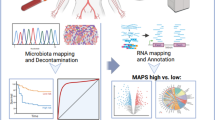
Abstract
Purpose
Given that prognosis of hepatocellular carcinoma (HCC) differs dramatically, it is imperative to uncover effective and available prognostic biomarker(s). The intratumor microbiome plays a significant role in the response to tumor microenvironment, we aimed to identify an intratumor microbiome signature for predicting the prognosis of HCC patients accurately and investigate its possible mechanisms subsequently.
Methods
The TCGA HCC microbiome data (TCGA-LIHC-microbiome) was downloaded from cBioPortal. To create an intratumor microbiome-related prognostic signature, univariate and multivariate Cox regression analyses were used to quantify the association of microbial abundance and patients’ overall survival (OS), as well as their diseases specific survival (DSS). The performance of the scoring model was evaluated by the area under the ROC curve (AUC). Based on the microbiome-related signature, clinical factors, and multi-omics molecular subtypes on the basis of “icluster” algorithm, nomograms were established to predict OS and DSS. Patients were further clustered into three subtypes based on their microbiome-related characteristics by consensus clustering. Moreover, deconvolution algorithm, weighted correlation network analysis (WGCNA) and gene set variation analysis (GSVA) were used to investigate the potential mechanisms.
Results
In TCGA LIHC microbiome data, the abundances of 166 genera among the total 1406 genera were considerably associated with HCC patients’ OS. From that filtered dataset we identified a 27-microbe prognostic signature and developed a microbiome-related score (MRS) model. Compared with those in the relatively low-risk group, patients in higher-risk group own a much worse OS (P < 0.0001). Besides, the time-dependent ROC curves with MRS showed excellent predictive efficacy both in OS and DSS. Moreover, MRS is an independent prognostic factor for OS and DSS over clinical factors and multi-omics-based molecular subtypes. The integration of MRS into nomograms significantly improved the efficacy of prognosis prediction (1-year AUC:0.849, 3-year AUC: 0.825, 5-year AUC: 0.822). The analysis of microbiome-based subtypes on their immune characteristics and specific gene modules inferred that the intratumor microbiome may affect the HCC patients’ prognosis via modulating the cancer stemness and immune response.
Conclusion
MRS, a 27 intratumor microbiome-related prognostic model, was successfully established to predict HCC patients overall survive independently. And the possible underlying mechanisms were also investigated to provide a potential intervention strategy.








Similar content being viewed by others
Data availability
All data used in this study are available on the cBioportal and UCSC Xena website.
Abbreviations
- HCC:
-
Hepatocarcinoma
- OS:
-
Overall survival
- DSS:
-
Disease-specific survival
- WGCNA:
-
Weighted correlation network analysis
- GSVA:
-
Gene set variation analysis
- MRS:
-
Microbiome-related score
- AUC:
-
Area under the ROC curve
- ROC:
-
Receiver-operating characteristic
References
Al-Qadami G, Van Sebille Y, Le H, Bowen J (2019) Gut microbiota: implications for radiotherapy response and radiotherapy-induced mucositis. Expert Rev Gastroenterol Hepatol 13(5):485–496. https://doi.org/10.1080/17474124.2019.1595586
Behary J, Amorim N, Jiang X-T, Raposo A, Gong L, McGovern E, Ibrahim R, Chu F, Stephens C, Jebeili H, Fragomeli V, Koay YC, Jackson M, O’Sullivan J, Weltman M, McCaughan G, El-Omar E, Zekry A (2021) Gut microbiota impact on the peripheral immune response in non-alcoholic fatty liver disease related hepatocellular carcinoma. Nat Commun 12(1):187. https://doi.org/10.1038/s41467-020-20422-7
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, Antipin Y, Reva B, Goldberg AP, Sander C, Schultz N (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2(5):401–404. https://doi.org/10.1158/2159-8290.CD-12-0095
Cheng C, Wang Z, Wang J, Ding C, Sun C, Liu P, Xu X, Liu Y, Chen B, Gu B (2020) Characterization of the lung microbiome and exploration of potential bacterial biomarkers for lung cancer. Transl Lung Cancer Res 9(3):693–704. https://doi.org/10.21037/tlcr-19-590
Chiba A, Bawaneh A, Velazquez C, Clear KYJ, Wilson AS, Howard-McNatt M, Levine EA, Levi-Polyachenko N, Yates-Alston SA, Diggle SP, Soto-Pantoja DR, Cook KL (2020) Neoadjuvant chemotherapy shifts breast tumor microbiota populations to regulate drug responsiveness and the development of metastasis. Mol Cancer Res MCR 18(1):130–139. https://doi.org/10.1158/1541-7786.MCR-19-0451
Dapito DH, Mencin A, Gwak G-Y, Pradere J-P, Jang M-K, Mederacke I, Caviglia JM, Khiabanian H, Adeyemi A, Bataller R, Lefkowitch JH, Bower M, Friedman R, Sartor RB, Rabadan R, Schwabe RF (2012) Promotion of hepatocellular carcinoma by the intestinal microbiota and TLR4. Cancer Cell 21(4):504–516. https://doi.org/10.1016/j.ccr.2012.02.007
Fu A, Yao B, Dong T, Chen Y, Yao J, Liu Y, Li H, Bai H, Liu X, Zhang Y, Wang C, Guo Y, Li N, Cai S (2022) Tumor-resident intracellular microbiota promotes metastatic colonization in breast cancer. Cell 185(8):1356-1372.e26. https://doi.org/10.1016/j.cell.2022.02.027
Gäbele E, Dostert K, Hofmann C, Wiest R, Schölmerich J, Hellerbrand C, Obermeier F (2011) DSS induced colitis increases portal LPS levels and enhances hepatic inflammation and fibrogenesis in experimental NASH. J Hepatol 55(6):1391–1399. https://doi.org/10.1016/j.jhep.2011.02.035
Gao Q, Zhu H, Dong L, Shi W, Chen R, Song Z, Huang C, Li J, Dong X, Zhou Y, Liu Q, Ma L, Wang X, Zhou J, Liu Y, Boja E, Robles AI, Ma W, Wang P, Li Y, Ding L, Wen B, Zhang B, Rodriguez H, Gao D, Zhou H, Fan J (2019) Integrated proteogenomic characterization of HBV-related hepatocellular carcinoma. Cell 179(2):561–577. https://doi.org/10.1016/j.cell.2019.08.052
Guo N, Zhou L-X, Meng N, Shi Y-P (2020) Associations of oral and intestinal florae and serum inflammatory factors with pathogenesis of oral cancer. Eur Rev Med Pharmacol Sci 24(21):11090–11095. https://doi.org/10.26355/eurrev_202011_23595
Hajj Hussein I, Dosh L, Al Qassab M, Jurjus R, El Masri J, Abi Nader C, Rappa F, Leone A, Jurjus A (2023) Highlights on two decades with microbiota and inflammatory bowel disease from etiology to therapy. Transplant Immunol 78:101835. https://doi.org/10.1016/j.trim.2023.101835
Helmink BA, Khan MAW, Hermann A, Gopalakrishnan V, Wargo JA (2019) The microbiome, cancer, and cancer therapy. Nat Med 25(3):377–388. https://doi.org/10.1038/s41591-019-0377-7
Hu X, Chen R, Wei Q, Xu X (2022) The landscape of alpha fetoprotein in hepatocellular carcinoma: where are we? Int J Biol Sci 18(2):536–551. https://doi.org/10.7150/ijbs.64537
Huang H, Ren Z, Gao X, Hu X, Zhou Y, Jiang J, Lu H, Yin S, Ji J, Zhou L, Zheng S (2020) Integrated analysis of microbiome and host transcriptome reveals correlations between gut microbiota and clinical outcomes in HBV-related hepatocellular carcinoma. Genome Med 12(1):102. https://doi.org/10.1186/s13073-020-00796-5
Huang Y, Fan X-G, Wang Z-M, Zhou J-H, Tian X-F, Li N (2004) Identification of helicobacter species in human liver samples from patients with primary hepatocellular carcinoma. J Clin Pathol 57(12):1273–1277. https://doi.org/10.1136/jcp.2004.018556
Jiang P, Gu S, Pan D, Fu J, Sahu A, Hu X, Li Z, Traugh N, Bu X, Li B, Liu J, Freeman GJ, Brown MA, Wucherpfennig KW, Liu XS (2018) Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med 24(10):1550–1558. https://doi.org/10.1038/s41591-018-0136-1
Johnson P, Zhou Q, Dao DY, Lo YMD (2022) Circulating biomarkers in the diagnosis and management of hepatocellular carcinoma. Nat Rev Gastroenterol Hepatol 19(10):670–681. https://doi.org/10.1038/s41575-022-00620-y
Kang Y, Cai Y, Yang Y (2022) The gut microbiome and hepatocellular carcinoma: implications for early diagnostic biomarkers and novel therapies. Liver Cancer 11(2):113–125. https://doi.org/10.1159/000521358
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinform 9:559. https://doi.org/10.1186/1471-2105-9-559
Li Q, Cao M, Lei L, Yang F, Li H, Yan X, He S, Zhang S, Teng Y, Xia C, Chen W (2022) Burden of liver cancer: from epidemiology to prevention. Chin J Cancer Res Chung-Kuo Yen Cheng Yen Chiu 34(6):554–566. https://doi.org/10.21147/j.issn.1000-9604.2022.06.02
Li R, Zhou R, Wang H, Li W, Pan M, Yao X, Zhan W, Yang S, Xu L, Ding Y, Zhao L (2019) Gut microbiota-stimulated cathepsin K secretion mediates TLR4-dependent M2 macrophage polarization and promotes tumor metastasis in colorectal cancer. Cell Death Differ 26(11):2447–2463. https://doi.org/10.1038/s41418-019-0312-y
Lin Z-F, Qin L-X, Chen J-H (2022) Biomarkers for response to immunotherapy in hepatobiliary malignancies. Hepatobiliary Pancreat Dis Int HBPD INT 21(5):413–419. https://doi.org/10.1016/j.hbpd.2022.08.002
Luo X-Y, Wu K-M, He X-X (2021) Advances in drug development for hepatocellular carcinoma: clinical trials and potential therapeutic targets. J Exp Clin Cancer Res CR 40(1):172. https://doi.org/10.1186/s13046-021-01968-w
Ma C, Han M, Heinrich B, Fu Q, Zhang Q, Sandhu M, Agdashian D, Terabe M, Berzofsky JA, Fako V, Ritz T, Longerich T, Theriot CM, McCulloch JA, Roy S, Yuan W, Thovarai V, Sen SK, Ruchirawat M, Korangy F, Wang XW, Trinchieri G, Greten TF (2018) Gut microbiome-mediated bile acid metabolism regulates liver cancer via NKT cells. Science (new York, NY) 360(6391):eaan5931. https://doi.org/10.1126/science.aan5931
Mo Q, Shen R, Guo C, Vannucci M, Chan KS, Hilsenbeck SG (2018) A fully Bayesian latent variable model for integrative clustering analysis of multi-type omics data. Biostatistics (oxford, England) 19(1):71–86. https://doi.org/10.1093/biostatistics/kxx017
Nejman D, Livyatan I, Fuks G, Gavert N, Zwang Y, Geller LT, Rotter-Maskowitz A, Weiser R, Mallel G, Gigi E, Meltser A, Douglas GM, Kamer I, Gopalakrishnan V, Dadosh T, Levin-Zaidman S, Avnet S, Atlan T, Cooper ZA et al (2020) The human tumor microbiome is composed of tumor type-specific intracellular bacteria. Science (new York, NY) 368(6494):973–980. https://doi.org/10.1126/science.aay9189
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, Hoang CD, Diehn M, Alizadeh AA (2015) Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 12(5):453–457. https://doi.org/10.1038/nmeth.3337
Nikitina D, Lehr K, Vilchez-Vargas R, Jonaitis LV, Urba M, Kupcinskas J, Skieceviciene J, Link A (2023) Comparison of genomic and transcriptional microbiome analysis in gastric cancer patients and healthy individuals. World J Gastroenterol 29(7):1202–1218. https://doi.org/10.3748/wjg.v29.i7.1202
Poore GD, Kopylova E, Zhu Q, Carpenter C, Fraraccio S, Wandro S, Kosciolek T, Janssen S, Metcalf J, Song SJ, Kanbar J, Miller-Montgomery S, Heaton R, Mckay R, Patel SP, Swafford AD, Knight R (2020) Microbiome analyses of blood and tissues suggest cancer diagnostic approach. Nature 579(7800):567–574. https://doi.org/10.1038/s41586-020-2095-1
Qu D, Wang Y, Xia Q, Chang J, Jiang X, Zhang H (2022) Intratumoral microbiome of human primary liver cancer. Hepatol Commun 6(7):1741–1752. https://doi.org/10.1002/hep4.1908
Rao B-C, Lou J-M, Wang W-J, Li A, Cui G-Y, Yu Z-J, Ren Z-G (2020) Human microbiome is a diagnostic biomarker in hepatocellular carcinoma. Hepatobiliary Pancreat Dis Int HBPD INT 19(2):109–115. https://doi.org/10.1016/j.hbpd.2020.01.003
Ren Z, Li A, Jiang J, Zhou L, Yu Z, Lu H, Xie H, Chen X, Shao L, Zhang R, Xu S, Zhang H, Cui G, Chen X, Sun R, Wen H, Lerut JP, Kan Q, Li L, Zheng S (2019) Gut microbiome analysis as a tool towards targeted non-invasive biomarkers for early hepatocellular carcinoma. Gut 68(6):1014–1023. https://doi.org/10.1136/gutjnl-2017-315084
Routy B, Le Chatelier E, Derosa L, Duong CPM, Alou MT, Daillère R, Fluckiger A, Messaoudene M, Rauber C, Roberti MP, Fidelle M, Flament C, Poirier-Colame V, Opolon P, Klein C, Iribarren K, Mondragón L, Jacquelot N, Qu B et al (2018) Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science (new York, NY) 359(6371):91–97. https://doi.org/10.1126/science.aan3706
Shen M, Di K, He H, Xia Y, Xie H, Huang R, Liu C, Yang M, Zheng S, He N, Li Z (2020) Progress in exosome associated tumor markers and their detection methods. Mol Biomed 1(1):3. https://doi.org/10.1186/s43556-020-00002-3
Sivan A, Corrales L, Hubert N, Williams JB, Aquino-Michaels K, Earley ZM, Benyamin FW, Lei YM, Jabri B, Alegre M-L, Chang EB, Gajewski TF (2015) Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science (new York, NY) 350(6264):1084–1089. https://doi.org/10.1126/science.aac4255
Vétizou M, Pitt JM, Daillère R, Lepage P, Waldschmitt N, Flament C, Rusakiewicz S, Routy B, Roberti MP, Duong CPM, Poirier-Colame V, Roux A, Becharef S, Formenti S, Golden E, Cording S, Eberl G, Schlitzer A, Ginhoux F et al (2015) Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science (new York, NY) 350(6264):1079–1084. https://doi.org/10.1126/science.aad1329
Wei X, Su R, Yang M, Pan B, Lu J, Lin H, Shu W, Wang R, Xu X (2022) Quantitative proteomic profiling of hepatocellular carcinoma at different serum alpha-fetoprotein level. Transl Oncol 20:101422–101431. https://doi.org/10.1016/j.tranon.2022.101422
Wilkerson MD, Hayes DN (2010) ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics (oxford, England) 26(12):1572–1573. https://doi.org/10.1093/bioinformatics/btq170
Xia C, Dong X, Li H, Cao M, Sun D, He S, Yang F, Yan X, Zhang S, Li N, Chen W (2022) Cancer statistics in China and United States, 2022: profiles, trends, and determinants. Chin Med J 135(5):584–590. https://doi.org/10.1097/CM9.0000000000002108
Xia C, Su J, Liu C, Mai Z, Yin S, Yang C, Fu L (2023) Human microbiomes in cancer development and therapy. MedComm 4(2):e221. https://doi.org/10.1002/mco2.221
Xiang Z, Wu J, Li J, Zheng S, Wei X, Xu X (2023) Gut microbiota modulation: a viable strategy to address medical needs in hepatocellular carcinoma and liver transplantation. Engineering. https://doi.org/10.1016/j.eng.2022.12.012
Xue C, Chu Q, Zheng Q, Yuan X, Su Y, Bao Z, Lu J, Li L (2023) Current understanding of the intratumoral microbiome in various tumors. Cell Rep Med 4(1):100884. https://doi.org/10.1016/j.xcrm.2022.100884
Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W, Treviño V, Shen H, Laird PW, Levine DA, Carter SL, Getz G, Stemke-Hale K, Mills GB, Verhaak RGW (2013) Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun 4:2612. https://doi.org/10.1038/ncomms3612
Yu G, Wang L-G, Han Y, He Q-Y (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS J Integr Biol 16(5):284–287. https://doi.org/10.1089/omi.2011.0118
Zheng H, Liu H, Li H, Dou W, Wang J, Zhang J, Liu T, Wu Y, Liu Y, Wang X (2022) Characterization of stem cell landscape and identification of stemness-relevant prognostic gene signature to aid immunotherapy in colorectal cancer. Stem Cell Res Ther 13(1):244. https://doi.org/10.1186/s13287-022-02913-0
Funding
This work was supported by the Major Research Plan of the National Natural Science Foundation of China (No. 92159202); National Key Research and Development Program of China (Nos. 2021YFA1100502, 2021YFA1100504).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study’s conception and design. Material preparation, data collection and analysis were performed by YS. The first draft of the manuscript was written by YS and ZX, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Figure S1
Kaplan–Meier OS for AFP positive or negative in HCC (PDF 7 KB)
Figure S2
Further stratification of HCC with different AFP levels by MRS. A, C The effect of MRS on OS and DSS in HCC with AFP > 400 ng/ml. B, D The effect of MRS on OS and DSS in HCC with AFP < 400 ng/ml (PDF 578 KB)
Figure S3
Differences of MRS among cluster C1, C2 and C3 (PDF 86 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Song, Y., Xiang, Z., Lu, Z. et al. Identification of a brand intratumor microbiome signature for predicting prognosis of hepatocellular carcinoma. J Cancer Res Clin Oncol 149, 11319–11332 (2023). https://doi.org/10.1007/s00432-023-04962-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-04962-1