Abstract
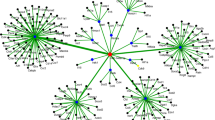
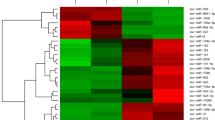
miR-124 is a significantly up-regulated miRNA in peripheral blood collected from piglets infected with Salmonella Typhimurium, suggesting that it may play an important role in Salmonella pathogenesis. This study focused on the transcriptomic analysis of peripheral blood mononuclear cells (PBMCs) isolated from miR-124 sponge and Salmonella Typhimurium-treated piglets, and trying to investigate the function of miR-124 in Salmonella infection. The transcriptome profiling analysis revealed that 2778 genes in miR-124 sponge + Salmonella Typhimurium treatment versus control, 2271 genes in Salmonella Typhimurium treatment versus control, and 1301 genes in miR-124 sponge + Salmonella Typhimurium versus Salmonella Typhimurium treatment, were differentially expressed, respectively (FDR < 0.05 and fold change > 2.0). Pathway analysis indicated that the MAPK signaling pathway, Ribosome pathway, and T-cell receptor signaling pathway were the most significantly enriched pathway in differentially expressed genes between miR-124 sponge + Salmonella Typhimurium and Salmonella Typhimurium along treatment (FDR < 0.05). Reporter assays and electrophoretic mobility shift assays showed that miR-124 is a crucial regulatory factor that targets IQ motif containing GTPase-activating protein 2 (IQGAP2). Cell culture experiment indicated that miR-124 attenuated the Salmonella Typhimurium-mediated activation of CDC42 and RAC1 (P < 0.05). Cultured PBMCs treated with miR-124 and IQGAP2-siRNA had higher intracellular Salmonella count than control samples, particularly 12 h post-infection (P < 0.05). Immunofluorescence analysis revealed that miR-124 treatment reduced the percentage of LAMP-1-positive phagosomes. The miR-124 could be an important regulator for IQGAP2/Rho GTPase pathway in Salmonella Typhimurium-infected PBMCs, and this pathway could be a target for Salmonella that support its infection in PBMCs in piglets.






Similar content being viewed by others
Availability of data and materials
The datasets generated and analyzed during the current study are available in the GEO repository (GSE189380).
References
Aldapa-Vega G, Moreno-Eutimio MA, Berlanga-Taylor AJ, Jimenez-Uribe AP, Nieto-Velazquez G, Lopez-Ortega O, Mancilla-Herrera I, Cortes-Malagon EM, Gunn JS, Isibasi A, Wong-Baeza I, Lopez-Macias C, Pastelin-Palacios R (2019) Structural variants of Salmonella Typhimurium lipopolysaccharide induce less dimerization of TLR4/MD-2 and reduced pro-inflammatory cytokine production in human monocytes. Mol Immunol 111:43–52
Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, Marshall KA, Phillippy KH, Sherman PM, Holko M, Yefanov A, Lee H, Zhang N, Robertson CL, Serova N, Davis S, Soboleva A (2013) NCBI GEO: archive for functional genomics data sets–update. Nucleic Acids Res 41:D991-995
Brill S, Li S, Lyman CW, Church DM, Wasmuth JJ, Weissbach L, Bernards A, Snijders AJ (1996) The Ras GTPase-activating-protein-related human protein IQGAP2 harbors a potential actin binding domain and interacts with calmodulin and Rho family GTPases. Mol Cell Biol 16:4869–4878
da Huang W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57
Guan TY, Holley RA (2003) Pathogen survival in swine manure environments and transmission of human enteric illness–a review. J Environ Qual 32:383–392
Herzog S (2018) Generation of microRNA sponge library. Bio Protoc 8:e2820
Hobbie S, Chen LM, Davis RJ, Galan JE (1997) Involvement of mitogen-activated protein kinase pathways in the nuclear responses and cytokine production induced by Salmonella Typhimurium in cultured intestinal epithelial cells. J Immunol 159:5550–5559
Huang TH, Uthe JJ, Bearson SM, Demirkale CY, Nettleton D, Knetter S, Christian C, Ramer-Tait AE, Wannemuehler MJ, Tuggle CK (2011) Distinct peripheral blood RNA responses to Salmonella in pigs differing in Salmonella shedding levels: intersection of IFNG, TLR and miRNA pathways. PLoS ONE 6:e28768
Huang T, Huang X, Shi B, Wang F, Feng W, Yao M (2018a) Regulators of Salmonella-host interaction identified by peripheral blood transcriptome profiling: roles of TGFB1 and TRP53 in intracellular Salmonella replication in pigs. Vet Res 49:121
Huang T, Huang X, Yao M (2018b) miR-143 inhibits intracellular salmonella growth by targeting ATP6V1A in macrophage cells in pig. Res Vet Sci 117:138–143
Huang T, Huang X, Chen W, Yin J, Shi B, Wang F, Feng W, Yao M (2019) MicroRNA responses associated with Salmonella enterica serovar typhimurium challenge in peripheral blood: effects of miR-146a and IFN-gamma in regulation of fecal bacteria shedding counts in pig. BMC Vet Res 15:195
Huang T, Jiang C, Yang M, Xiao H, Huang X, Wu L, Yao M (2020) Salmonella enterica serovar Typhimurium inhibits the innate immune response and promotes apoptosis in a ribosomal/TRP53-dependent manner in swine neutrophils. Vet Res 51:105
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25
Lathrop SK, Binder KA, Starr T, Cooper KG, Chong A, Carmody AB, Steele-Mortimer O (2015) Replication of Salmonella enterica Serovar Typhimurium in Human Monocyte-Derived Macrophages. Infect Immun 83:2661–2671
LeCour L Jr, Boyapati VK, Liu J, Li Z, Sacks DB, Worthylake DK (2016) The structural basis for Cdc42-induced dimerization of IQGAPs. Structure 24:1499–1508
Lemichez E, Aktories K (2013) Hijacking of Rho GTPases during bacterial infection. Exp Cell Res 319:2329–2336
Luckheeram RV, Zhou R, Verma AD, Xia B (2012) CD4(+)T cells: differentiation and functions. Clin Dev Immunol 2012:925135
Muroi M, Tanamoto K (2002) The polysaccharide portion plays an indispensable role in Salmonella lipopolysaccharide-induced activation of NF-kappaB through human toll-like receptor 4. Infect Immun 70:6043–6047
O’Leary NA, Wright MW, Brister JR, Ciufo S, Haddad D, McVeigh R, Rajput B, Robbertse B, Smith-White B, Ako-Adjei D, Astashyn A, Badretdin A, Bao Y, Blinkova O, Brover V, Chetvernin V, Choi J, Cox E, Ermolaeva O, Farrell CM, Goldfarb T, Gupta T, Haft D, Hatcher E, Hlavina W, Joardar VS, Kodali VK, Li W, Maglott D, Masterson P, McGarvey KM, Murphy MR, O’Neill K, Pujar S, Rangwala SH, Rausch D, Riddick LD, Schoch C, Shkeda A, Storz SS, Sun H, Thibaud-Nissen F, Tolstoy I, Tully RE, Vatsan AR, Wallin C, Webb D, Wu W, Landrum MJ, Kimchi A, Tatusova T, DiCuccio M, Kitts P, Murphy TD, Pruitt KD (2016) Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res 44:D733-745
Ozdemir ES, Jang H, Gursoy A, Keskin O, Li Z, Sacks DB, Nussinov R (2018) Unraveling the molecular mechanism of interactions of the Rho GTPases Cdc42 and Rac1 with the scaffolding protein IQGAP2. J Biol Chem 293:3685–3699
Patel JC, Galan JE (2006) Differential activation and function of Rho GTPases during Salmonella-host cell interactions. J Cell Biol 175:453–463
Pramanik D, Campbell NR, Karikari C, Chivukula R, Kent OA, Mendell JT, Maitra A (2011) Restitution of tumor suppressor microRNAs using a systemic nanovector inhibits pancreatic cancer growth in mice. Mol Cancer Ther 10:1470–1480
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43:e47
Sole A, Mencia N, Villalobos X, Noe V, Ciudad CJ (2013) Validation of miRNA-mRNA interactions by electrophoretic mobility shift assays. BMC Res Notes 6:454
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550
Vinayaka AC et al (2019) "Rapid detection of Salmonella enterica in food samples by a novel approach with combination of sample concentration and direct PCR." Biosens Bioelectron 129:224–230
Wu Z, Zhang Z, Lei Z, Lei P (2019) CD14: biology and role in the pathogenesis of disease. Cytokine Growth Factor Rev 48:24–31
Yao M et al (2016) "Regulation signature of miR-143 and miR-26 in porcine Salmonella infection identified by binding site enrichment analysis." Mol Genet Genomics 291(2):789–799
Zhou M, Li X, Hou W, Wang H, Paoli GC, Shi X (2019) Incidence and characterization of salmonella isolates from raw meat products sold at small markets in Hubei Province. China Front Microbiol 10:2265
Acknowledgements
This project was funded by the National Natural Science Foundation of China [NSFC Grant No. 31902231 and 31402055], the College Students' Innovation and Entrepreneurship Training Program of Yangtze University [Grant No. 2020084], the Teaching research project of Yangtze University [Grant No. JY2020125], and the Graduate Students Teaching Program of Yangtze University [Grant No. YAL202108]. We thank Editage for professional English editing service.
Author information
Authors and Affiliations
Contributions
TH, JY, and MY designed the study, performed the experiments and data analysis, and wrote the manuscript. QT, ZH, HX, CY, and ZL helped conduct the pig infection experiments, and TH performed the RNA extraction. All authors have read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Ethical approval and consent to participate
The animal study was approved by the Animal Care and Use Committee of Hubei Province (China, YZU-2018-0031).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, T., Tian, Q., He, Z. et al. Transcriptome analysis of PBMCs isolated from piglets treated with a miR-124 sponge construct identified miR124/IQGAP2/Rho GTPase as a target pathway support Salmonella Typhimurium infection. Mol Genet Genomics 298, 213–227 (2023). https://doi.org/10.1007/s00438-022-01976-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-022-01976-1