Abstract
Quantitative structure activity relationship (QSAR) modeling has been in use for several decades now. One branch of it, in silico ADMET, became more and more important since the late 1990s as studies indicated that poor pharmacokinetics and toxicity were important causes of costly late-stage failures in drug development. In this paper we describe some of the available methods and best practice for the different stages of the in silico model building process. We also describe some more recent developments, like automated model building and the prediction probability. Finally we will discuss the use of in silico ADMET for “big data” and the importance and possible further development of interpretable models.

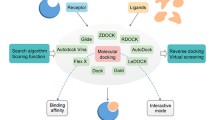
Schematic ADMET model generation process








Similar content being viewed by others
References
Richet MC (1893) Note sur le rapport entre la toxicité et les propriétés physiques des corps. C R Soc Biol 45:775–776
Fischer E (1894) Einfluss der Configuration auf die Wirkung der Enzyme. Ber Dtsch Chem Ges 273:2985–2993
Overton E (1901) Studien über die Narkose. Gustav Fischer, Jena
Meyer H (1899) Zur Theorie der Alkoholnarkose. Arch Exp Pathol Pharmakol 42:109–118
Hansch C, Maloney P, Fujita T, Muir R (1962) Correlation of biological activity of phenoxyacetic acids with Hammett substituent constants and partition coefficients. Nature 194:178–180
Hansch C, Sammes PG, Taylor JB (1990) Quantitative Drug Design. In: Ramsden CA (ed) Comprehensive Medicinal Chemistry. Pergamon Press, Oxford
Kubiniyi H (1993) QSAR. Hansch analysis and related approaches. 1. In: Mangold R, Krosgaard Larsen P, Timmermann H (eds) Methods and principles in medicinal chemistry. VCH, Weinheim
Leo A, Jow PY, Silipo C, Hansch C (1975) Calculation of hydrophobic constant (log P) from pi and f constants. J Med Chem 18:865–868
Cramer RD, Patterson DE, Bunce JD (1988) Comparative molecular field analysis (CoMFA). 1. Effect of shape on binding of steroids to carrier proteins. J Am Chem Soc 110:5959–5967
Beck B, Glen RC, Clark T (1996) The inhibition of alpha-chymotrypsin predicted using theoretically derived molecular properties. J Mol Graph 14(130–5):142
Breindl A, Beck B, Clark T, Glen RC (1997) Prediction of the n-octanol/water partition coefficient, logP, using a combination of semiempirical mo-calculations and a neural network. J Mol Model 3:142–155
TSAR, Oxford Molecular Limited, The Magdalen Centre, Oxford Science Park, Sandford on Thames, Oxford OX4 4GA, UK
van de Waterbeemd H, Gifford E (2003) ADMET in silico modelling: towards prediction paradise? Nat Rev Drug Discov 2:192–204
Kennedy T (1997) Managing the drug discovery/development interface. Drug Discov Today 210:436–444
Lessel U, Wellenzohn B, Lilienthal M, Claussen H (2009) Searching Fragment Spaces with Feature Trees. J Chem Inf Model 49:270–279
Nicolaou CA, Watson I, Wang J (2013) The Lilly Reachable Chemical Space System: bridging chemical synthesis potential with discovery chemistry, sixth jointSheffield Conference on Chemoinformatics
FastROCS v1.4, OpenEye Scientific Software, Inc.: Santa Fe, NM, 2012
Rarey M, Dixon JS (1998) Feature trees: a new molecular similarity measure based on tree matching. J Comput Aided Mol Des 12:471–490
Michel JB, Shen YK, Aiden AP, Veres A, Gray MK, Pickett JP, Hoiberg D, Clancy D, Norvig P, Orwant J, Pinker S, Nowak MA, Aiden EL (2011) Quantitative analysis of culture using millions of digitized books. Science 331:176–182
World Drug Index. Reuters, New York
Comprehensive Medicinal Chemistry, Accelrys Software Inc., San Diego
MDDR, Accelrys Software Inc. San Diego
Bolton EE, Wang Y, Thiessen PA, Bryant SH (2008) Chapter 12 PubChem: integrated platform of small molecules and biological activities. In: Ralph AWaD, (ed) Annual reports in computational chemistry, Elsevier, pp. 217-241.
Gaulton A, Bellis LJ, Bento AP, Chambers J, Davies M, Hersey A, Light Y, McGlinchey S, Michalovich D, Al-Lazikani B, Overington JP (2012) ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic Acids Res 40:D1100–D1107
Chambers J, Davies M, Gaulton A, Hersey A, Velankar S, Petryszak R, Hastings J, Bellis L, McGlinchey S, Overington J (2013) UniChem: a unified chemical structure cross-referencing and identifier tracking system. J Chem Inf 5:3
Integrity, Thomson Reuters, New York
Young D, Martin T, Venkatapathy R, Harten P (2008) Are the chemical structures in your QSAR correct? QSAR Comb Sci 27:1337–1345
Fourches D, Muratov E, Tropsha A (2010) Trust, but verify: on the importance of chemical structure curation in cheminformatics and QSAR modeling research. J Chem Inf Model 50:1189–1204
Olah MM, Bologa CG, Oprea TI (2004) Strategies for compound selection. Curr Drug Discov Technol 1:211–220
Tiikkainen P, Franke L (2011) Analysis of commercial and public bioactivity databases. J Chem Inf Model 52:319–326
Tropsha A (2010) Best practices for QSAR model development, validation, and exploitation. Mol Inf 29:476–488
Cherkasov A, Muratov EN, Fourches D, Varnek A, Baskin II, Cronin M, Dearden J, Gramatica P, Martin YC, Todeschini R, Consonni V, Kuzmin VE, Cramer R, Benigni R, Yang C, Rathman J, Terfloth L, Gasteiger J, Richard A, Tropsha A (2013) QSAR modeling: where have you been? Where are you going to? J Med Chem. doi:10.1021/jm4004285
Todeschini R and Consonni V (2000) Frontmatter. In: Handbook of molecular descriptors. Wiley-VCH, Weinheim, pp i-xxi
Dragon 6, TALETE srl, Via V. Pisani, 13 - 20124 Milano – Italy, 2013
Kramer C & Clark T (2012) New types of descriptors and models in QSAR/QSPR. In: Statistical modelling of molecular descriptors in QSAR/QSPR. Wiley-VCH, Weinheim, pp 293-305
Hopfinger AJ (1980) A QSAR investigation of dihydrofolate reductase inhibition by Baker triazines based upon molecular shape analysis. J Am Chem Soc 102:7196–7206
Hopfinger AJ (1981) Inhibition of dihydrofolate reductase: structure-activity correlations of 2,4-diamino-5-benzylpyrimidines based upon molecular shape analysis. J Med Chem 24:818–822
Hopfinger AJ, Wang S, Tokarski JS, Jin B, Albuquerque M, Madhav PJ, Duraiswami C (1997) Construction of 3D-QSAR models using the 4D-QSAR analysis formalism. J Am Chem Soc 119:10509–10524
Hopfinger AJ, Reaka A, Venkatarangan P, Duca JS, Wang S (1999) Construction of a virtual high throughput screen by 4D-QSAR analysis: application to a combinatorial library of glucose inhibitors of glycogen phosphorylase b. J Chem Inf Comput Sci 39:1151–1160
Kier LB, Hall LH (1976) Molecular connectivity in chemistry and drug research. Academic, New York
Beck B, Horn A, Carpenter JE, Clark T (1998) Enhanced 3D-databases: a fully electrostatic database of AM1-optimized structures. J Chem Inf Comput Sci 38:1214–1217
Beck B, Breindl A, Clark T (2000) QM/NN QSPR models with error estimation: vapor pressure and logP. J Chem Inf Comput Sci 40:1046–1051
Brüstle M, Beck B, Schindler T, King W, Mitchell T, Clark T (2002) Descriptors, physical properties, and drug-likeness. J Med Chem 45:3345–3355
Ehresmann B, de Groot MJ, Clark T (2005) Surface-integral QSPR models: local energy properties. J Chem Inf Model 45:1053–1060
Kramer C, Beck B, Kriegl J, Clark T (2008) A composite model for hERG blockade. Chem Med Chem 3:254–265
Hennemann M, Friedl A, Lobell M, Keldenich J, Hillisch A, Clark T, Göller A (2009) CypScore: quantitative prediction of reactivity toward cytochromes P450 based on semiempirical molecular orbital theory. Chem Med Chem 4:657–669
Kramer C, Beck B, Clark T (2010) A Surface-Integral Model for Log POW. J Chem Inf Model 50:429–436
Shahlaei M (2013) Descriptor selection methods in quantitative structure activity relationship studies: a review study. Chem Rev 113:8093–8103
Breiman L (2001) Random Forests. Mach Learn 5–32
Cortes C, Vapnik V (1995) Support-vector networks. Mach Learn 20:273–297
Rosenblatt F (1958) The perceptron: a probabilistic model for information storage and organization in the brain. Psychol Rev 65:386–408
Legendre AM (1806) Nouvelles méthodes pour la détermination des orbites des comètes: avec un supplément contenant divers perfectionnements perfectionnements. de ces méthodes et leur application aux deux comètes de 1805. Courcier, Paris
Gauss KF (1857) Theory of the motion of the heavenly bodies moving about the sun in conic sections. Dover, Phoenix
Livingstone DJ (2000) The characterization of chemical structures using molecular properties. A survey. J Chem Inf Comput Sci 40:195–209
Tarca AL, Carey VJ, Xw C, Romero R, Drâghici S (2007) Machine learning and its applications to biology. PLoS Comput Biol 3:e116
Rasmussen CE (2004) Gaussian processes in machine learning. Lect Notes Comput Sci 3176:63–71
Kohonen T (1982) Self-organized formation of topologically correct feature maps. Biol Cybern 43:59–69
Schneider P, Stutz K, Kasper L, Haller S, Reutlinger M, Reisen F, Geppert T, Schneider G (2011) Target profile prediction and practical evaluation of a Biginelli-type dihydropyrimidine compound library. Pharmaceuticals 4:1236–1247
Tropsha A, Gramatica P, Gombar V (2003) The importance of being earnest: validation is the absolute essential for successful application and interpretation of QSPR models. QSAR Comb Sci 22:69–77
McNemar Q (1947) Note on the sampling error of the difference between correlated proportions or percentages. Psychometrika 12:153–157
Cumming JG, Davis AM, Muresan S, Haeberlein M, Chen H (2013) Chemical predictive modelling to improve compound quality. Nat Rev Drug Discov 12:948–962
ChemAxon (2013) J Chem 5.11.5 http://www.chemaxon.com
Molecular Operating Environment (MOE), 2013.08; Chemical Computing Group Inc., 1010 Sherbooke St. West, Suite #910, Montreal, QC, Canada, H3A 2R7, 2013
CORINA—Generation of 3D coordinates, version 3.0, Molecular Networks GmbH, Erlangen, Germany
Deng H, Runger G (2013) Gene selection with guided regularized random forest. Pattern Recogn 46:3483–3489
Friedman JH, Hastie T, Tibshirani R (2008) Regularization paths for generalized linear models via coordinate descent. J Stat Softw 33:1–22
Schneider G, Neidhart W, Giller T, Schmid G (1999) Scaffold-hopping by topological pharmacophore search: a contribution to virtual screening. Angew Chem Int Ed 38:2894–2896
Berthold MR, Cebron M, Dill F, Gabriel TR, Kötter T, Meinl T, Ohl P, Sieb C, Thiel K, Wiswedel B (2007) KNIME: the konstanz information miner. In:Studies in classification, data analysis, and knowledge organization. Springer, Heidelberg
Pipeline Pilot, Version 8.0. Accelrys Software Inc. San Diego: s.n.; 2011
R Core Team (2013) R: A Language and Environment for statistical computing. R Foundation for Statistical Computing. (http://www.R-project.org)
Kriegl JM, Arnhold T, Beck B, Fox T (2005) A support vector machine approach to classify human cytochrome P450 3A4 inhibitors. J Comput Aided Mol Des 19:189–201
Veith H, Southall N, Huang R, James T, Fayne D, Artemenko N, Shen M, Inglese J, Austin CP, Lloyd DG, Auld DS (2009) Comprehensive characterization of cytochrome P450 isozyme selectivity across chemical libraries. Nat Biotechnol 27:1050–1055
Cheng F, Yu Y, Shen J, Yang L, Li W, Liu G, Lee PW, Tang Y (2011) Classification of cytochrome P450 inhibitors and noninhibitors using combined classifiers. J Chem Inf Model 51:996–1011
Sun H, Veith H, Xia M, Austin CP, Huang R (2011) Predictive models for cytochrome P450 isozymes based on quantitative high throughput screening data. J Chem Inf Model 51:2474–2481
OECD (2005) Principles for the validation, for regulatory purposes, of (quantitative) structure-activity relationship models
OPS (2000) TOPKAT OPS. US patent no. 6 036 349
Sheridan R, Feuston RP, Maiorov VN, Kearsley S (2004) Similarity to molecules in the training set is a good discriminator for prediction accuracy in QSAR. J Chem Inf Comput Sci 44:1912–1928
Sahigara F, Ballabio D, Todeschini R, Consonni V (2013) Defining a novel k-nearest neighbours approach to assess the applicability domain of a QSAR model for reliable predictions. J Chem Inf 5:27
Netzeva TI, Worth A, Aldenberg T, Benigni R, Cronin MT, Gramatica P, Jaworska JS, Kahn S, Klopman G, Marchant CA, Myatt G, Nikolova-Jeliazkova N, Patlewicz GY, Perkins R, Roberts D, Schultz T, Stanton DW, van de Sandt JJ, Tong W, Veith G, Yang C (2005) Current status of methods for defining the applicability domain of (quantitative) structure-activity relationships. The report and recommendations of ECVAM Workshop 52. Altern Lab Anim 33:155–173
Jaworska J, Nikolova-Jeliazkova N, Aldenberg T (2005) QSAR applicabilty domain estimation by projection of the training set descriptor space: a review. Altern Lab Anim 33:445–459
Wold S, Esbensen K, Geladi P (1987) Principal component analysis. Chemom Intell Lab 2:37–52
Worth AP, Bassan A, Gallegos A, Netzeva TI, Patlewicz G, Pavan M, Tsakovska I, Vra-ìko M (2005) The Characterisation of (quantitative) structure-activity relationships: preliminary guidance. Institute for Health and Consumer Protection, Toxicology and Chemical Substances Unit, European Chemical Bureau
Sahigara F, Mansouri K, Ballabio D, Mauri A, Consonni V, Todeschini R (2012) Comparison of different approaches to define the applicability domain of QSAR models. Molecules 17:4791–4810
Clark RD, Liang W, Waldman M, Fraczkiewicz R (2013) Estimating classification confidence for ensemble models. Sixth Joint Sheffield Conference on Chemoinformatics
Pence HE, Williams A (2010) ChemSpider: an online chemical information resource. J Chem Educ 87:1123–1124
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 46:3–26
Duewer DL (1990) The free–Wilson paradigm redux: Significance of the free–Wilson coefficients, insignificance of coefficient ‘uncertainities’ and statistical sins. J Chemometr 4:299–321
Stardrop, Optibrium Ltd., Cambridge Research, Cambridge, UK
Acknowledgments
We want to thank our Colleagues from Computational Chemistry, Drug Discovery Support and Medicinal Chemistry. Several of the mentioned approaches have been established together with our colleagues from these departments. Without their continuous commitment, assistance and support most of the described research could have not been done.
B. Beck wants to thank Prof. T. Clark for the very good and productive time starting with the Diploma thesis followed by a PhD thesis and with a short intersection a 18 month PostDoc time.
T. Geppert wants to thank Prof. T. Clark for the productive collaborations since his time as a PhD candidate within Prof. G. Schneider’s lab at the ETH Zurich.
Author information
Authors and Affiliations
Corresponding author
Additional information
This paper belongs to a Topical Collection on the occasion of Prof. Tim Clark’s 65th birthday
Rights and permissions
About this article
Cite this article
Beck, B., Geppert, T. Industrial applications of in silico ADMET. J Mol Model 20, 2322 (2014). https://doi.org/10.1007/s00894-014-2322-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-014-2322-5