DETSs). Of the 185 transcripts, 31 showed a significant drought effect in both genotypes, of which 16 showed higher expression in the R genotype and 15 showed higher expression in the S genotype; thus, they were regarded as DETRs and DETSs, respectively. The results of gene filtering in the two genotypes is illustrated by a Venn diagram (Fig. 1b).
Venn diagrams summarizing regulation patterns of DETs and further filtering of DETRs and DETSs in wild emmer wheat. a Differentially and commonly expressed (up- and downregulated) transcripts in the resistance (R) and sensitive (S) genotypes; b DETs filtering procedure and the resulted DETSs and DETRs, each showing significant G, E and G × E effects (P < 0.05) and FC > 1.4 in the comparisons between: (a) R and S genotypes under water-stress (in the dashed red circles) and (b) water-stress vs. well-watered control in each of the genotypes
Gene enrichment analysis and functional classification of DETs
Gene enrichment analysis
The very large and complex genome of wheat is not fully sequenced yet, and it is poorly annotated (Choulet et al. 2010). Therefore, gene annotation and further enrichment analysis were based on sequence comparisons with the fully sequenced rice genome (Oryza sativa L.). As a result, the list of DETs that was used for GO term enrichment analyses was reduced twice, first by using only transcripts that had rice annotation and then by excluding redundant rice accession numbers. Therefore, the list of DETs was reduced by about 45% to 1,388 genes in the R genotype (837 upregulated and 551 downregulated) and by 40% to 1,007 genes (514 upregulated and 493 downregulated) in the S genotype. The enrichment analyses of GO terms based on biological processes identified 30 enriched GOs among the upregulated DETs (Table 1); 19 of them were enriched in both genotypes, such as ‘embryonic development’ which includes seed maturation and late embryogenesis abundant (LEA) proteins that are involved in response to drought stress and drought resistance (Choi et al. 1999; Suprunova et al. 2004). Notably, these genes were not only highly represented, but most of them were characterized by high fold change in response to drought. For example, 26 transcripts annotated as LEA genes and 17 transcripts annotated as dehydrin genes were upregulated, most of them in both genotypes. The expression pattern of such LEA gene (Ta.14145.S1._at) was confirmed in the two genotypes by qPCR (Fig. S1). Interestingly, most of the GOs of group C (Table 1) that are associated with multilevel regulation were enriched only in the R genotype. The enrichment analyses based on molecular function revealed that ‘nucleotide binding’ and ‘RNA binding’ were exclusively enriched in the R genotype. Additionally, ‘translation’ GO group that was enriched in both genotypes included a 56% higher number of genes in the R genotype.
The annotation analyses were further focused on the filtered DETRs (Fig. 2), of which only 84 could be annotated and classified into functional groups; these were compared with the 30 annotated DETSs (Tables S2 and S3, respectively). The DETRs were classified into three main functional categories (Fig. 2): (a) multilevel regulation (39%), included transcriptional and post-transcriptional regulation, translation activity, signalling and plant hormone metabolism and signalling. The relatively high proportion of multilevel regulation and signalling genes, in particular genes involved in plant hormones, is much higher than those identified previously in leaves (26%) of the same genotype under terminal drought (Krugman et al. 2010); (b) metabolism (29%), included protein, lipid and carbohydrate metabolism, metabolic and catalytic activities and (c) binding, transport and energy proteins (26%). The remaining genes (6%) were classified as stress and membrane proteins. Of the 30 annotated DETSs, 25% were classified as associated with multilevel regulation genes (25%) while 20% were classified as stress-induced proteins (20%). We further review the genes known to be involved in multilevel regulation, in particular in plant hormone homeostasis.
Gene products of 84 DETRs classified into functional categories. The main categories are marked in colours: (1) multilevel regulation in red, including transcription and post-transcriptional regulation, signalling, kinase activity and hormone signalling; (2) metabolic and catalytic activity in blue, including proteins, carbohydrates and lipids; (3) transport and energy transfer in green; (4) stress-induced proteins in black; (5) other in brown
Changes in the expression of genes encoding hormone-associated pathways
A comparison between the DETRs and DETSs revealed differential expression of genes encoding enzymes mediating hormone biosynthesis and homeostasis in addition to downstream hormone-mediated processes. In general, ABA and GA were differentially expressed between the R and S genotype, while genes involved in IAA were differentially expressed in response to drought but not between genotypes. Based on these findings, we further searched our full data of the non-filtered DETs for additional genes that may be associated with these three plant hormones. The identified hormone-related transcripts are described in Table 2, and their pattern of expression in the two genotypes is illustrated by the heat map Fig. 3.
Expression pattern of hormone-associated genes in the resistance (R) and sensitive (S) genotypes in response to water-stress. a abscisic acid (ABA), b gibbereline (GA) and c auxin (IAA). Well-watered control (C), water-stress (D). Red and green represent high and low relative expression when compared to the mean value of expression across all samples, respectively. Scale is log2 of mean expression value. Annotation of transcripts and full names of the putative genes are detailed in Table 2
ABA
Genes associated with ABA biosynthesis, signalling or response were up- or downregulated in the R genotype under drought stress (Fig. 3a; Table 2). The biosynthesis of ABA is associated with increased rates of carotenoid and epoxycarotenoid synthesis, in addition to increased epoxy carotenoid cleavage by 9-cis-epoxycarotenoid dioxygenase (NCED) that are needed to elevate ABA biosynthesis. In the current study, while NCED was not identified as differentially expressed, other essential genes in the ABA biosynthesis pathway were highly upregulated in the R genotype, including zeaxanthin epoxidase (ZEP) which is known to be activated upstream of NECD and aldehyde oxidase AAO1 and AAO3, which catalyse the final step in the biosynthesis of ABA through abscisic aldehyde into ABA. Interestingly, NECD was not differentially expressed between these two genotypes also in the leaves (Krugman et al. 2010). The expression patterns of AAO1 as well as the ABA-responsive gene PM19 were validated by qPCR (Fig. S1). In addition to ABA metabolism, important regulators were identified in both genotypes, including PP2C genes (Ta.12348.1.A1_at, Ta.12348.1.S1_at and Ta.26201.1.S1_at) that are structurally related to ABI1 and HAB2 known as negative regulators of the ABA response, OST1/SNRK2E (Ta.6040.1.S1_a_at; open stomata 1 SNF1-related protein kinase 2.6) and the nuclear cap-binding protein CBP80/ABH1 known as important regulation of ABA modulation via post-transcriptional mRNA processing (Table 2). Furthermore, several ABA-responsive genes were highly upregulated in both genotypes, such as dehydration-responsive genes and several members of the dehydrin gene family (e.g. RD20, RAB18, WZY, dhn1, dhn2 and dhn2). Further metabolic analysis confirmed that ABA content in the R genotype was significantly higher in the well-watered treatment and further increased in response to drought, while in the S genotype ABA was invariant in comparison with the control treatment (Fig. 4). Interestingly, the dehydration-responsive gene RD22 (Ta.26910.1.S1_at), which was commonly used in many studies as an indicator for ABA response (Yamaguchi-Shinozaki and Shinozaki 1993; Abe et al. 2003), was highly upregulated in the roots of the R genotype under the well-watered conditions and downregulated under the drought treatment. RD22 was previously highly expressed under drought in the leaves of the R genotype; therefore, we confirmed its pattern of regulation by qPCR, using the same PCR primers that were used in the leaves. The reduced expression of RD22 that was indentified in the roots of the R genotype under drought may be related to IAA and ABA cross talk that is modulated by the transcription factor MYB96 in the roots (Seo et al. 2009). A second gene that was expected to be upregulated under drought is PM19 (Ta.24312.1.S1_at), which is also well-known as an ABA-responsive gene (Ranford et al. 2002). Surprisingly, PM19 was downregulated in the current study only in roots of the R genotype but was highly upregulated in the S genotype under drought (Table 2). It has been shown by Mashiguchi et al. (2008) that GA can downregulate several genes related to ABA function including PM19; therefore, the reduction of the ABA-responsive gene PM19 in the roots of the R genotype may be explained by ABA and GA cross talk.
GA
Genes involved in GA biosynthesis, signalling and response were differentially expressed in the R and S genotyped under stress. GA 2-beta-dioxygenase (GA2OX1) was downregulated in both genotypes (Table 2); as was shown by Yamaguchi et al. (2007), downregulation of GA2OX1 may cause an increase in the active GA. Other GA-responsive and regulation genes listed in Table 2 showed higher upregulation in the R genotype. These include: GID1L2 which encodes the receptor that binds GA, and O-linked N-acetyl glucosamine (O-GlcNAc) transferase that may be involved in the activity or stability of DELLA proteins by modification or phosphorylation via the action of SPINDLY (SPY) as a negative regulator of GA signalling. The dissection of GA biosynthesis, signalling and further downstream transcriptional events is highly complex also due to gene-specific expression, cell types and developmental stages within the roots, as was described in Arabidopsis (Birnbaum et al. 2003). This complexity is indicated in the current study by the alteration of several genes, including the chitin-inducible GA-responsive genes (CIGR1), which is a member of the GRAS gene family in rice that was upregulated while another CIGR gene showing similarity to SCARECROW (SCR; Ta.4968.1.S1_at) was downregulated. The SCR gene is a key regulator of the asymmetric cell division, which is one of the most important mechanisms in the diversification of cell function and fate. Additionally, the transcription factor GAMYB (Ta.19664.1.A1_at) was downregulated in both genotypes. GAMYB is responsible for the tissue specific expression of GA-regulated genes in cereal aleurone cells (Gubler et al. 1999).
IAA
Genes involved in IAA transport and binding had higher or lower expression in response to drought in both genotypes (Table 3; Fig. 3c). Nonetheless, several IAA efflux carriers had a slightly higher expression in the R genotype. The IAA efflux carrier (PIN3; Ta.5171.1.S1_at) was highly downregulated in both genotypes (Fig. 3c; Table 2). PIN3 has been hypothesized to function in the redirection of auxin toward the new lower flank of gravi-stimulated roots. In gravi-stimulated roots, PIN3 re-localizes to the lower side of statocytes, a process that is thought to play a role in the asymmetrical redistribution of auxin (Harrison and Masson 2008).
Transcriptional and post-transcriptional regulation
Transcriptional regulation
Large number of DETs annotated as transcription factors (TFs) belonging to different gene families were identified in the roots in response to drought. The most abundant TFs were members of MYB and bZIP gene families. Of the MYB gene family, six members were down- and four were upregulated in response to drought; the Myb1 (Ta.26049.1.S1_a_at), a transcription activator involved in the regulation of flavonoid biosynthesis, was one of the most highly upregulated TF (FC log2 4.1 and 3.87 in S and R genotypes, respectively) in both R and S genotypes. Myb1 was previously reported to be induced in roots of wheat in response to hypoxia and other abiotic stresses (Lee et al. 2007). Of this gene family MCB2; (DETR TaAffx.5070.1.S1_at) was the only differentially expressed gene in the R genotype. Members of the bZIP family have an important role in ABA response and regulation of oxidative and pathogen defence responses (Lu et al. 2009). Two of the bZIP transcripts (Ta.30801.1.S1_s_at and TaAffx.132154.1.S1_at) were up- or downregulated in both R and S genotypes, respectively. These transcripts were annotated as AREB3 (ABA-responsive element binding protein 3) in Arabidopsis. MADS-box genes are key components of the networks that control the transition to flowering and flower development, but their role in vegetative development is poorly understood. One member of this gene family, MADS-box26 (DETR Ta.26215.2.S1_s_at), was upregulated in the R genotype (FC > 2). It shows similarity to MIKCc type-box26 in rice and to AGL12 (AGAMOUS-LIKE 12/XAL1), which is required for normal root development and proper transition to flowering in Arabidopsis, and was suggested to mediate auxin participation in root meristematic cells and in root cell elongation (Tapia-Lopez et al. 2008).
Post-transcriptional regulation
Genes encoding helicases and RNA binding proteins were significantly enriched (Table 1) and showed a trend of higher expression in the R genotype in response to drought (Fig. 5a; supplementary Table S4). Notably, 15 transcripts annotated as DEAD-box ATP-dependent RNA helicase were upregulated in the roots, while none were downregulated. Ta.7609.1.S1_at that was upregulated in the R genotype was annotated as DEAD-box RNA helicase STRS1 (stress response suppressor 1). It was shown that mutations in this gene may attenuate the expression of stress-responsive transcriptional activators that function in ABA-dependent and ABA-independent abiotic stress-signalling networks (Kant et al. 2007). An additional type of post-transcriptional regulation in the R genotype is related to the nuclear cap-binding protein CBP80/ABH1 (ABA hypersensitive 1; DETR Ta.5831.2.S1_at) which encodes the large subunit (CBP80) of the nuclear cap-binding complex. RNA processing by ABH1/CBP80 contributes to ABA signal transduction as a negative regulator (Hugouvieux et al. 2001, 2002; Kuhn et al. 2008; Table 2).
Expression patterns of DETs associated with post-transcriptional regulation. a RNA binding proteins; b tRNAs. Well-watered control (C), water-stress (D). Red and green represent high and low relative expression when compared to the mean value of expression across all samples, respectively. Scale is log2 of mean expression value. Annotation of transcripts and full names of the putative genes are detailed in Table S4
Translation efficiency
‘RNA charging’ was one of the most significantly changed pathways in the R genotype (analysis conducted by the Plant MetGenMAP package). Therefore, the non-filtered DETs data were searched for the genes involved. Higher abundance of tRNA ligases were identified (Table S4); their pattern of regulation is presented by heat map (Fig. 5b). As described by Kwapisz et al. (2002), a higher level of tRNA is indicated to increase translational fidelity, but not the rate of translation, and independently may affect other cellular processes that contribute to the respiratory growth. An additional indication that may also contribute to translation fidelity is the downregulation of DETR Ta.2959.1.A1_at, annotated as Maf1, which is an essential and specific mediator of transcriptional repression in the RNA polymerase III system. Maf1-dependent repression occurs in response to a wide range of conditions, suggesting that the protein itself is targeted by the major nutritional and stress-signalling pathways (Ciesla et al. 2007; Hopper and Shaheen 2008).
Signalling and second messengers
Calcium signalling
Genes encoding proteins known as second messengers or as involved in signal transduction showed higher abundance in the R genotype in response to drought, including 14 transcripts annotated as calcium-dependent protein kinases and several types of calcium sensors, such as a calmodulin-related calcium sensor (DETR TaAffx.107453.1.S1_at) that was further validated by qPCR (Table S1 and Fig. S1), calcium binding protein (Ta.7626.3.S1_at), caleosine (Ta.9830.1.A1_at and Ta.7279.1.S1_at) and annexin (Ta.1577.1.S1_at and Ta.14590.3.S1_at). Caleosine is known as a calcium and phosphate binding protein, responsive to a range of environmental stresses and ABA in leaves and roots (Partridge and Murphy 2009). Annexin acts as a target of calcium signal in eukaryotic cells, binding phospholipid membranes in a calcium-dependent manner under drought stress in Arabidopsis (Konopka-Postupolska et al. 2009) and cold stress in wheat (Breton et al. 2000).
Phospholipid signalling
The phosphatidylinositol-4-phosphate 5-Kinase (PIP5K; TaAffx.66666.1.A1_at) showed higher upregulation in the R genotype. PIP5K is involved in signal transduction pathways and is also related to drought stress stimuli in leaves and roots (Lee et al. 2008b; Chen and Xiong 2010; Munnik and Vermeer 2010). Phosphatidylinositol (PtdIns) is a major phospholipid molecule in eukaryotic cells, and its phosphorylated derivatives are involved in many cellular processes in animal and plant cells, mainly in lipid signalling. It catalyses the phosphorylation of phosphatidylinositol-4-phosphate (PIP) to produce phosphatidylinositol-4,5-bisphosphate (PIP2). PIP2 is hydrolyzed by phosphatidylinositol-specific phospholipase C to produce two second messengers, inositol-1,4,5-triphosphate, which is involved in the release of Ca2+ from intracellular stores, and diacylglycerol (Mikami et al. 1998). Additionally, three transcripts (Affx.83351.1_at, Ta.24282.1.S1_at and Ta.465.1.S1_at) annotated as SEC14 proteins/phosphatidylinositol/phosphatidylcholine transfer proteins were upregulated in response to drought in the current study. Quantitative PCR (Fig. S1) showed that although the expression of DETR TaAffx.66666.1.A1_at was higher in the stressed R genotype, the difference was not significant. Nevertheless, metabolic analysis indicated a 1.5-fold increase in myo-inositol 1 phosphate in the R genotype (Supplementary Table S6), while there was no change in the S genotype, supporting the expression results and suggesting that myo-inositol metabolism was increased in the R genotype.
WD-40 proteins
Genes encoding proteins containing WD-40 repeats showed higher abundance in the R genotype under drought conditions; 23 WD-40 proteins were upregulated in the R genotype, as compared to only two in the S genotype. WD-40 repeat proteins play a key role in signal transduction, cytoskeletal dynamics, protein trafficking, nuclear export, ribosomal RNA biogenesis and especially in chromatin modification and transcription (van Nocker and Ludwig 2003), also recognized as involved in stress (Huang et al. 2008). In all proteins with WD-40 domains, the domains act as sites for interactions with other proteins, but the range of proteins with which they interact is large. Genetic analyses and yeast two-hybrid experiments have identified these partners as belonging to the bHLH and MYB classes of proteins (Ramsay and Glover 2005) that were also found to be up- or downregulated in response to drought in the current study.
Metabolomic profiling in wild emmer roots under well-watered and water-stress conditions
The GC-MS-based metabolic analysis data (presented as supplementary data Table S5) were first analysed by PCA (Saeed et al. 2004) which revealed a clear separation of the two genotypes based mainly on water regimes. The first component (PC1) represented the effect of drought, accounting for about 62% of the variance within the dataset (Fig. S2). Differences between genotypes were identified also under the well-watered conditions.
Metabolic differences between genotypes under well-watered conditions
One of the most significant differences between the genotypes included a 2-fold higher level of malate and related succinate in the R genotype as compared to the S genotype (Table 3). The R genotype was also characterized by very high levels (10-fold higher than the S genotype) of polyol that is recognized by the MS library as threitol or erythritol (Schauer et al. 2005). Polyols which are considered to be primary photosynthesis products are known to be produced under stress (Farooq et al. 2009). Therefore, their increased levels in the well-watered R genotype were unexpected. Plants have been found to accumulate polyols to compensate for reduced cell water potential under drought conditions (Noiraud et al. 2001). On the other hand, there are examples in the literature of resistant plants that have constitutively higher levels of stress-related metabolites, a feature possibly genetically evolved in the regulation of stress-related metabolic pathways (Kant et al. 2006). Interestingly, under the well-watered condition the amount of trehalose together with sugars such as maltose, raffinose and sucrose were relatively high in the S genotype as compared with the R genotype (Table S5). Many other metabolites had higher content in the S genotype under well-watered condition, including arginine, asparagine, cystathionine, galactinol, glucoheptose, glucuronic acid, glucopyranose and glycine (Table 3).
Metabolic differences between genotypes in response to drought
Drought caused significant (P < 0.05) alteration in the level of metabolites in both genotypes (Table 3): While the content of 23 metabolites increased significantly in the R genotype, the level of only 17 metabolites increased in the S genotype. Some of the metabolites displayed an opposite pattern of change, accumulating in the R genotype while decreasing in the S genotype, including citrate, glutarate, pyruvate, trehalose, glycine, maltitol and proline (Fig. 6). Whilst metabolite to transcript relations have been shown as non-linear in previous studies combining omics technologies, the accumulation of the above-mentioned metabolites could be partly supported by induction of metabolic genes. For example, accumulation of glucose was supported by the induction of sucrose synthase (SUS3 and SUS4; TaAffx.51190.1.S1_at) and sucrose-phosphate synthase (Ta.3972.1.S1_at). It was shown that the accumulation of glucose and increased SUS enzymes consume less energy, a feature that might be relevant under stress conditions (Baena-González 2010). The stress-related disaccharide trehalose increased 3-fold in the drought treated R genotype, twice the change measured in the S genotype (Fig. 6). Nonetheless, several transcripts (e.g. Ta.5372.1.S1_at, Ta.30416.1.S1_at) encoding for trehalose-6-phosphate phosphatase (TPP), which is the second enzyme in the trehalose biosynthesis, were downregulated in both genotypes growing under stress. The non-correlated transcriptomic and metabolomic results (e.g. accumulation of trehalose under drought and reduction of TPP) could aim at maintaining trehalose-6-phosphate (T6P) levels, which were shown to regulate sugar phosphates and sugar metabolism in general. Indeed, Schluepmann et al. (2003) showed that upon depletion of T6P, sugar phosphates accumulate. An alternative explanation to the opposite change of metabolite and transcript levels could involve post-translational factors in the regulation of metabolism (Fernie et al. 2004).
Changes in the contents of drought-related metabolites in the roots of the resistance (R) and the sensitive (S) genotypes in response to water-stress. Values represent the fold change to the mean response of the well-watered control, where the control is given the value 0. Values are representative of at least five biological replicates of each genotype. Pro proline, Gly glycine
Proline was accumulated (1.5-fold) in response to drought in the roots of the R genotype while a slight reduction was observed in the S genotype (Table S6). The results of transcriptome analysis indicated that transcript Ta.7091.1.S1_at annotated as delta 1 pyrroline-5-carboxylate synthase 2 (P5CS) was highly upregulated in both genotypes. P5CS is the first and the rate-limiting enzyme in proline biosynthesis in plants. This transcript was also highly upregulated in both genotypes in the leaves under drought. The non-correlated trend between the upregulation of P5CS in the S genotype and the lower accumulation of proline in this genotype may be also related to complex control mechanisms, at the transcriptional, post-transcriptional and post-translational levels of proline biosynthesis, not only by the activation of P5CS but also by feedback regulation of proline, the end product of the pathway that can influence the amount of proline in the cell (Hong et al. 2000). The accumulation of glycine (Table S5), which is the precursor of glycine betaine, was correlated with the activation of several transcripts annotated as betaine-aldehyde dehydrogenases (e.g. Ta.2585.1.S1_at) in the R genotype.
Alteration in the tricarboxylic acid cycle
The metabolic analysis indicated that the response to drought of the R genotype was characterized by altered levels of tricarboxylic acid (TCA) intermediates, including increased isocitrate, citrate, an outstanding increase of aconitate (5-fold increase) and a reduction in the amount of succinate and malate (Fig. 7a). High accumulation of citrate and concomitant decrease in malate content were also found in Arabidopsis roots under oxidative stress (Lehmann et al. 2009) and in drought-tolerant cotton under severe drought (Levi et al. 2011). As presented in Fig. 7a, b, the alteration in metabolites was correlated with the upregulation of the involved genes, including citrate synthase (Ta.970.1.S1_at), aconitate hydratase (TaAffx.6397.1.S1_s_at and Ta.10193.1.S1_at), malate dehydrogenase (Ta.1039.1.S1_at) and succinate dehydrogenase (Ta.23753.1.A1_at). The latter is related to electron transport which might be altered under stress conditions. Citrate, aconitate and isocitrate are upstream intermediates of 2-oxoglutarate which was also upregulated (Ta.28698.1.S1_at).
The TCA cycle—a metabolite accumulation and b gene expression patterns, identified in the roots of R and S genotypes in response to drought. A simplified figure of the TCA cycle, based on KEGG (http://www.genome.jp/kegg-bin/show_pathway?ko00020+C00417). The metabolite fold change level of the drought treatment relative to the control in the R and S genotypes are indicated by circle and square, respectively. The increase, decreased and no change fold levels of metabolites and transcripts indicated with colours green, red and black, respectively. The oval-shaped numbers indicate transcripts 1–6 that are presented by the heat map (in b) as revealed by microarray analysis correspond to the metabolic enzymes: 1 citrate hydro-lyase/aconitase, 2 2-oxoglutarate dehydrogenase, 3 citrate hydro-lyase/aconitase, 4 citrate synthase/CSY3, 5 lactate/malate dehydrogenase, 6 succinate dehydrogenase
Discussion
Comparative transcriptome and metabolome profiling revealed that drought stress induced major changes in the expression of genes involved in multilevel regulation, metabolism, energy and transport as well as in metabolite content, mainly in the roots of the R genotype. A similar trend of gene expression was previously found in the leaves of the same genotype (Krugman et al. 2010). However, a major shift in expression of genes associated with plant hormones, signalling and post-transcriptional regulation, were observed in the roots as compared with the leaves.
Cross talk between plant hormones results in synergetic or antagonist interactions play crucial roles in plant development and responses to abiotic stress (Wolters and Jurgens 2009). The beneficial effect of modifying hormone homeostasis for enhancement of abiotic stress tolerance in crop-plants was recently discussed by Peleg and Blumwald (2011). The biosynthesis of ABA in response to abiotic stress is one of the most studied pathways in plants. ABA is also involved in roots to shoots long-range signalling (Wilkinson and Davies 2010). In the current study, a higher abundance of genes related to ABA metabolism (ZEP, AAO1 and AAO3) and regulation (OST1, PP2C, HAB1 and HAB2) was found in the R genotype. Quantification of ABA in the roots further supported higher ABA levels in the roots of R genotype under both irrigation regimes, with increased levels under drought stress (Table 2; Figs. 3 and 4). ABA was shown to stimulate changes in Ca2+ in a number of plant tissues, including stomata guard cells and in roots. Furthermore, phosphoinositide signalling in plants is known to be involved in the release of Ca2+ from cellular stores (Stenzel et al. 2008; Munnik and Vermeer 2010). In agreement with that, several types of calcium binding proteins (calmodulin, caleosine and annexin) and phosphatidylinositol signalling (PIP5K and SEC14) were also upregulated in the R genotype.
The R genotype also exhibited an alteration in expression of genes associated with GA biosynthesis (GA2OX1) and response (CIGR1; XERICO). Induction of the GA receptor (GID1L2) in the R genotype suggests that GA may play a role in stress tolerance in the roots. Binding of GA to GID1 promotes interaction of GID1 with DELLA proteins, a subsequent poly-ubiquitination of DELLAs via the E3 ubiquitin-ligase SCF SLY1 and the eventual destructions of DELLAs. Furthermore, it is known that non-destructed DELLAs exert general plant inhibitory activity by both reducing cell proliferation and expansion rates and that accumulation of DELLA proteins repress root meristem size (Murase et al. 2008; Achard et al. 2009; Ubeda-Tomás et al. 2008; Sun 2010). Fundamental research has revealed that most of the dwarfing genes were involved in the GA biosynthetic and signal transduction pathway in cereals (Jia et al. 2009). The effects of GA on wheat root systems are not clear (Wojciechowski et al. 2009 and reference therein), and further study will be needed to unravel its role. It is worth noting that genes annotated as auxin-related genes (e.g. OsSAUR37- and OsSAUR38-responsive genes; auxin efflux carrier), being key compounds in the fine tuning of hormone signalling in the roots (Malamy 2005; Seo and Park 2009; Herder et al. 2010), showed in the current study differential expression in both genotypes in response to drought (Table 2). Recently, it was shown that the initiation and elongation of lateral roots is associated with a cross talk between auxin and ABA (Park et al. 2009) and between GA and IAA (Gou et al. 2010). The alteration in expression of genes related to biosynthesis and signalling of ABA, GA and IAA identified in the current study can support hormonal shifts in the roots in response to drought. Furthermore, cross talk between the three plant hormones is indicated by the expression patterns of RD22 and PM19. The transcription factor MADS-box26 was induced under drought stress in the roots, as well as in the leaves (Krugman et al. 2010) of the R genotype. Similarly, MADS-box26 was upregulated in rice plants exposed to dehydration stress (Arora et al. 2007; Diaz-Riquelme et al. 2009). MAD-box26 is required for normal root development and was reported to be induced by auxin (Tapia-Lopez et al. 2008). Transgenic rice plants overexpressing the OsMADS26 gene showed increased expression of JA and ethylene biosynthesis (Lee et al. 2008a). The transgenics were similar to those previously reported for plants exposed to various stresses including diminished leaf, shoot and root elongation, as well as enhanced formation of lateral roots. These authors reported that, unexpectedly, a gene with high homology to GA β-hydroxylase, which converts GA20 to the active GA1, was also upregulated in the OsMADS26-overexpressing plants. However, they did not study the relationship between GA and OsMADS26 because GA is rarely indicated as involved in stress-related responses.
Genes involved in post-transcriptional regulation and translation efficiency had higher representation in the roots of the same R genotype under drought stress as compared with the leaves (Krugman et al. 2010). These genes included RNA binding proteins and RNA helicases, which are the two main protein families determining the fate of pre-mRNAs and mRNAs from transcription to translation (reviewed by Mazzucotelli et al. 2008). Interestingly, the cap-binding protein CBP80/ABH1 was upregulated in the roots of the R genotype. This gene was shown previously to be involved in ABA signal transduction as a negative regulator (Hugouvieux et al. 2001, 2002; Kuhn et al. 2008).
The R genotype was previously shown to have higher photosynthetic capacity compared with the S genotype (Avneri 2009). The accumulation of malate and polyols found in the roots of R genotype may reflect this advantage. Malate has a central role in controlling the activity of enzymes of primary metabolism through the TCA cycle (Charlton et al. 2008). The TCA cycle is a central element of carbon metabolism in higher plants, which provides among other things, electrons for oxidative phosphorylation in the inner mitochondrial membrane, intermediates for amino acid biosynthesis and oxaloacetate for gluconeogenesis from succinate derived from fatty acids via the glyoxylate cycle in glyoxysomes (Fernie et al. 2004). Several studies have shown a correlation between TCA cycle activity and root growth (e.g. Carrari et al. 2003; Nunes-Nesi et al. 2005, 2007; Studart-Guimarães et al. 2007; Sienkiewicz-Porzucek et al. 2008). Metabolomic analysis of Arabidopsis plants treated with several individual phytohormones, such as ABA, GA, IAA and salicylic acid, identified that ABA treatment caused a major metabolic change including reduction in malate, fumarate and succinate and increased citrate and isocitrate (Okamoto et al. 2009), similar to the findings of the current study.
Integrating transcriptome and metabolome profiles reveal that ABA signalling lead to appropriate physiological, metabolic, transcriptional and translational modifications (Urano et al. 2009; Rao et al. 2010). In the current study, several metabolites, including glucose, trehalose and amino acids proline and glycine (a precursor of glycine betaine), that were shown to be associated with protecting cellular functions or with maintaining the structure of cellular components under stress were increased in the R genotype (Elbein et al. 2003; Seki et al. 2007; Foito et al. 2009). Proline which is ABA-dependent (Urano et al. 2009) and glycine betaine are known as stress-related metabolites, and in general, the increased amino acid content represents a response of the plant to stress (Bartels and Sunkar 2005; Chen and Murata 2008). The metabolites accumulating under water-stress conditions can function also as osmolytes, signalling molecules, antioxidants, scavengers and membrane-protective molecules that help plants to avoid or to tolerate dehydration stress (Bartels and Sunkar 2005).
In conclusion, integration of transcriptome and metabolome profiling reveal genomic changes involved in drought-adaptive mechanisms in wild emmer wheat. Alteration in expression of genes involved in hormonal balance in the roots were associated with improved drought tolerance. Our results demonstrate the potential of the wild emmer genepool as a source for novel genes to improve drought tolerance in modern wheat cultivars. Since the wheat genome remains mostly unknown, future advancement is wheat sequencing will enable the identification of novel genes for drought tolerance.
References
Abe H, Urao T, Ito T, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signalling. Plant Cell 15:63–78
Achard P, Gusti A, Cheminant S, Alioua M, Dhondt S, Coppens F, Beemster GTS, Genschik P (2009) Gibberellin signalling controls cell proliferation rate in Arabidopsis. Curr Biol 19:1188–1193
Arora R, Agarwal P, Ray S, Singh AK, Singh VP, Tyagi AK, Kapoor S (2007) MADS-box gene family in rice: genome-wide identification, organization and expression profiling during reproductive development and stress. BMC Genomics 8:242
Avneri A (2009) Drought resistance and water use efficiency in wild emmer wheat (Triticum turgidum ssp. dicoccoides). M.Sc. dissertation, The Robert H. Smith Faculty of Agriculture Food and Environment, The Hebrew University of Jerusalem (Hebrew with English abstract)
Baena-González E (2010) Energy signalling in the regulation of gene expression during stress. Mol Plant 3:300–313
Bartels D, Sunkar R (2005) Drought and salt tolerance in plants. Crit Rev Plant Sci 24:23–58
Benfey PN, Bennett M, Schiefelbein J (2010) Getting to the root of plant biology: impact of the Arabidopsis genome sequence on root research. Plant J 61:992–1000
Birnbaum K, Shasha DE, Wang JY, Jung JW, Lambert GM, Galbraith DW, Benfey PN (2003) A gene expression map of the Arabidopsis root. Science 302:1956–1960
Brady SM, Orlando DA, Lee JY, Wang JY, Koch J, Dinneny JR, Mace D, Ohler U, Benfey PN (2007) A high-resolution root spatiotemporal map reveals dominant expression patterns. Science 318:801–806
Breton G, Vazquez-Tello A, Danyluk J, Sarhan F (2000) Two novel intrinsic annexins accumulate in wheat membranes in response to low temperature. Plant Cell Physiol 41:177–184
Carrari F, Nunes-Nesi A, Gibon Y, Lytovchenko A, Loureiro ME, Fernie AR (2003) Reduced expression of aconitase results in an enhanced rate of photosynthesis and marked shifts in carbon partitioning in illuminated leaves of wild species tomato. Plant Physiol 133:1322–1335
Cattivelli L, Rizza F, Badeck FW, Mazzucotelli E, Mastrangelo AM, Francia E, Mare C, Tondelli A, Stanca AM (2008) Drought tolerance improvement in crop plants: an integrated view from breeding to genomics. Field Crops Res 105:1–14
Charlton AJ, Donarski JA, Harrison M, Jones SA, Godward J, Oehlschlager S, Arques JL, Ambrose M, Chinoy C, Mullineaux PM (2008) Responses of the pea (Pisum sativum L.) leaf metabolome to drought stress assessed by nuclear magnetic resonance spectroscopy. Metabolomics 4:312–327
Chen THH, Murata N (2008) Glycinebetaine: an effective protectant against abiotic stress in plants. Trends Plant Sci 13:499–505
Chen H, Xiong L (2010) myo-Inositol-1-phosphate synthase is required for polar auxin transport and organ development. Biol Chem 285:24238–24247
Choi DW, Zhu B, Close T (1999) The barley (Hordeum vulgare L.) dehydrin multigene family: sequences, allele types, chromosome assignments, and expression characteristics of 11 Dhn genes of cv Dicktoo. Theor Appl Genet 98:1234–1247
Choulet F, Wicker T, Rustenholz C, Paux E, Salse J, Leroy P, Schlub S, Le Paslier MC, Magdelenat G, Gonthier C (2010) Megabase level sequencing reveals contrasted organization and evolution patterns of the wheat gene and transposable element spaces. Plant Cell 22:1686–1701
Ciesla M, Towpik J, Graczyk D, Oficjalska-Pham D, Harismendy O, Suleau A, Balicki K, Conesa C, Lefebvre O, Boguta M (2007) Maf1 is involved in coupling carbon metabolism to RNA polymerase III transcription. Mol Cell Biol 27:7693–7702
Cui H, Benfey PN (2009) Interplay between SCARECROW, GA and like heterochromatin protein 1 in ground tissue patterning in the Arabidopsis root. Plant J 58:1016–1027
Day RB, Tanabe S, Koshioka M, Mitsui T, Itoh H, Ueguchi-Tanaka M, Matsuoka M, Kaku H, Shibuya N, Minami E (2004) Two rice GRAS family genes responsive to N-acetylchitooligosaccharide elicitor are induced by phytoactive gibberellins: evidence for cross-talk between elicitor and gibberellin signalling in rice cells. Plant Mol Biol 54:261–272
Diaz-Riquelme J, Lijavetzky D, Martinez-Zapater JM, Carmona MJ (2009) Genome-wide analysis of MIKCC-type MADS box genes in grapevine. Plant Physiol 149:354–369
Elbein AD, Pan Y, Pastuszak I, Carroll D (2003) New insights on trehalose: a multifunctional molecule. Glycobiol 13:17R
Erban A, Schauer N, Fernie AR, Kopka J (2006) Nonsupervised construction and application of mass spectral and retention time index libraries from time-of-flight gas chromatography-mass spectrometry metabolite profiles. Methods Mol Biol 358:19–38
Farooq M, Wahid A, Kobayashi N, Fujita D, Basra S (2009) Plant drought stress: effects, mechanisms and management. Sustainable Agric 1:153–188
Feldman M (2001) The origin of cultivated wheat. In: Bonjean A, Angus W (eds) The world wheat book. Lavoisier Technical and Doc, Paris, pp 3–56
Fernie AR, Carrari F, Sweetlove LJ (2004) Respiratory metabolism: glycolysis, the TCA cycle and mitochondrial electron transport. Curr Opin Plant Biol 7:254–261
Foito A, Byrne SL, Shepherd T, Stewart D, Barth S (2009) Transcriptional and metabolic profiles of Lolium perenne L. genotypes in response to a PEG-induced water stress. Plant Biotech J 7:719–732
Gou J, Strauss SH, Tsai CJ, Fang K, Chen Y, Jiang X, Busov VB (2010) Gibberellins regulate lateral root formation in Populus through interactions with auxin and other hormones. Plant Cell 22:623–639
Gubler F, Raventos D, Keys M, Watts R, Mundy J, Jacobsen JV (1999) Target genes and regulatory domains of the GAMYB transcriptional activator in cereal aleurone. Plant J 17:1–9
Harrison BR, Masson PH (2008) ARL2, ARG1 and PIN3 define a gravity signal transduction pathway in root statocytes. Plant J 53:380–392
Herder GD, Van Isterdael G, Beeckman T, De Smet I (2010) The roots of a new green revolution. Trends Plant Sci 15:600–607
Hochberg Y (1988) A sharper Bonferroni procedure for multiple tests of significance. Biometrika 75:800–803
Hong Z, Lakkineni K, Zhang Z, Verma DPS (2000) Removal of feedback inhibition of delta 1-pyrroline-5-carboxylate synthetase results in increased proline accumulation and protection of plants from osmotic stress. Plant Physiol 122:1129–1136
Hopper AK, Shaheen HH (2008) A decade of surprises for tRNA nuclear–cytoplasmic dynamics. Trends Cell Biol 18:98–104
Huang J, Wang MM, Bao YM, Sun SJ, Pan LJ, Zhang HS (2008) SRWD: a novel WD40 protein subfamily regulated by salt stress in rice (Oryza sativa L.). Gene 424:71–79
Hugouvieux V, Kwak JM, Schroeder JI (2001) An mRNA cap binding protein, ABH1, modulates early abscisic acid signal transduction in Arabidopsis. Cell 106:477–487
Hugouvieux V, Murata Y, Young JJ, Kwak JM, Mackesy DZ, Schroeder JI (2002) Localization, ion channel regulation, and genetic interactions during abscisic acid signalling of the nuclear mRNA cap-binding protein, ABH1. Plant Physiol 130:1276–1287
Iglesias MJ, Terrile MC, Bartoli CG, D’Ippólito S, Casalongué CA (2010) Auxin signaling participates in the adaptative response against oxidative stress and salinity by interacting with redox metabolism in Arabidopsis. Plant Mol Biol 74:215–222
Jia Q, Zhang J, Westcott S, Zhang XQ, Bellgard M, Lance R, Li C (2009) GA-20 oxidase as a candidate for the semidwarf gene sdw1/denso in barley. Funct Integr Genomics 9:255–262
Jiao Y, Tausta SL, Gandotra N, Sun N, Liu T, Clay NK, Ceserani T, Chen M, Ma L, Holford M (2009) A transcriptome atlas of rice cell types uncovers cellular, functional and developmental hierarchies. Nat Genet 41:258–263
Joung JG, Corbett AM, Fellman SM, Tieman DM, Klee HJ, Giovannoni JJ, Fei Z (2009) Plant MetGenMAP: an integrative analysis system for plant systems biology. Plant Physiol 151:1758–1768
Kamiya N, Itoh JI, Morikami A, Nagato Y, Matsuoka M (2003) The SCARECROW gene’s role in asymmetric cell divisions in rice plants. Plant J 36:45–54
Kant S, Kant P, Raveh E, Barak S (2006) Evidence that differential gene expression between the halophyte, Thellungiella halophila, and Arabidopsis thaliana is responsible for higher levels of the compatible osmolyte proline and tight control of Na+ uptake in T. halophila. Plant Cell Environ 29:1220–1234
Kant P, Kant S, Gordon M, Shaked R, Barak S (2007) STRESS RESPONSE SUPPRESSOR1 and STRESS RESPONSE SUPPRESSOR2, two DEAD-box RNA helicases that attenuate Arabidopsis responses to multiple abiotic stresses. Plant Physiol 145:814–830
Koiwai H, Nakaminami K, Seo M, Mitsuhashi W, Toyomasu T, Koshiba T (2004) Tissue-specific localization of an abscisic acid biosynthetic enzyme, AAO3, in Arabidopsis 1. Plant Physiol 134:1697–1707
Konopka-Postupolska D, Clark G, Goch G, Debski J, Floras K, Cantero A, Fijolek B, Roux S, Hennig J (2009) The role of annexin 1 in drought stress in Arabidopsis. Plant Physiol 150:1394–1410
Krugman T, Chagué V, Peleg Z, Balzergue S, Just J, Korol AB, Nevo E, Saranga Y, Chalhoub B, Fahima T (2010) Multilevel regulation and signalling processes associated with adaptation to terminal drought in wild emmer wheat. Funct Integr Genomic 10:167–186
Kuhn J, Hugouvieux V, Schroeder J (2008) mRNA cap binding proteins: effects on abscisic acid signal transduction, mRNA processing, and microarray analyses. Curr Top Microbiol Immunol 326:139–150
Kwapisz M, Smagowicz WJ, Oficjalska D, Hatin I, Rousset JP, Żołądek T, Boguta M (2002) Up-regulation of tRNA biosynthesis affects translational readthrough in maf1-Δ mutant of Saccharomyces cerevisiae. Curr Genet 42:147–152
Lee TG, Jang CS, Kim JY, Kim DS, Park JH, Kim DY, Seo YW (2007) A Myb transcription factor (TaMyb1) from wheat roots is expressed during hypoxia: roles in response to the oxygen concentration in root environment and abiotic stresses. Physiol Plant 129:375–385
Lee S, Woo YM, Ryu SI, Shin YD, Kim WT, Park KY, Lee IJ, An G (2008a) Further characterization of a rice AGL12 group MADS-box gene, OsMADS26. Plant Physiol 147:156–168
Lee Y, Bak G, Choi Y, Chuang WI, Cho HT, Lee Y (2008b) Roles of phosphatidylinositol 3-kinase in root hair growth. Plant Physiol 147:624–635
Legay S, Lefèvre I, Lamoureux D, Barreda C, Tincopa R, Raymundo L, Quiroz R, Hoffmann L, Hausman JF, Bonierbale M, Evers D, Schafleitner R (2011) Carbohydrate metabolism and cell protection mechanisms differentiate drought tolerance and sensitivity in advanced potato clones (Solanum tuberosum L.). Funct Integr Genomic 11:275–291
Lehmann M, Schwarzlander M, Obata T, Sirikantaramas S, Burow M, Olsen CE, Tohge T, Fricker MD, Moller BL, Fernie AR (2009) The metabolic response of Arabidopsis roots to oxidative stress is distinct from that of heterotrophic cells in culture and highlights a complex relationship between the levels of transcripts, metabolites, and flux. Mol Plant 2:390–406
Levi A, Paterson AH, Cakmak I, Saranga Y (2011) Metabolite and mineral analyses of cotton near-isogenic lines introgressed with QTLs for productivity and drought-related traits. Physiol Plant 141:265–275
Levin M, Resnick N, Rosianskey Y, Kolotilin I, Wininger S, Lemcoff JH, Cohen S, Galili G, Koltai H, Kapulnik Y (2009) Transcriptional profiling of Arabidopsis thaliana plants’ response to low relative humidity suggests a shoot–root communication. Plant Sci 177:450–459
Lisec J, Schauer N, Kopka J, Willmitzer L, Fernie AR (2006) Gas chromatography mass spectrometry-based metabolite profiling in plants. Nat Protoc 1:387–396
López-Carbonell M, Jáuregui O (2005) A rapid method for the analysis of abscisic acid (ABA) in crude extracts of water stressed Arabidopsis thaliana plants by liquid chromatography-mass spectrometry in tandem mode. Plant Physiol Biochem 43:407–411
Lu G, Gao C, Zheng X, Han B (2009) Identification of OsbZIP72 as positive regulator of ABA response and drought tolerance in rice. Planta 229:605–615
Luedemann A, Strassburg K, Erban A, Kopka J (2008) TagFinder for the quantitative analysis of gas chromatography–mass spectrometry (GC-MS)-based metabolite profiling experiments. Bioinformatics 24:732–737
Malamy J (2005) Intrinsic and environmental response pathways that regulate root system architecture. Plant Cell Environ 28:67–77
Manschadi AM, Hammer GL, Christopher JT, Devoil P (2008) Genotypic variation in seedling root architectural traits and implications for drought adaptation in wheat (Triticum aestivum L.). Plant Soil 303:115–129
Mashiguchi K, Urakami E, Hasegawa M, Sanmiya K, Matsumoto I, Yamaguchi I, Asami T, Suzuki Y (2008) Defense-related signalling by interaction of arabinogalactan proteins and β-glucosyl Yariv reagent inhibits gibberellin signalling in barley aleurone cells. Plant Cell Physiol 49:178–190
Mazzucotelli E, Mastrangelo AM, Crosatti C, Guerra D, Stanca AM, Cattivelli L (2008) Abiotic stress response in plants: when post-transcriptional and post-translational regulations control transcription. Plant Sci 174:420–431
Merlot S, Leonhardt N, Fenzi F, Valon C, Costa M, Piette L, Vavasseur A, Genty B, Boivin K, Müller A (2007) Constitutive activation of a plasma membrane H(+)-ATPase prevents abscisic acid-mediated stomatal closure. EMBO J 26:3216
Mikami K, Katagiri T, Iuchi S, Yamaguchi-Shinozaki K, Shinozaki K (1998) A gene encoding phosphatidylinositol–4–phosphate 5–kinase is induced by water stress and abscisic acid in Arabidopsis thaliana. Plant J 15:563–568
Mittler R, Blumwald E (2010) Genetic engineering for modern agriculture: challenges and perspectives. Ann Rev Plant Biol 61:443–462
Miyazawa Y, Takahashi A, Kobayashi A, Kaneyasu T, Fujii N, Takahashi H (2009) GNOM-mediated vesicular trafficking plays an essential role in hydrotropism of Arabidopsis roots. Plant Physiol 149:835–840
Miyazono K, Miyakawa T, Sawano Y, Kubota K, Kang HJ, Asano A, Miyauchi Y, Takahashi M, Zhi Y, Fujita Y (2009) Structural basis of abscisic acid signalling near-final version. Nature 462:609–614
Munnik T, Vermeer JEM (2010) Osmotic stress-induced phosphoinositide and inositol phosphate signalling in plants. Plant Cell Environ 33:655–669
Murase K, Hirano Y, Sun T, Hakoshima T (2008) Gibberellin-induced DELLA recognition by the gibberellin receptor GID1. Nature 456:459–463
Nemhauser JL, Hong F, Chory J (2006) Different plant hormones regulate similar processes through largely nonoverlapping transcriptional responses. Cell 126:467–475
Nevo E, Korol AB, Beiles A, Fahima T (2002) Evolution of wild emmer and wheat improvement: population genetics, genetic resources and genome organization of wheat’s progenitor, Triticum dicoccoides. Springer, Berlin
Nibau C, Gibbs D, Coates J (2008) Branching out in new directions: the control of root architecture by lateral root formation. New Phytol 179:595–614
Noiraud N, Maurousset L, Lemoine R (2001) Transport of polyols in higher plants. Plant Physiol Biochem 39:717–728
Nunes-Nesi A, Carrari F, Lytovchenko A, Smith AMO, Loureiro ME, Ratcliffe RG, Sweetlove LJ, Fernie AR (2005) Enhanced photosynthetic performance and growth as a consequence of decreasing mitochondrial malate dehydrogenase activity in transgenic tomato plants. Plant Physiol 137:611–622
Nunes-Nesi A, Carrari F, Gibon Y, Sulpice R, Lytovchenko A, Fisahn J, Graham J, Ratcliffe RG, Sweetlove LJ, Fernie AR (2007) Deficiency of mitochondrial fumarase activity in tomato plants impairs photosynthesis via an effect on stomatal function. Plant J 50:1093–1106
Okamoto M, Tsuboi Y, Chikayama E, Kikuchi J, Hirayama T (2009) Metabolic movement upon abscisic acid and salicylic acid combined treatments. Plant Biotech 26:551–560
Paolacci AR, Tanzarella OA, Porceddu E, Ciaffi M (2009) Identification and validation of reference genes for quantitative RT-PCR normalization in wheat. BMC Mol Biol 10:11
Park SY, Fung P, Nishimura N, Jensen DR, Fujii H, Zhao Y, Lumba S, Santiago J, Rodrigues A, Chow TF (2009) Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 324:1068–1071
Partridge M, Murphy DJ (2009) Roles of a membrane-bound caleosin and putative peroxygenase in biotic and abiotic stress responses in Arabidopsis. Plant Physiol Biochem 47:796–806
Peleg Z, Blumwald E (2011) Hormone balance and abiotic stress tolerance in crop plants. Curr Op Plant Biol. doi:10.1016/j.pbi.2011.02.001
Peleg Z, Fahima T, Abbo S, Krugman T, Nevo E, Yakir D, Saranga Y (2005) Genetic diversity for drought resistance in wild emmer wheat and its ecogeographical associations. Plant Cell Environ 28:176–191
Peleg Z, Fahima T, Saranga Y (2007) Drought resistance in wild emmer wheat: physiology, ecology, and genetics. Israel J Plant Sci 55:289–296
Peleg Z, Fahima T, Krugman T, Abbo S, Yakir D, Korol AB, Saranga Y (2009) Genomic dissection of drought resistance in durum wheat × wild emmer wheat RIL population. Plant Cell Environ 10:1365–3040
Pysh LD, Wysocka-Diller JW, Camilleri C, Bouchez D, Benfey PN (2002) The GRAS gene family in Arabidopsis: sequence characterization and basic expression analysis of the SCARECROW-LIKE genes. Plant J 18:111–119
Ramsay NA, Glover BJ (2005) MYB-bHLH-WD40 protein complex and the evolution of cellular diversity. Trends Plant Sci 10:63–70
Ranford JC, Bryce JH, Morris PC (2002) PM19, a barley (Hordeum vulgare L.) gene encoding a putative plasma membrane protein, is expressed during embryo development and dormancy. Exp Bot 53:147–148
Rao SR, Ford KL, Cassin AM, Roessner U, Patterson JH, Bacic A (2010) Proteomic and metabolic profiling of rice suspension culture cells as a model to study abscisic acid (ABA) signalling response pathways in plants. Proteome Res 9:6623–6634
Roessner U, Luedemann A, Brust D, Fiehn O, Linke T, Willmitzer L, Fernie AR (2001) Metabolic profiling allows comprehensive phenotyping of genetically or environmentally modified plant systems. Plant Cell 13:11–29
Saeed AI, Sharov V, White J, Li J, Liang W, Bhagabati N, Braisted J, Klapa M, Currier T, Thiagarajan M, Sturn A, Snuffin M, Rezantsev A, Popov D, Ryltsov A, Kostukovich E, Borisovsky I, Liu Z, Vinsavich A, Trush V, Quackenbush J (2004) TM4: a free, open-source system for microarray data management and analysis. Bio-technique 34:374–378
Saeed AI, Hagabati NK, Braisted JC, Liang W, Sharov V, Howe EA, Li JW, Thiagarajan M, White JA, Quackenbush J (2006) TM4 microarray software suite. In: DNA microarrays, part B: databases and statistics vol 411. Academic, New York, pp 134–193
Schauer N, Zamir, Fernie AR (2005) Metabolic profiling of leaves and fruit of wild species tomato: a survey of the Solanum lycopersicum complex. J Exp Bot 56:297–307
Schluepmann H, Pellny T, van Dijken A, Smeekens S, Paul M (2003) Trehalose 6-phosphate is indispensable for carbohydrate utilization and growth in Arabidopsis thaliana. Proc Natl Acad Sci USA 100:6849–6854
Schroeder JI, Kwak JM, Allen GJ (2001) Guard cell abscisic acid signalling and engineering drought hardiness in plants. Nature 410:327–330
Seki M, Umezawa T, Urano K, Shinozaki K (2007) Regulatory metabolic networks in drought stress responses. Curr Opin Plant Biol 10:296–302
Seo PJ, Park CM (2009) Auxin homeostasis during lateral root development under drought condition. Plant Signalling Behav 4:1002–1004
Seo PJ, Xiang F, Qiao M, Park JY, Lee YN, Kim SG, Lee YH, Park WJ, Park CM (2009) The MYB96 transcription factor mediates abscisic acid signalling during drought stress response in Arabidopsis. Plant Physiol 151:275–289
Sienkiewicz-Porzucek A, Nunes-Nesi A, Sulpice R, Lisec J, Centeno DC, Carillo P, Leisse A, Urbanczyk-Wochniak E, Fernie AR (2008) Mild reductions in mitochondrial citrate synthase activity result in a compromised nitrate assimilation and reduced leaf pigmentation but have no effect on photosynthetic performance or growth. Plant Physiol 147:115–127
Skern R, Frost P, Nilsen F (2005) Relative transcript quantification by quantitative PCR: roughly right or precisely wrong? BMC Mol Biol 6:10
Stenzel I, Ischebeck T, Konig S, Holubowska A, Sporysz M, Hause B, Heilmann I (2008) The type B phosphatidylinositol-4-phosphate 5-kinase 3 is essential for root hair formation in Arabidopsis thaliana. Plant Cell 20:124–141
Studart-Guimarães FA, Nunes-Nesi A, Carrari F, Usadel B, Fernie AR (2007) Reduced expression of succinyl-coenzyme A ligase can be compensated for by up-regulation of the γ-aminobutyrate shunt in illuminated tomato leaves. Plant Physiol 145:626–639
Sun T (2010) Gibberellin-GID1-DELLA: a pivotal regulatory module for plant growth and development. Plant Physiol 154:567–570
Suprunova T, Krugman T, Fahima T, Chen G, Shams I, Korol A, Nevo E (2004) Differential expression of dehydrin genes in wild barley, Hordeum spontaneum, associated with resistance to water deficit. Plant Cell Environ 27:1297–1308
Tanksley SD, McCouch SR (1997) Seed banks and molecular maps: unlocking genetic potential from the wild. Science 277:1063–1066
Tapia-Lopez R, Garcia-Ponce B, Dubrovsky JG, Garay-Arroyo A, Perez-Ruiz RV, Kim SH, Acevedo F, Pelaz S, Alvarez-Buylla ER (2008) An AGAMOUS-related MADS-box gene, XAL1 (AGL12), regulates root meristem cell proliferation and flowering transition in Arabidopsis. Plant Physiol 146:1182–1192
Thompson AJ, Andrews J, Mulholland BJ, McKee JMT, Hilton HW, Horridge JS, Farquhar GD, Smeeton RC, Smillie IRA, Black CR, Taylor IB (2007) Overproduction of abscisic acid in tomato increases transpiration efficiency and root hydraulic conductivity and influences leaf expansion. Plant Physiol 143:1905–1917
Thornton TM, Swain SM, Olszewski NE (1999) Gibberellin signal transduction presents… the SPY who O-GlcNAc’d me. Trends Plant Sci 4:424–428
Ubeda-Tomás S, Swarup R, Coates J, Swarup K, Laplaze L, Beemster GTS, Hedden P, Bhalerao R, Bennett MJ (2008) Root growth in Arabidopsis requires gibberellin/DELLA signalling in the endodermis. Nat Cell Biol 10:625–628
Urano K, Maruyama K, Ogata Y, Morishita Y, Takeda M, Sakurai N, Suzuki H, Saito K, Shibata D, Kobayashi M (2009) Characterization of the ABA–regulated global responses to dehydration in Arabidopsis by metabolomics. Plant J 57:1065–1078
van Nocker S, Ludwig P (2003) The WD-repeat protein superfamily in Arabidopsis: conservation and divergence in structure and function. BMC Genomics 4:50
Wilkinson S, Davies WJ (2010) Drought, ozone, ABA and ethylene: new insights from cell to plant to community. Plant Cell Environ 33:510–525
Wojciechowski T, Gooding M, Ramsay L, Gregory P (2009) The effects of dwarfing genes on seedling root growth of wheat. J Exp Bot 60:2565–2573
Wolters H, Jurgens G (2009) Survival of the flexible: hormonal growth control and adaptation in plant development. Nat Rev Genet 10:305–317
Xu H, Li Y, Yan Y, Wang K, Gao Y, Hu Y (2010) Genome-scale identification of soybean BURP domain-containing genes and their expression under stress treatments. BMC Plant Biol 10:197
Yamaguchi Y, Takeda-Kamiya N, Hanada A, Ogawa M, Kuwahara A, Seo M, Kamiya Y, Yamaguchi S (2007) Contribution of gibberellin deactivation by AtGA2ox2 to the suppression of germination of dark-imbibed Arabidopsis thaliana seeds. Plant Cell Physiol 48:555–561
Yamaguchi-Shinozaki K, Shinozaki K (1993) The plant hormone abscisic acid mediates the drought-induced expression but not the seed-specific expression of rd22, a gene responsive to dehydration stress in Arabidopsis thaliana. Mol Gen Genet 238:17–25
Yoshida T, Fujita Y, Sayama H, Kidokoro S, Maruyama K, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2010) AREB1, AREB2, and ABF3 are master transcription factors that cooperatively regulate ABRE-dependent ABA signalling involved in drought stress tolerance and require ABA for full activation. Plant J 61:672–685
Acknowledgments
This project was supported by the Program for Sustainable Agriculture funded by the Israel Ministry of Science (# 01-21-00048), the French Ministry for Foreign Affairs and the French Ministry for Education and Research. We also acknowledge the Israel Science Foundation grant #1089/04 and equipment grants #048/99 and 1478/04. Z. Peleg is indebted to the Israel Council for the Higher Education Postdoctoral Fellowships Award. The authors thank A. Fahum, M. Goldshmit and S. Khalifa for their excellent technical assistance.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary materials
Below is the link to the electronic supplementary material.
Fig. S2
Principal component analysis (PCA) of GC-MS data of two emmer wheat genotypes (R and S) subjected to control (well watered) and drought (withholding water for 7 days). PCA is presented as the combinations of first three dimensions. Rectangle shape filled and not filled represent R control and drought, respectively; circle filled and not filled represent S control and drought, respectively. Values in bracket represent the value of addition of the two components contribution to the variance. Each data point represents an independent sample and biological replicate of at least five samples. (PDF 18 kb)
Table S1
Probe-set primers of DETs used to validate expression patterns by quantitative real-time PCR (PDF 15 kb)
ESM 1
(XLSX 57 kb)
ESM 2
(XLSX 48 kb)
ESM 3
(XLSX 20 kb)
Rights and permissions
About this article
Cite this article
Krugman, T., Peleg, Z., Quansah, L. et al. Alteration in expression of hormone-related genes in wild emmer wheat roots associated with drought adaptation mechanisms. Funct Integr Genomics 11, 565–583 (2011). https://doi.org/10.1007/s10142-011-0231-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-011-0231-6