Abstract
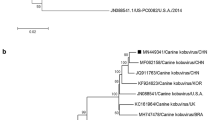
Canine bufavirus (CBuV) is a novel protoparvovirus of dogs that was first reported in 2018 in Italy. The prevalence and genetic diversity of CBuV in China are not clear. In this study, a total of 115 canine fecal samples were collected from northern China in 2019, and two of the samples tested positive for CBuV DNA by PCR. These two field CBuV strains were designated Henan38 and Henan44. The complete genomic sequences of Henan38 and Henan44 were obtained by gap-filling PCR, sequenced, and assembled. Phylogenetic analysis demonstrated that the two strains clustered together in a novel group that was distant from previously reported CBuV strains. This study will strengthen our understanding of the epidemiology and genetic diversity of CBuV in China.

Similar content being viewed by others
References
Martella V, Lanave G, Mihalov-Kovacs E, Marton S, Varga-Kugler R, Kaszab E, Di Martino B, Camero M, Decaro N, Buonavoglia C, Banyai K (2018) Novel parvovirus related to primate Bufaviruses in dogs. Emerg Infect Dis 24:1061–1068
Hargitai R, Pankovics P, Kertész AM, Bíró H, Boros Á, Phan TG, Delwart E, Reuter G (2016) Detection and genetic characterization of a novel parvovirus distantly related to human bufavirus in domestic pigs. Arch Virol 161:1033–1037. https://doi.org/10.1007/s00705-015-2732-4
Yang S, Liu D, Wang Y, Qu F, He Y, Sun Z, Shen Q, Li W, Fu X, Deng X et al (2016) Bufavirus protoparvovirus in feces of wild rats in China. Virus Genes 52:130–133. https://doi.org/10.1007/s11262-015-1262-1
Sasaki M, Orba Y, Anindita PD, Ishii A, Ueno K, Hang'Ombe BM, Mweene AS, Ito K, Sawa H (2015) Distinct lineages of bufavirus in wild shrews and nonhuman primates. Emerg Infect Dis 21:1230–1233
Sun W, Zhang S, Huang H, Wang W, Cao L, Zheng M, Yin Y, Zhang H, Lu H, Jin N (2019) First identification of a novel parvovirus distantly related to human bufavirus from diarrheal dogs in China. Virus Res 265:127–131
Sun Y, Chen Y, Cai Y, Zhu D, Pan H, Wei Y, Han X, Ji C, Lu G, Wang H et al (2020) First report and genetic diversity of porcine bufavirus in China. Virol J. https://doi.org/10.1186/s12985-019-1278-6
Kemenesi G, Dallos B, Gorfol T, Estok P, Boldogh S, Kurucz K, Oldal M, Marton S, Banyai K, Jakab F (2015) Genetic diversity and recombination within bufaviruses: detection of a novel strain in Hungarian bats. Infect Genet Evol 33:288–292
Liu L, Schwarz L, Ullman K, Ahola H, Qiu Y, Ma Z, Hennig-Pauka I (2016) Identification of a novel bufavirus in domestic pigs by a viral metagenomic approach. J Gen Virol 97:1592–1596. https://doi.org/10.1099/jgv.0.000476
Li J, Cui L, Deng X, Yu X, Zhang Z, Yang Z, Delwart E, Zhang W, Hua X (2019) Canine bufavirus in faeces and plasma of dogs with diarrhoea, China. Emerg Microbes Infect 8:245–247. https://doi.org/10.1080/22221751.2018.1563457
Funding
This work was supported by the National Natural Science Foundation of China (31702271) and the Guangdong Provincial Natural Science Foundation (2017A030310367).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they do not have a conflict of interest.
Ethical approval
This research did not involve any human participants or experiments on live animals.
Accession numbers
The complete genome sequences of Henan38 and Henan44 were deposited in the NCBI nucleotide database under the accession numbers MT364251 and MT364252, respectively.
Additional information
Handling Editor: Ana Cristina Bratanich
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shao, R., Zheng, F., Cai, S. et al. Genomic sequencing and characterization of a novel group of canine bufaviruses from Henan province, China. Arch Virol 165, 2699–2702 (2020). https://doi.org/10.1007/s00705-020-04785-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-020-04785-2