Abstract
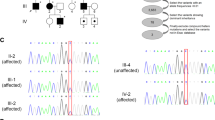
In this study, we aimed to determine the genetic basis of a Turkish family related to hereditary spastic paraplegia (HSP) by exome sequencing. HSP is a progressive neurodegenerative disorder and displays genetic and clinical heterogeneity. The major symptoms are muscle weakness and spasticity, especially in the lower extremities. We studied seven affected and seven unaffected family members, as well as a clinically undetermined member, to identify the disease-causing gene. Exome sequencing was performed for four affected and two unaffected individuals. The variants were firstly filtered for HSP-associated genes, and we found a common variant in the ZFYVE27 gene, which has been previously implied for association with HSP. Due to the incompletely penetrant segregation pattern of the ZFYVE27 variant, revealed by Sanger sequencing, with the disease in this family, filtering was re-performed according to the mode of inheritance and allelic frequencies. The resulting 14 rare variants were further evaluated in terms of their cellular functions, and three candidate variants in ATAD3C, VPS16, and MYO1H genes were selected as possible causative variants, which were analyzed for their familial segregation. ATAD3C and VPS16 variants were eliminated due to incomplete penetrance. Eventually, the MYO1H variant NM_001101421.3:c.2972_2974del (p.Glu992del, rs372231088) was found as the possible disease-causing deletion for HSP in this family. This is the first study reporting the possible role of a MYO1H variant in HSP pathogenesis. Further studies on the cellular roles of Myo1h protein are needed to validate the causality of MYO1H gene at the onset of HSP.




Similar content being viewed by others

References
Adams RJ, Pollard TD (1989) Binding of myosin I to membrane lipids. Nature 340:565–568. https://doi.org/10.1038/340565a0
Adzhubei IA, Schmidt S, Peshkin L et al (2010) A method and server for predicting damaging missense mutations. Nat Methods 7:248–249
Arun RM, Lakkakula BVKS, Chitharanjan AB (2016) Role of myosin 1H gene polymorphisms in mandibular retrognathism. Am J Orthod Dentofac Orthop 149:699–704. https://doi.org/10.1016/j.ajodo.2015.10.028
Bateman A, Martin MJ, Orchard S et al (2021) UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res 49:D480–D489. https://doi.org/10.1093/nar/gkaa1100
Benesh AE, Nambiar R, McConnell RE et al (2010) Differential localization and dynamics of class I myosins in the enterocyte microvillus. Mol Biol Cell 21:970–978. https://doi.org/10.1091/mbc.E09-07-0638
Blackstone C (2012) Cellular pathways of hereditary spastic paraplegia. Annu Rev Neurosci 35:25–47
Blackstone C (2018) Converging cellular themes for the hereditary spastic paraplegias. Curr Opin Neurobiol. https://doi.org/10.1016/j.conb.2018.04.025
Brzeska H, Gonzalez J, Korn ED, Titus MA (2020) Basic-hydrophobic sites are localized in conserved positions inside and outside of PH domains and affect localization of Dictyostelium myosin 1s. Mol Biol Cell 31:101–117. https://doi.org/10.1091/mbc.E19-08-0475
Cheney RE, Mooseker MS (1992) Unconventional myosins. Curr Opin Cell Biol 4:27–35. https://doi.org/10.1016/0955-0674(92)90055-H
Choi Y, Chan AP (2015) PROVEAN web server: a tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 31:2745–2747. https://doi.org/10.1093/bioinformatics/btv195
Coudrier E, Almeida CG (2011) Myosin 1 controls membrane shape by coupling F-Actin to membrane. BioArchitecture 1:230–235. https://doi.org/10.4161/bioa.18406
da Fontoura CSG, Miller SF, Wehby GL et al (2015) Candidate gene analyses of skeletal variation in malocclusion. J Dent Res 94:913–920. https://doi.org/10.1177/0022034515581643
Dai J, Ting-Beall HP, Hochmuth RM et al (1999) Myosin I contributes to the generation of resting cortical tension. Biophys J 77:1168–1176. https://doi.org/10.1016/S0006-3495(99)76968-7
Fink JK (2003) Advances in the hereditary spastic paraplegias. Exp Neurol 184(Suppl 1):106–110. https://doi.org/10.1016/j.expneurol.2003.08.005
Fowler PC, Garcia-Pardo ME, Simpson JC, O’Sullivan NC (2019) NeurodegenERation: the Central Role for ER Contacts in Neuronal Function and Axonopathy, lessons from hereditary spastic paraplegias and related diseases. Front Neurosci 13:1051
Harding AE (1981) Hereditary “pure” spastic paraplegia: a clinical and genetic study of 22 families. J Neurol Neurosurg Psychiatry 44:871–883. https://doi.org/10.1136/jnnp.44.10.871
Harding AE (1983) Classification of the hereditary ataxias and paraplegias. Lancet 321:1151–1155
Hofmann WA, Richards TA, de Lanerolle P (2009) Ancient animal ancestry for nuclear myosin. J Cell Sci 122:636–643. https://doi.org/10.1242/jcs.030205
Iuliano O, Yoshimura A, Prospéri M et al (2018) Myosin 1b promotes axon formation by regulating actin wave propagation and growth cone dynamics. J Cell Biol 217(6):2033–2046
Jia X, Madireddy L, Caillier S et al (2018) Genome sequencing uncovers phenocopies in primary progressive multiple sclerosis. Ann Neurol 84:51–63. https://doi.org/10.1002/ana.25263
Joensuu M, Belevich I, Rämö O et al (2014) ER sheet persistence is coupled to myosin 1c–regulated dynamic actin filament arrays. Mol Biol Cell 25(7):1111–1126
Karczewski KJ, Francioli LC, Tiao G et al (2020) (2020) The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581:7809. https://doi.org/10.1038/s41586-020-2308-7 (581:434–443)
Kars ME, Başak AN, Onat OE et al (2021) The genetic structure of the Turkish population reveals high levels of variation and admixture. Proc Natl Acad Sci USA. https://doi.org/10.1073/PNAS.2026076118/-/dcsupplemental
Kittelberger N, Breunig M, Martin R et al (2016) The role of myosin 1c and myosin 1b in surfactant exocytosis. J Cell Sci 129:1685–1696. https://doi.org/10.1242/jcs.181313
Kumar P, Henikoff S, Ng PC (2009) Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc 4:1073–1082. https://doi.org/10.1038/nprot.2009.86
Lo Giudice T, Lombardi F, Santorelli FM et al (2014) Hereditary spastic paraplegia: clinical-genetic characteristics and evolving molecular mechanisms. Exp Neurol 261:518–539
Lu Q, Li J, Ye F, Zhang M (2015) Structure of myosin-1c tail bound to calmodulin provides insights into calcium-mediated conformational coupling. Nat Struct Mol Biol 22:81–87. https://doi.org/10.1038/nsmb.2923
Mackay-Sim A (2021) Hereditary spastic paraplegia: from genes, cells and networks to novel pathways for drug discovery. Brain Sci. https://doi.org/10.3390/brainsci11030403
Mannan AU, Krawen P, Sauter SM et al (2006) ZFYVE27 (SPG33), a novel spastin-binding protein, is mutated in hereditary spastic paraplegia. Am J Hum Genet 79:351–357. https://doi.org/10.1086/504927
Mazzolini R, Dopeso H, Mateo-Lozano S et al (2012) Brush border Myosin Ia has tumor suppressor activity in the intestine. Proc Natl Acad Sci USA 109:1530–1535. https://doi.org/10.1073/pnas.1108411109
McConnell RE, Tyska MJ (2010) Leveraging the membrane—cytoskeleton interface with myosin-1. Trends Cell Biol 20:418–426
McIntosh BB, Ostap EM (2016) Myosin-I molecular motors at a glance. J Cell Sci 129:2689–2695. https://doi.org/10.1242/jcs.186403
Meyyazhagan A, Orlacchio A (2022) Hereditary Spastic Paraplegia: An Update. Int J Mol Sci 23(3):1697
Murala S, Nagarajan E, Bollu PC (2021) Hereditary spastic paraplegia. Neurol Sci 42(3):883–894
Nambiar R, McConnell RE, Tyska MJ (2009) Control of cell membrane tension by myosin-I. Proc Natl Acad Sci USA 106:11972–11977. https://doi.org/10.1073/pnas.0901641106
Rump A, Scholz T, Thiel C et al (2011) Myosin-1C associates with microtubules and stabilizes the mitotic spindle during cell division. J Cell Sci 124(15):2521–2528
Salinas S, Proukakis C, Crosby A, Warner TT (2008) Hereditary spastic paraplegia: clinical features and pathogenetic mechanisms. The Lancet Neurology 7:1127–1138. https://doi.org/10.1016/S1474-4422(08)70258-8
Schüle R, Bonin M, Dürr A et al (2009) Autosomal dominant spastic paraplegia with peripheral neuropathy maps to chr12q23-24. Neurology 72:1893–1898. https://doi.org/10.1212/wnl.0b013e3181a6086c
Schwarz JM, Rödelsperger C, Schuelke M, Seelow D (2010) MutationTaster evaluates disease-causing potential of sequence alterations. Nat Methods 7:575–576
Spielmann M, Hernandez-Miranda LR, Ceccherini I et al (2017) Mutations in MYO1H cause a recessive form of central hypoventilation with autonomic dysfunction. J Med Genet 54:754–761. https://doi.org/10.1136/jmedgenet-2017-104765
Sun R, Wang Y, Jin M et al (2018) Identification and functional studies of MYO1H for mandibular prognathism. J Dent Res 97:1501–1509. https://doi.org/10.1177/0022034518784936
Tassopoulou-Fishell M, Deeley K, Harvey EM et al (2012) Genetic variation in Myosin 1H contributes to mandibular prognathism. Am J Orthod Dentofac Orthop 141:51–59. https://doi.org/10.1016/j.ajodo.2011.06.033
Tyska MJ, Mackey AT, Huang JD et al (2005) Myosin-1a is critical for normal brush border structure and composition. Mol Biol Cell 16:2443–2457. https://doi.org/10.1091/mbc.e04-12-1116
van der Auwera GA, Carneiro MO, Hartl C et al (2013) From fastQ data to high-confidence variant calls: the genome analysis toolkit best practices pipeline. Curr Protoc Bioinformatics. https://doi.org/10.1002/0471250953.bi1110s43
Wang K, Li M, Hakonarson H (2010) ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. https://doi.org/10.1093/nar/gkq603
Yang Y, Faraggi E, Zhao H, Zhou Y (2011) Improving protein fold recognition and template-based modeling by employing probabilistic-based matching between predicted one-dimensional structural properties of query and corresponding native properties of templates. Bioinformatics 27:2076. https://doi.org/10.1093/bioinformatics/btr350
Acknowledgements
This work was supported by Istanbul Technical University BAP 1239-39586 to AK for ES.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors do not have any conflict of interest.
Ethical approval
Ethical approval was obtained by the Ethics Committee of Istanbul Technical University and Biruni University Hospital.
Informed consent
Informed consent was obtained from all participants included in the study.
Additional information
Communicated by Shuhua Xu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary Information:
Supplementary Table shows the shared variants (except for the variants on ATAD3C, VSP16, and MYO1H) among the cases who were exome sequenced. The elimination criteria used to determine the causative candidate gene are also given in this table (DOCX 193 kb)
Rights and permissions
About this article
Cite this article
Selçuk, E., Kırımtay, K., Temizci, B. et al. MYO1H is a novel candidate gene for autosomal dominant pure hereditary spastic paraplegia. Mol Genet Genomics 297, 1141–1150 (2022). https://doi.org/10.1007/s00438-022-01910-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-022-01910-5