Abstract
Background
Ewing's sarcoma (ES) is a kind of malignant tumor, which often occurs in the long bone, pelvis, and other bone tissues, as well as some soft tissues. It often occurs in children and adolescents, second only to osteosarcoma and rhabdomyosarcoma. In the past 30 years, little progress has been made on the genomic mechanism of ES metastasis.
Methods
The gene expression sequence of ES metastasis samples was compared with that of primary tumor samples to obtain differentially expressed genes (DEGs). Subsequently, we annotated the gene functions and enriched pathways of DEGs. Additionally, the protein and protein interaction network were constructed to screen key genes that can lead to the metastasis in ES. Then, cell and molecular biology experiments were conducted to verify the results obtained from the bioinformatics analysis. Finally, we assessed the correlation of expression between the key genes EWSR and FLI1, and conducted a survival analysis of ICAM1.
Results
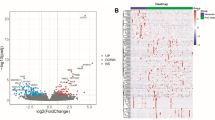
Our study revealed 153 DEGs. Of these, 82 (53.59%) were upregulated and the remaining 71 (46.41%) were downregulated. The bioinformatics analysis showed that ICAM1 was the key gene leading to the invasion and metastasis of ES. Through cell biology and molecular biology experiments, inactivation of ICAM1 inhibited the metastasis of ES cells. The survival and correlation analyses showed that ICAM1 was a risk factor in patients with ES, and that ICAM1 expression was correlated with EWSR and FLI1 expression.
Conclusion
Our study shows that inactivation of ICAM1 inhibits metastasis and improves the prognosis of ES. Additionally, our findings provide a better understanding of the underlying mechanisms of metastatic ES, a basis for an accurate diagnosis, and therapeutic targets for ES patients.






Similar content being viewed by others
Change history
06 January 2022
A Correction to this paper has been published: https://doi.org/10.1007/s00432-021-03869-z
References
Antal CE et al (2015) Cancer-associated protein kinase C mutations reveal kinase’s role as tumor suppressor. Cell 160:489–502. https://doi.org/10.1016/j.cell.2015.01.001
Bonin G, Scamps C, Turc-Carel C, Lipinski M (1993) Chimeric EWS-FL11 transcript in a ewing cell line with a complex t(11; 22; 14). Translocat Cancer Res 53:3655–3657
Broggini T et al (2015) ICAM1 depletion reduces spinal metastasis formation in vivo and improves neurological outcome. Eur Spine J 24:2173–2181. https://doi.org/10.1007/s00586-015-3811-7
Cao H, Xu E, Liu H, Wan L, Lai M (2015) Epithelial–mesenchymal transition in colorectal cancer metastasis: a system review. Pathol Res Pract 211:557–569
Castagna M, Takai Y, Kaibuchi K, Sano K, Kikkawa U, Nishizuka Y (1982) Direct activation of calcium-activated, phospholipid-dependent protein kinase by tumor-promoting phorbol esters. J Biol Chem 257:7847–7851
Chaturvedi A, Hoffman LM, Welm AL, Lessnick SL, Beckerle MC (2012) The EWS/FLI oncogene drives changes in cellular morphology, adhesion, and migration in Ewing sarcoma. Genes Cancer 3:102–116
Chin C-H, Chen S-H, Wu H-H, Ho C-W, Ko M-T, Lin C-Y (2014) cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol 8:S11
Choi E-YK, Gardner JM, Lucas DR, McHugh JB, Patel RM (2014) Ewing sarcoma. Seminars in diagnostic pathology, vol 1. Elsevier, Amsterdam, pp 39–47
De Alava E et al (1998) EWS–FLI1 fusion transcript structure is an independent determinant of prognosis in Ewing’s sarcoma. J Clin Oncol 16:1248–1255
Delattre O et al (1992) Gene fusion with an ETS DNA-binding domain caused by chromosome translocation in human tumours. Nature 359:162
Diboun I, Wernisch L, Orengo CA, Koltzenburg M (2006) Microarray analysis after RNA amplification can detect pronounced differences in gene expression using limma. BMC Genomics 7:252
Dries DR, Gallegos LL, Newton AC (2007) A single residue in the C1 domain sensitizes novel protein kinase C isoforms to cellular diacylglycerol production. J Biol Chem 282:826–830. https://doi.org/10.1074/jbc.C600268200
Gaspar N et al (2015) Ewing sarcoma: current management and future approaches through collaboration. J Clin Oncol 33:3036–3046
Gilkes DM, Semenza GL, Wirtz D (2014) Hypoxia and the extracellular matrix: drivers of tumour metastasis. Nat Rev Cancer 14:430–439. https://doi.org/10.1038/nrc3726
Gorlick R, Janeway K, Lessnick S, Randall RL, Marina N, Committee COGBT (2013) Children’s Oncology Group’s 2013 blueprint for research: bone tumors. Pediatr Blood Cancer 60:1009–1015. https://doi.org/10.1002/pbc.24429
Grier HE et al (2003) Addition of ifosfamide and etoposide to standard chemotherapy for Ewing’s sarcoma and primitive neuroectodermal tumor of bone. N Engl J Med 348:694–701
Howarth MM et al (2014) Long noncoding RNA EWSAT1-mediated gene repression facilitates Ewing sarcoma oncogenesis. J Clin Investig 124:5275–5290
Huang X et al (2016) High-density lipoprotein of patients with breast cancer complicated with type 2 diabetes mellitus promotes cancer cells adhesion to vascular endothelium via ICAM-1 and VCAM-1 upregulation. Breast Cancer Res Treat 155:441–455. https://doi.org/10.1007/s10549-016-3696-0
Hubbard AK, Rothlein R (2000) Intercellular adhesion molecule-1 (ICAM-1) expression and cell signaling cascades. Free Radic Biol Med 28:1379–1386. https://doi.org/10.1016/s0891-5849(00)00223-9
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4:249–264
Lawlor ER, Sorensen PH (2015) Twenty years on: what do we really know about Ewing sarcoma and what is the path forward? Crit Rev Oncog 20:155–171
Leek J, Johnson W, Parker H, Jaffe A, Storey J (2014) sva: surrogate variable analysis. R package version 3.10.0
Liu S et al (2013) Expression of intercellular adhesion molecule 1 by hepatocellular carcinoma stem cells and circulating tumor cells. Gastroenterology 144(1031–1041):e1010. https://doi.org/10.1053/j.gastro.2013.01.046
Lung HL et al (2005) THY1 is a candidate tumour suppressor gene with decreased expression in metastatic nasopharyngeal carcinoma. Oncogene 24:6525–6532. https://doi.org/10.1038/sj.onc.1208812
Luo L et al (2018) miR-335-5p targeting ICAM-1 inhibits invasion and metastasis of thyroid cancer cells. Biomed Pharmacother 106:983–990. https://doi.org/10.1016/j.biopha.2018.07.046
Meyers PA (2015) Systemic therapy for osteosarcoma and Ewing sarcoma. American Society of Clinical Oncology educational book. In: American Society of Clinical Oncology annual meeting, pp e644–647. https://doi.org/10.14694/EdBook_AM.2015.35.e644
Niedan S et al (2014) Suppression of FOXO1 is responsible for a growth regulatory repressive transcriptional sub-signature of EWS–FLI1 in Ewing sarcoma. Oncogene 33:3927–3938
Okegawa T, Pong RC, Li Y, Hsieh JT (2004) The role of cell adhesion molecule in cancer progression and its application in cancer therapy. Acta Biochim Pol 51:445–457. https://doi.org/10.18388/abp.2004_3583
Ozaki T (2015) Diagnosis and treatment of Ewing sarcoma of the bone: a review article. J Orthop Sci 20:250–263
Rosette C, Roth RB, Oeth P, Braun A, Kammerer S, Ekblom J, Denissenko MF (2005) Role of ICAM1 in invasion of human breast cancer cells. Carcinogenesis 26:943–950. https://doi.org/10.1093/carcin/bgi070
Schroder C et al (2011) Prognostic value of intercellular adhesion molecule (ICAM)-1 expression in breast cancer. J Cancer Res Clin Oncol 137:1193–1201. https://doi.org/10.1007/s00432-011-0984-2
Selvanathan SP et al (2019) EWS–FLI1 modulated alternative splicing of ARID1A reveals novel oncogenic function through the BAF complex. Nucl Acids Res 47:9619–9636
Therneau T, Lumley T (2013) R survival package
Wei T, Simko V, Levy M, Xie Y, Jin Y, Zemla J (2017) Package ‘corrplot.’ Statistician 56:316–324
Yan C et al (2018) Identification of key genes and pathways in Ewing’s sarcoma using bioinformatics analysis. J BU ON 23:1472–1480
Yang L, Hu H-M, Zielinska-Kwiatkowska A, Chansky HA (2010) FOXO1 is a direct target of EWS–FLI1 oncogenic fusion protein in Ewing’s sarcoma cells. Biochem Biophys Res Commun 402:129–134
Yu G, Wang L-G, Han Y, He Q-Y (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. Omics J Integr Biol 16:284–287
Zeng LQ, Peng ZL, Duan ZL (2009) Expression of THY1 gene in epithelial ovarian cancer. Chin J Oncol 31:118–120
Acknowledgements
The present study was supported by the Youth Program of National Natural Science Foundation of China (81801213), China Postdoctoral Science Foundation, No. 65 General Fund (2019M651967), Jiangsu Planned Projects for Postdoctoral Research Funds (2018K176C) and China Postdoctoral start-up fund (2018107007), Science and technology project of yili kazak autonomous prefecture (YZ2019D006). The authors would also like to acknowledge the family and colleagues for giving up their time and energy to support this study.
Funding
The Youth Program of National Natural Science Foundation of China (81801213); China Postdoctoral Science Foundation, No. 65 General Fund (2019M651967); Jiangsu Planned Projects for Postdoctoral Research Funds (2018K176C); China Postdoctoral start-up fund (2018107007); Science and technology project of yili kazak autonomous prefecture (YZ2019D006).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised due to correction in figure 5.
Rights and permissions
About this article
Cite this article
Pan, B., Bu, X., Cao, M. et al. Inactivation of ICAM1 inhibits metastasis and improves the prognosis of Ewing's sarcoma. J Cancer Res Clin Oncol 147, 393–401 (2021). https://doi.org/10.1007/s00432-020-03431-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-020-03431-3