Abstract
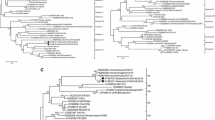
Forty-four Newcastle disease virus (NDV) strains, obtained between 2002 and 2007 from different poultry species in Nigeria, Niger, Burkina Faso and Cameroon, were phylogenetically analysed based on partial F sequences. Lineage 2 viruses were genetically identical or similar to the locally used LaSota vaccine strain and were mostly detected in commercial farms. Lineage 1, 3 and 4 strains were only sporadically found, and their origin was less clear. Twenty-one strains from backyard farms and live bird markets formed three new clusters within lineage 5, tentatively named 5f, 5g and 5h. All of these strains were predicted to be virulent based on their F protein cleavage site sequence. Minimal genetic distances between new and previously established sublineages ranged from 9.4 to 15.9%, and minimal distances between the new sublineages were 11.5 to 17.3%. Their high genetic diversity and their presence in three different Sub-Saharan countries suggest that these new sublineages represent the NDV variants indigenous to West Africa.


Similar content being viewed by others
References
Abolnik C, Horner RF, Bisschop SP, Parker ME, Romito M, Viljoen GJ (2004) A phylogenetic study of South African Newcastle disease virus strains isolated between 1990 and 2002 suggests epidemiological origins in the Far East. Arch Virol 149:603–619
Adu FD, Oyejide O, Ikede BO (1985) Characterization of Nigerian strains of Newcastle disease virus. Avian Dis 29:829–831
Agbede G, Demey F, Verhulst A, Bell JG (1992) Prevalence of Newcastle disease in traditional breeding facilities for chickens in Cameroon. Rev Sci Tech 11:805–811
Aldous EW, Mynn JK, Banks J, Alexander DJ (2003) A molecular epidemiological study of avian paramyxovirus type 1 (Newcastle disease virus) isolates by phylogenetic analysis of a partial nucleotide sequence of the fusion protein gene. Avian Pathol 32:239–256
Aldous EW, Fuller CM, Mynn JK, Alexander DJ (2004) A molecular epidemiological investigation of isolates of the variant avian paramyxovirus type 1 virus (PPMV-1) responsible for the 1978 to present panzootic in pigeons. Avian Pathol 33:258–269
Alexander DJ (2003) Newcastle disease, other avian paramyxoviruses, and pneumovirus infections. In: Saif JM, Barnes HJ, Glisson JR, Fadly AM, McDougald LR, Swayne DE (eds) Diseases of poultry, 11th edn. Ames, Iowa, pp 63–99
Alexander DJ, Manvell RJ, Banks J, Collins MS, Parsons G, Cox B, Frost KM, Speidel EC, Ashman S, Aldous EW (1999) Experimental assessment of the pathogenicity of the Newcastle disease viruses from outbreaks in Great Britain in 1997 for chickens and turkeys, and the protection afforded by vaccination. Avian Pathol 28:501–511
Awan MA, Otte MJ, James AD (1994) The epidemiology of Newcastle disease in rural poultry: a review. Avian Pathol 23:405–423
Ballagi-Pordany A, Wehmann E, Herczeg J, Belak S, Lomniczi B (1996) Identification and grouping of Newcastle disease virus strains by restriction site analysis of a region from the F gene. Arch Virol 141:243–261
Courtecuisse C, Japiot F, Bloch N, Diallo I (1990) Serological survey on Newcastle and Gumboro diseases, pasteurellosis and pullorosis in local hens in Niger. Rev Elev Med Vet Pays Trop 43:27–29
Czegledi A, Wehmann E, Lomniczi B (2003) On the origins and relationships of Newcastle disease virus vaccine strains Hertfordshire and Mukteswar, and virulent strain Herts’33. Avian Pathol 32:271–276
Domingo E, Holland JJ (1997) RNA virus mutations and fitness for survival. Annu Rev Microbiol 51:151–178
Ducatez MF, Olinger CM, Owoade AA, Tarnagda Z, Tahita MC, Sow A, De Landtsheer S, Ammerlaan W, Ouedraogo JB, Osterhaus AD, Fouchier RA, Muller CP (2007) Molecular and antigenic evolution and geographical spread of H5N1 highly pathogenic avian influenza viruses in western Africa. J Gen Virol 88:2297–2306
Echeonwu GO, Iroegbu CU, Emeruwa AC (1993) Recovery of velogenic Newcastle disease virus from dead and healthy free-roaming birds in Nigeria. Avian Pathol 22:383–387
Fuller CM, Collins MS, Easton AJ, Alexander DJ (2007) Partial characterisation of five cloned viruses differing in pathogenicity, obtained from a single isolate of pigeon paramyxovirus type 1 (PPMV-1) following passage in fowls’ eggs. Arch Virol 152:1575–1582
Glickman RL, Syddall RJ, Iorio RM, Sheehan JP, Bratt MA (1988) Quantitative basic residue requirements in the cleavage-activation site of the fusion glycoprotein as a determinant of virulence for Newcastle disease virus. J Virol 62:354–356
Goebel SJ, Taylor J, Barr BC, Kiehn TE, Castro-Malaspina HR, Hedvat CV, Rush-Wilson KA, Kelly CD, Davis SW, Samsonoff WA, Hurst KR, Behr MJ, Masters PS (2007) Isolation of avian paramyxovirus 1 from a patient with a lethal case of pneumonia. J Virol 81:12709–12714
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acid Symp Ser 41:95–98
Herczeg J, Wehmann E, Bragg RR, Travassos Dias PM, Hadjiev G, Werner O, Lomniczi B (1999) Two novel genetic groups (VIIb and VIII) responsible for recent Newcastle disease outbreaks in Southern Africa, one (VIIb) of which reached Southern Europe. Arch Virol 144:2087–2099
Iritani Y, Aoyama S, Takigami S, Hayashi Y, Ogawa R, Yanagida N, Saeki S, Kamogawa K (1991) Antibody response to Newcastle disease virus (NDV) of recombinant fowlpox virus (FPV) expressing a hemagglutinin-neuraminidase of NDV into chickens in the presence of antibody to NDV or FPV. Avian Dis 35:659–661
Kho CL, Mohd-Azmi ML, Arshad SS, Yusoff K (2000) Performance of an RT-nested PCR ELISA for detection of Newcastle disease virus. J Virol Methods 86:71–83
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Liu XF, Wan HQ, Ni XX, Wu YT, Liu WB (2003) Pathotypical and genotypical characterization of strains of Newcastle disease virus isolated from outbreaks in chicken and goose flocks in some regions of China during 1985–2001. Arch Virol 148:1387–1403
Lomniczi B, Wehmann E, Herczeg J, Ballagi-Pordany A, Kaleta EF, Werner O, Meulemans G, Jorgensen PH, Mante AP, Gielkens AL, Capua I, Damoser J (1998) Newcastle disease outbreaks in recent years in Western Europe were caused by an old (VI) and a novel genotype (VII). Arch Virol 143:49–64
Mayo MA (2002) Virus taxonomy—Houston 2002. Arch Virol 147:1071–1076
Meulemans G, van den Berg TP, Decaesstecker M, Boschmans M (2002) Evolution of pigeon Newcastle disease virus strains. Avian Pathol 31:515–519
Oakeley RD (2000) The limitations of a feed/water based heat-stable vaccine delivery system for Newcastle disease-control strategies for backyard poultry flocks in sub-Saharan Africa. Prev Vet Med 47:271–279
Otim MO, Christensen H, Jorgensen PH, Handberg KJ, Bisgaard M (2004) Molecular characterization and phylogenetic study of Newcastle disease virus isolates from recent outbreaks in eastern Uganda. J Clin Microbiol 42:2802–2805
Peeters BP, de Leeuw OS, Koch G, Gielkens AL (1999) Rescue of Newcastle disease virus from cloned cDNA: evidence that cleavability of the fusion protein is a major determinant for virulence. J Virol 73:5001–5009
Qin ZM, Tan LT, Xu HY, Ma BC, Wang YL, Yuan XY, Liu WJ (2008) Pathotypical characterization and molecular epidemiology of Newcastle disease virus isolates from different hosts in China from 1996 to 2005. J Clin Microbiol 46:601–611
Senne DA, King DJ, Kapczynski DR (2004) Control of Newcastle disease by vaccination. Dev Biol (Basel) 119:165–170
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Toyoda T, Sakaguchi T, Imai K, Inocencio NM, Gotoh B, Hamaguchi M, Nagai Y (1987) Structural comparison of the cleavage-activation site of the fusion glycoprotein between virulent and avirulent strains of Newcastle disease virus. Virology 158:242–247
Wang Z, Liu H, Xu J, Bao J, Zheng D, Sun C, Wei R, Song C, Chen J (2006) Genotyping of Newcastle disease viruses isolated from 2002 to 2004 in China. Ann N Y Acad Sci 1081:228–239
Wise MG, Suarez DL, Seal BS, Pedersen JC, Senne DA, King DJ, Kapczynski DR, Spackman E (2004) Development of a real-time reverse-transcription PCR for detection of newcastle disease virus RNA in clinical samples. J Clin Microbiol 42:329–338
Yu L, Wang Z, Jiang Y, Chang L, Kwang J (2001) Characterization of newly emerging Newcastle disease virus isolates from the People’s Republic of China and Taiwan. J Clin Microbiol 39:3512–3519
Yusoff K, Nesbit M, McCartney H, Meulemans G, Alexander DJ, Collins MS, Emmerson PT, Samson AC (1989) Location of neutralizing epitopes on the fusion protein of Newcastle disease virus strain Beaudette C. J Gen Virol 70(Pt 11):3105–3109
Acknowledgments
We thank Aurélie Sausy, Emilie Charpentier and Sebastien De Landtsheer for technical help. The authors gratefully acknowledge support from University of Ibadan Senate research grant SRG/FVM/2006/10A, the cooperation of live-bird poultry market vendors in Oyo and Sokoto states, and of the numerous small subsistence farmers as well as the large commercial farms. We also acknowledge the contribution of Shaiibu Samuel for the sample collection in Sokoto. The authors wish to thank the diagnostic and epidemiological survey service from the LABOCEL in Niger. The authors also thank the Ministry of Cooperation of Luxembourg, the Ministry of Health, the Ministry of Research and the Centre de Recherche Public-Santé for their generous financial and moral support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Nucleotide sequence data reported are available in the DDBJ/EMBL/GenBank databases under the accession numbers FM200796 to FM200839.
Rights and permissions
About this article
Cite this article
Snoeck, C.J., Ducatez, M.F., Owoade, A.A. et al. Newcastle disease virus in West Africa: new virulent strains identified in non-commercial farms. Arch Virol 154, 47–54 (2009). https://doi.org/10.1007/s00705-008-0269-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-008-0269-5