Abstract
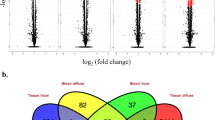
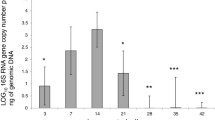
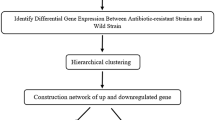
Our aim was to investigate the in vivo gene expression pattern of the Guillain-Barre syndrome (GBS) with DNA microarrays and bioinformatics tools. Oral-infusion model animals mimicking human infection of GBS were analyzed. Tissue samples and body fluids were collected to perform antibody tests and biopsy assays. Gene-expression microarray was conducted with nerve tissues and GBS-related genes were elucidated via bioinformatics tools. Model animals showed typical symptoms of GBS in that mild demyelination was shown by cerebellar white matter and by lumbar enlargement of model animals. Then, 81.25% of the model animals were positive with GM1-IgG antibodies by ELISA. In the microarray analysis, 1,261 genes were identified with statistically different expression (P < 0.05), 21 of which were associated with gene function analysis, gene pathway identification, signal transduction and co-expression network construction. Furthermore, quantitative PCR was used to characterize the gene expression level. We found that genes of HPRT1, PKC and PPARGC-1 were in the core of the network, while the expression of PPARGC-1, SUS2DD and AMPKA2 were significantly inhibited. A total of 21 genes were found to be actively involved in the process of protein transportation, transcriptional regulation, antigen identification and cell cycle regulation during the GBS infection period. The co-expression network indicated an important association between GBS and the 21 genes, especially the down-regulated ones. In conclusion, we demonstrated that GBS-affected hosts had a specific gene expression profile, which may guide the direction of GBS research and therapy.







Similar content being viewed by others
References
Ashbumer M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G (2000) Gene ontology: tool for the unification of biology. The gene ontology consortium. J Nature Genetics 25:25–29
Calderwood MA, Venkatesan K, Xing L, Chase MR, Vazquez A, Holthaus AM, Ewence AE, Li N, Hirozane-Kishikawa T, Hill DE, Vidal M, Kieff E, Johannsen E (2007) Epstein-Barr virus and virus human protein interaction maps. Proc Natl Acad Sci USA 104:7606–7611
Carlson MR, Zhang B, Fang Z, Mischel PS, Horvath S, Nelson SF (2006) Gene connectivity, function, and sequence conservation: predictions from modular yeast co-expression networks. BMC Genomics 7:40
Draghici S, Khatri P, Tarca AL, Amin K, Done A, Voichita C, Georgescu C, Romero R (2007) A systems biology approach for pathway level analysis. Genome Res 17:1537–1545
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32:D277–280
Li C, Li H (2008) Network-constrained regularization and variable selection for analysis of genomic data. Bioinformatics 24:1175–1182
Li CY, Xue P, Tian WQ, Liu RC, Yang C (1996) Experimental campylobacter jejuni infection in the chicken: an animal model of axonal guillain-barre syndrome. J Neurol Neurosurg Psychiatry 61:279–284
Prieto C, Risueno A, Fontanillo C, De las Rivas J (2008) Human gene coexpression landscape: confident network derived from tissue transcriptomic profiles. PLoS One 3:e3911
Ryan MM (2005) Guillain-Barre syndrome in childhood. J Paediatr Child Health 41:237–241
Schmidt-Ott R, Schmidt H, Feldmann S, Brass F, Krone B, Gross U (2006) Improved serological diagnosis stresses the major role of campylobacter jejuni in triggering guillain-barre syndrome. Clin Vaccine Immunol 13:779–783
Tian X, Li C, Zeng X, Li Z, Zhang J (2006) Sequencing of WLAX gene in guillain-barré syndrome-associated campylobacter jejuni strains. Basic Clin Med 26:823–826
Vermeirssen V, Barrasa MI, Hidalgo CA, Babon JA, Sequerra R, Doucette-Stamm L, Barabasi AL, Walhout AJ (2007) Transcription factor modularity in a gene-centered C. elegans core neuronal protein-DNA interaction network. Genome Res 17:1061–1071
Wang X, Shu X, Yang B (2011) Study on the critical antigen that induces immume damage of peripheral nerve following campylobacter jejun infection. J Clin Pediatr 29:68–72
Wright GW, Simon RM (2003) A random variance model for detection of differential gene expression in small microarray experiments. Bioinformatics 19:2448–2455
Yi M, Horton JD, Cohen JC, Hobbs HH, Stephens RM (2006) WholePathwayScope: a comprehensive pathway-based analysis tool for high-throughput data. BMC Bioinforma 7:30
Yuki N (2001) Infectious origins of, and molecular mimicry in, guillain-barre and fisher syndromes. Lancet Infect Dis 1:29–37
Acknowledgements
The author thanks the members of the research group, as well as Prof Brendan W. Wren from the London School of Hygiene & Tropical Medicine for helpful suggestions. Research at the author’s laboratory is supported by the AQSIQ research projects (No. 2009IK135、2010IK128); 12th Five Years Key Programs for Science and Technology Development of China.
Author information
Authors and Affiliations
Corresponding author
Additional information
Feng Xue, Dexin Zeng, Rui Zhang and Fei Xu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Xue, F., Zeng, D., Xu, F. et al. Gene expression profile of campylobacter jejuni-induced GBS in bama miniature pigs. Cell Tissue Res 348, 523–536 (2012). https://doi.org/10.1007/s00441-012-1382-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-012-1382-z