Abstract
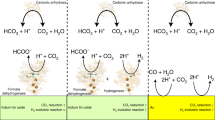
The enzyme carbon monoxide dehydrogenase is capable of efficiently converting \(\hbox {CO}_{2}\) to CO and, therefore, can enable an affordable \(\hbox {CO}_{2}\) recycling strategy. The reduction of \(\hbox {CO}_{2}\) occurs at a peculiar nickel–iron–sulfur cluster, following a mechanism that remains little understood. In this study, we have used ab initio molecular dynamics simulations to explore the free energy landscape of the reaction. We predict the existence of a COOH ligand that strongly interacts with the surrounding protein residues and favours a mechanism where a \(\hbox {H}_{2}\hbox {O}\) molecule is eliminated before CO. We have taken advantages of the insights offered by our simulations to revisit the catalytic mechanism and the role of the residues surrounding the active centre in particular, thus assisting in the design of inorganic catalysts that mimic the enzyme.







Similar content being viewed by others
Change history
24 September 2021
Supplementary file has been included to the article.
References
Wang W, Wang S, Ma X, Gong J (2011) Recent advances in catalytic hydrogenation of carbon dioxide. Chem Soc Rev 40:3703–3727
Appel AM, Bercaw JE, Bocarsly AB, Dobbek H, DuBois DL, Dupuis M, Ferry JG, Fujita E, Hille R, Kenis PJ (2013) Frontiers, opportunities, and challenges in biochemical and chemical catalysis of CO2 fixation. Chem Rev 113:6621–6658
Zheng T, Jiang K, Wang H (2018) Recent advances in electrochemical CO2-to-CO conversion on heterogeneous catalysts. Adv Mater 30:1802066
Can M, Armstrong FA, Ragsdale SW (2014) Structure, function, and mechanism of the nickel metalloenzymes, CO dehydrogenase, and acetyl-CoA synthase. Chem Rev 114:4149–4174
Shin W, Lee SH, Shin JW, Lee SP, Kim Y (2003) Highly selective electrocatalytic conversion of CO2 to CO at -0.57 V (NHE) by carbon monoxide dehydrogenase from Moorella thermoacetica. J Am Chem Soc 125:14688–14689
Parkin A, Seravalli J, Vincent KA, Ragsdale SW, Armstrong FA (2007) Rapid and efficient electrocatalytic CO2/CO interconversions by Carboxydothermus hydrogenoformans CO dehydrogenase I on an electrode. J Am Chem Soc 129:10328–10329
Drennan CL, Heo J, Sintchak MD, Schreiter E, Ludden PW (2001) Life on carbon monoxide: X-ray structure of Rhodospirillum rubrum Ni-Fe-S carbon monoxide dehydrogenase. Proc Natl Acad Sci USA 98:11973–11978
Dobbek H, Svetlitchnyi V, Gremer L, Huber R, Meyer O (2001) Crystal structure of a carbon monoxide dehydrogenase reveals a [Ni-4Fe-5S] cluster. Science 293:1281–1285
Darnault C, Volbeda A, Kim EJ, Legrand P, Vernède X, Lindahl PA, Fontecilla-Camps JC (2003) Ni-Zn-[Fe 4-S 4] and Ni-Ni-[Fe 4-S 4] clusters in closed and open \(\alpha\) subunits of acetyl-CoA synthase/carbon monoxide dehydrogenase. Nat Struct Mol Biol 10:271–279
Dobbek H, Svetlitchnyi V, Liss J, Meyer O (2004) Carbon monoxide induced decomposition of the active site [Ni- 4Fe- 5S] cluster of CO dehydrogenase. J Am Chem Soc 126:5382–5387
Jeoung J-H, Dobbek H (2007) Carbon dioxide activation at the Ni, Fe-cluster of anaerobic carbon monoxide dehydrogenase. Science 318:1461–1464
Jeoung J-H, Dobbek H (2009) Structural basis of cyanide inhibition of Ni, Fe-containing carbon monoxide dehydrogenase. J Am Chem Soc 131:9922–9923
Conover RC, Park JB, Adams MW, Johnson MK (1990) Formation and properties of an iron-nickel sulfide (NiFe3S4) cluster in Pyrococcus furiosus ferredoxin. J Am Chem Soc 112:4562–4564
Lindahl PA (2002) The Ni-containing carbon monoxide dehydrogenase family: light at the end of the tunnel? Biochemistry 41:2097–2105
Rebelein JG, Stiebritz MT, Lee CC, Hu Y (2017) Activation and reduction of carbon dioxide by nitrogenase iron proteins. Nat Chem Biol 13:147–149
Gong W, Hao B, Wei Z, Ferguson DJ, Tallant T, Krzycki JA, Chan MK (2008) Structure of the \(\alpha 2\varepsilon 2\) Ni-dependent CO dehydrogenase component of the Methanosarcina barkeri acetyl-CoA decarbonylase/synthase complex. Proc Natl Acad Sci USA 105:9558–9563
Woolerton TW, Sheard S, Reisner E, Pierce E, Ragsdale SW, Armstrong FA (2010) Efficient and clean photoreduction of CO2 to CO by enzyme-modified TiO2 nanoparticles using visible light. J Am Chem Soc 132:2132–2133
Woolerton TW, Sheard S, Pierce E, Ragsdale SW, Armstrong FA (2011) CO2 photoreduction at enzyme-modified metal oxide nanoparticles. Energy Environ Sci 4:2393–2399
Zhang L, Can M, Ragsdale SW, Armstrong FA (2018) Fast and selective photoreduction of CO2 to CO catalyzed by a complex of carbon monoxide dehydrogenase, TiO2, and Ag nanoclusters. ACS Catal 8:2789–2795
Majumdar A (2014) Bioinorganic modeling chemistry of carbon monoxide dehydrogenases: description of model complexes, current status and possible future scopes. Dalton Trans 43:12135–12145
Le JM, Bren KL (2019) Engineered enzymes and bioinspired catalysts for energy conversion. ACS Energy Lett 4:2168–2180
Volbeda A, Fontecilla-Camps JC (2005) Structural bases for the catalytic mechanism of Ni-containing carbon monoxide dehydrogenases. Dalton Trans 21:3443–3450
Jeon WB, Singer SW, Ludden PW, Rubio LM (2005) New insights into the mechanism of nickel insertion into carbon monoxide dehydrogenase: analysis of Rhodospirillum rubrum carbon monoxide dehydrogenase variants with substituted ligands to the [Fe 3 S 4] portion of the active-site C-cluster. JBIC J Biol Inorg Chem 10:903–912
Seravalli J, Ragsdale SW (2008) 13C NMR characterization of an exchange reaction between CO and CO2 catalyzed by carbon monoxide dehydrogenase. Biochemistry 47:6770–6781
Amara P, Mouesca J-M, Volbeda A, Fontecilla-Camps JC (2011) Carbon monoxide dehydrogenase reaction mechanism: a likely case of abnormal CO2 insertion to a Ni-H bond. Inorg Chem 50:1868–1878
Liao R-Z, Siegbahn PE (2019) Energetics for the mechanism of nickel-containing carbon monoxide dehydrogenase. Inorg Chem 58:7931–7938
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38
Kühne TD, Iannuzzi M, Del Ben M, Rybkin VV, Seewald P, Stein F, Laino T, Khaliullin RZ, Schütt O, Schiffmann F (2020) CP2K: an electronic structure and molecular dynamics software package-Quickstep: efficient and accurate electronic structure calculations. J Chem Phys 152:194103
Perdew JP, Burke K, Ernzerhof M (1996) Generalized gradient approximation made simple. Phys Rev Lett 77:3865
VandeVondele J, Hutter J (2007) Gaussian basis sets for accurate calculations on molecular systems in gas and condensed phases. J Chem Phys 127:114105
Goedecker S, Teter M, Hutter J (1996) Separable dual-space Gaussian pseudopotentials. Phys Rev B 54:1703
Noodleman L (1981) Valence bond description of antiferromagnetic coupling in transition metal dimers. J Chem Phys 74:5737–5743
Spangler NJ, Meyers MR, Gierke KL, Kerby RL, Roberts GP, Ludden PW (1998) Substitution of valine for histidine 265 in carbon monoxide dehydrogenase from Rhodospirillum rubrum affects activity and spectroscopic states. J Biol Chem 273:4059–4064
Byrd RH, Lu P, Nocedal J, Zhu C (1995) A limited memory algorithm for bound constrained optimization. SIAM J Sci Comput 16:1190–1208
Nosé S (1984) A unified formulation of the constant temperature molecular dynamics methods. J Chem Phys 81:511
Himmetoglu B, Floris A, Gironcoli S, Cococcioni M (2014) Hubbard-corrected DFT energy functionals: the LDA+U description of correlated systems. Int J Quantum Chem 114:14–49
Dudarev S, Botton G, Savrasov S, Humphreys C, Sutton A (1998) Electron-energy-loss spectra and the structural stability of nickel oxide: An LSDA+U study. Phys Rev B 57:1505–1509
Nair NN, Ribas-Arino J, Staemmler V, Marx D (2010) Magnetostructural dynamics from Hubbard-U corrected spin-projection:[2Fe- 2S] complex in ferredoxin. J Chem Theory Comput 6:569–575
Terranova U, de Leeuw NH (2014) Aqueous Fe2S2 cluster: structure, magnetic coupling, and hydration behaviour from Hubbard U density functional theory. Phys Chem Chem Phys 16:13426–13433
Laio A, Parrinello M (2002) Escaping free-energy minima. Proc Natl Acad Sci USA 99:12562–12566
Laio A, Rodriguez-Fortea A, Gervasio FL, Ceccarelli M, Parrinello M (2005) Assessing the accuracy of metadynamics. J Phys Chem B 109:6714–6721
Laio A, Gervasio FL (2008) Metadynamics: a method to simulate rare events and reconstruct the free energy in biophysics, chemistry and material science. Rep Progr Phys 71:126601
Farkas B, Terranova U, De Leeuw NH (2020) Binding modes of carboxylic acids on cobalt nanoparticles. Phys Chem Chem Phys 22:985–996
Yoo C, Kim J, Lee Y (2013) Synthesis and reactivity of nickel (II) hydroxycarbonyl species, NiCOOH-\(\kappa\) C. Organometallics 32:7195–7203
Fesseler J, Jeoung J-H, Dobbek H (2015) How the [NiFe4S4] cluster of CO dehydrogenase activates CO2 and NCO-. Angew Chem Int Ed 54:8560–8564
Edgcomb SP, Murphy KP (2002) Variability in the pKa of histidine side-chains correlates with burial within proteins. Proteins Struct Funct Genet 49:1–6
Breglia R, Arrigoni F, Sensi M, Greco C, Fantucci P, De Gioia L, Bruschi M (2021) First-principles calculations on Ni, Fe-containing carbon monoxide dehydrogenases reveal key stereoelectronic features for binding and release of CO2 to/from the C-cluster. Inorg Chem 60:387–402
Siegbahn PE (2021) A quantum chemical approach for the mechanisms of redox-active metalloenzymes. RSC Adv 11:3495–3508
Jeoung J-H, Dobbek H (2012) n-Butyl isocyanide oxidation at the [NiFe 4 S 4 OH x] cluster of CO dehydrogenase. JBIC J Biol Inorg Chem 17:167–173
Stiebritz MT, Hiller CJ, Sickerman NS, Lee CC, Tanifuji K, Ohki Y, Hu Y (2018) Ambient conversion of CO 2 to hydrocarbons by biogenic and synthetic [Fe 4 S 4] clusters. Nat Catal 1:444–451
Ciaccafava A, Tombolelli D, Domnik L, Fesseler J, Jeoung J-H, Dobbek H, Mroginski MA, Zebger I, Hildebrandt P (2016) When the inhibitor tells more than the substrate: the cyanide-bound state of a carbon monoxide dehydrogenase. Chem Sci 7:3162–3171
Ciaccafava A, Tombolelli D, Domnik L, Jeoung J-H, Dobbek H, Mroginski M-A, Zebger I, Hildebrandt P (2017) Carbon monoxide dehydrogenase reduces cyanate to cyanide. Angew Chem Int Ed 56:7398–7401
Wang P-H, Bruschi M, De Gioia L, Blumberger J (2013) Uncovering a dynamically formed substrate access tunnel in carbon monoxide dehydrogenase/acetyl-CoA synthase. J Am Chem Soc 135:9493–9502
Kim EJ, Feng J, Bramlett MR, Lindahl PA (2004) Evidence for a proton transfer network and a required persulfide-bond-forming cysteine residue in Ni-containing carbon monoxide dehydrogenases. Biochemistry 43:5728–5734
Acknowledgements
Via our membership of the UK’s HEC Materials Chemistry Consortium, which is funded by EPSRC (EP/L000202, EP/R029431), this work used the ARCHER UK National Supercomputing Service (http://www.archer.ac.uk). All data supporting this study (the XYZ coordinates of all structures and the metadynamics hills output) are provided as Supplementary Information accompanying this paper.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Terranova, U. Residues surrounding the active centre of carbon monoxide dehydrogenase are key in converting CO2 to CO. J Biol Inorg Chem 26, 617–624 (2021). https://doi.org/10.1007/s00775-021-01878-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-021-01878-4