Abstract
Main conclusion
AS events affect genes encoding protein domain composition and make the single gene produce more proteins with a certain number of genes to satisfy the establishment of photosynthesis during de-etiolation.
Abstract
The drastic switch from skotomorphogenic to photomorphogenic development is an excellent system to elucidate rapid developmental responses to environmental stimuli in plants. To decipher the effects of different light wavelengths on de-etiolation, we illuminated etiolated maize seedlings with blue, red, blue–red mixed and white light, respectively. We found that blue light alone has the strongest effect on photomorphogenesis and that this effect can be attributed to the higher number and expression levels of photosynthesis and chlorosynthesis proteins. Deep sequencing-based transcriptome analysis revealed gene expression changes under different light treatments and a genome-wide alteration in alternative splicing (AS) profiles. We discovered 41,188 novel transcript isoforms for annotated genes, which increases the percentage of multi-exon genes with AS to 63% in maize. We provide peptide support for all defined types of AS, especially retained introns. Further in silico prediction revealed that 58.2% of retained introns have changes in domains compared with their most similar annotated protein isoform. This suggests that AS acts as a protein function switch allowing rapid light response through the addition or removal of functional domains. The richness of novel transcripts and protein isoforms also demonstrates the potential and importance of integrating proteomics into genome annotation in maize.






Similar content being viewed by others
Abbreviations
- AS:
-
Alternative splicing
- BP:
-
Biological process
- CC:
-
Cellular component
- DEG:
-
Differentially expressed protein
- DEU:
-
Differential exon usage
- GO:
-
Gene ontology
- MF:
-
Molecular function
- NTR:
-
Novel transcript region
- PTC:
-
Premature termination codon
References
Anders S, Reyes A, Huber W (2012) Detecting differential usage of exons from RNA-seq data. Genome Res 22:2008–2017
Au KF, Jiang H, Lin L, Xing Y, Wong WH (2010) Detection of splice junctions from paired-end RNA-seq data by SpliceMap. Nucleic Acids Res 38:4570–4578
Bourbousse C, Mestiri I, Zabulon G, Bourge M, Formiggini F, Koini MA, Brown SC, Fransz P, Bowler C, Barneche F (2015) Light signaling controls nuclear architecture reorganization during seedling establishment. Proc Natl Acad Sci USA 112:E2836–2844
Brett D, Pospisil H, Valcarcel J, Reich J, Bork P (2002) Alternative splicing and genome complexity. Nat Genet 30:29–30
Brooks AN, Duff MO, May G, Yang L, Bolisetty M, Landolin J, Wan K, Sandler J, Booth BW, Celniker SE, Graveley BR, Brenner SE (2015) Regulation of alternative splicing in Drosophila by 56 RNA binding proteins. Genome Res 25:1771–1780
Bu TT, Shen J, Chao Q, Shen Z, Yan Z, Zheng HY, Wang BC (2017) Dynamic N-glycoproteome analysis of maize seedling leaves during de-etiolation using Concanavalin A lectin affinity chromatography and a nano-LC–MS/MS-based iTRAQ approach. Plant Cell Rep 36:1943–2195
Buza TJ, McCarthy FM, Burgess SC (2007) Experimental-confirmation and functional-annotation of predicted proteins in the chicken genome. BMC Genomics 8:425
Campbell MA, Haas BJ, Hamilton JP, Mount SM, Buell CR (2006) Comprehensive analysis of alternative splicing in rice and comparative analyses with Arabidopsis. BMC Genomics 7:327
Castillon A, Shen H, Huq E (2009) Blue light induces degradation of the negative regulator phytochrome interacting factor 1 to promote photomorphogenic development of Arabidopsis seedlings. Genetics 182:161–171
Chory J (1992) A genetic model for light-regulated seedling development in Arabidopsis. Development 115:337–354
Covshoff S, Majeran W, Liu P, Kolkman JM, van Wijk KJ, Brutnell TP (2008) Deregulation of maize C4 photosynthetic development in a mesophyll cell-defective mutant. Plant Physiol 146:1469–1481
Craig R, Beavis RC (2004) TANDEM: matching proteins with tandem mass spectra. Bioinform 20:1466–1467
de Souza GA, Malen H, Softeland T, Saelensminde G, Prasad S, Jonassen I, Wiker HG (2008) High accuracy mass spectrometry analysis as a tool to verify and improve gene annotation using Mycobacterium tuberculosis as an example. BMC Genomics 9:316
Fellner M, Horton LA, Cocke AE, Stephens NR, Ford ED, Van Volkenburgh E (2003) Light interacts with auxin during leaf elongation and leaf angle development in young corn seedlings. Planta 216:366–376
Finn RD, Clements J, Eddy SR (2011) HMMER web server: interactive sequence similarity searching. Nucleic Acids Res 39:W29–W37
Gan CS, Chong PK, Pham TK, Wright PC (2007) Technical, experimental, and biological variations in isobaric tags for relative and absolute quantitation (iTRAQ). J Proteome Res 6:821–825
Guan Q, Zheng W, Tang S, Liu X, Zinkel RA, Tsui KW, Yandell BS, Culbertson MR (2006) Impact of nonsense-mediated mRNA decay on the global expression profile of budding yeast. PLoS Genet 2:e203
Hansey CN, Vaillancourt B, Sekhon RS, de Leon N, Kaeppler SM, Buell CR (2012) Maize (Zea mays L.) genome diversity as revealed by RNA-sequencing. PLoS ONE 7:e33071
Hartmann L, Drewe-Boss P, Wiessner T, Wagner G, Geue S, Lee HC, Obermüller DM, Kahles A, Behr J, Sinz FH, Rätsch G, Wachter A (2016) Alternative splicing substantially diversifies the transcriptome during early photomorphogenesis and correlates with the energy availability in Arabidopsis. Plant Cell 28:2715–2734
Huang X, Ouyang X, Yang P, Lau OS, Li G, Li J, Chen H, Deng XW (2012) Arabidopsis FHY3 and HY5 positively mediate induction of COP1 transcription in response to photomorphogenic UV-B light. Plant Cell 24:4590–4606
Jang IC, Henriques R, Chua NH (2013) Three transcription factors, HFR1, LAF1 and HY5, regulate largely independent signaling pathways downstream of phytochrome A. Plant Cell Physiol 54:907–916
Jiang F, Guo M, Yang F, Duncan K, Jackson D, Rafalski A, Wang S, Li B (2012) Mutations in an AP2 transcription factor-like gene affect internode length and leaf shape in maize. PLoS ONE 7:e37040
Jiao YL, Ma LG, Strickland E, Deng XW (2005) Conservation and divergence of light-regulated genome expression patterns during seedling development in rice and Arabidopsis. Plant Cell 17:3239–3256
Jiao Y, Lau OS, Deng XW (2007) Light-regulated transcriptional networks in higher plants. Nat Rev Genet 8:217–230
Jin J, Zhang H, Kong L, Gao G, Luo J (2014) PlantTFDB 3.0: a portal for the functional and evolutionary study of plant transcription factors. Nucleic Acids Res 42:D1182–D1187
Kersey PJ, Allen JE, Christensen M, Davis P, Falin LJ, Grabmueller C et al (2014) Ensembl Genomes 2013: scaling up access to genome-wide data. Nucleic Acids Res 42:D546–D552
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL (2013) TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14:R36
Komeno Y, Yan M, Matsuura S, Lam K, Lo MC, Huang YJ, Tenen DG, Downing JR, Zhang DE (2014) Runx1 exon 6-related alternative splicing isoforms differentially regulate hematopoiesis in mice. Blood 123:3760–3769
Kong L, Zhang Y, Ye ZQ, Liu XQ, Zhao SQ, Wei L, Gao G (2007) CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res 35:W345–W349
Kurihara Y, Matsui A, Hanada K, Kawashima M, Ishida J, Morosawa T, Tanaka M, Kaminuma E, Mochizuki Y, Matsushima A, Toyoda T, Shinozaki K, Seki M (2009) Genome-wide suppression of aberrant mRNA-like noncoding RNAs by NMD in Arabidopsis. Proc Natl Acad Sci USA 106:2453–2458
Law M, Childs KL, Campbell MS, Stein JC, Olson AJ, Holt C et al (2015) Automated update, revision, and quality control of the maize genome annotations using MAKER-P improves the B73 RefGen_v3 gene models and identifies new genes. Plant Physiol 167:25–39
Lee J, He K, Stolc V, Lee H, Figueroa P, Gao Y, Tongprasit W, Zhao H, Lee I, Deng XW (2007) Analysis of transcription factor HY5 genomic binding sites revealed its hierarchical role in light regulation of development. Plant Cell 19:731–749
Leivar P, Quail PH (2011) PIFs: pivotal components in a cellular signaling hub. Trends Plant Sci 16:19–28
Lonosky PM, Zhang XS, Honavar VG, Dobbs DL, Fu A, Rodermel SR (2004) A proteomic analysis of maize chloroplast biogenesis. Plant Physiol 134:560–574
Lu T, Lu G, Fan D, Zhu C, Li W, Zhao Q, Feng Q, Zhao Y, Guo Y, Li W, Huang X, Han B (2010) Function annotation of the rice transcriptome at single-nucleotide resolution by RNA-seq. Genome Res 20:1238–1249
Ma LG, Li JM, Qu LJ, Hager J, Chen Z, Zhao H, Deng XW (2001) Light control of Arabidopsis development entails coordinated regulation of genome expression and cellular pathways. Plant Cell 13:2589–2607
Mandadi KK, Scholthof KB (2015) Genome-wide analysis of alternative splicing landscapes modulated during plant–virus interactions in Brachypodium distachyon. Plant Cell 27:71–85
Maquat LE (2005) Nonsense-mediated mRNA decay in mammals. J Cell Sci 118:3213–3213
Markelz NH, Costich DE, Brutnell TP (2003) Photomorphogenic responses in maize seedling development. Plant Physiol 133:1578–1591
Marquez Y, Brown JW, Simpson C, Barta A, Kalyna M (2012) Transcriptome survey reveals increased complexity of the alternative splicing landscape in Arabidopsis. Genome Res 22:1184–1195
Marquez Y, Hoepfler M, Ayatollahi Z, Barta A, Kalyna M (2015) Unmasking alternative splicing inside protein-coding exons defines exitrons and their role in proteome plasticity. Genome Res 25:995–1007
Min XJ, Powell B, Braessler J, Meinken J, Yu F, Sablok G (2015) Genome-wide cataloging and analysis of alternatively spliced genes in cereal crops. BMC Genomics 16:721
Nagasaki H, Arita M, Nishizawa T, Suwa M, Gotoh O (2005) Species-specific variation of alternative splicing and transcriptional initiation in six eukaryotes. Gene 364:53–62
Nawrocki EP, Kolbe DL, Eddy SR (2009) Infernal 1.0: inference of RNA alignments. Bioinform 25:1335–1337
Nawrocki EP, Burge SW, Bateman A, Daub J, Eberhardt RY, Eddy SR, Floden EW, Gradner PP, Jones TA, Tate J, Finn RD (2015) Rfam 12.0: updates to the RNA families database. Nucleic Acids Res 43:D130–D137
Ni ZF, Kim ED, Chen ZJ (2009) Chlorophyll and starch assays. Protocol Exchange. https://doi.org/10.1038/nprot.2009.12
Peterson ES, McCue LA, Schrimpe-Rutledge AC, Jensen JL, Walker H, Kobold MA et al (2012) VESPA: software to facilitate genomic annotation of prokaryotic organisms through integration of proteomic and transcriptomic data. BMC Genomics 13:131
Petrillo E, Godoy Herz MA, Fuchs A, Reifer D, Fuller J, Yanovsky MJ et al (2014) A chloroplast retrograde signal regulates nuclear alternative splicing. Science 344:427–430
Ponting CP (2008) The functional repertoires of metazoan genomes. Nat Rev Genet 9:689–698
Rehwinkel A, Letunic I, Raes J, Bork K, Izaurralde E (2005) Nonsense-mediated mrna decay factors act in concert to regulate common mrna targets. RNA 11:1530–1544
Resch A, Xing Y, Modrek B, Gorlick M, Riley R, Lee C (2004) Assessing the impact of alternative splicing on domain interactions in the human proteome. J Proteome Res 3:76–83
Roberts A, Pimentel H, Trapnell C, Pachter L (2011) Identification of novel transcripts in annotated genomes using RNA-Seq. Bioinformatics 27:2325–2329
Robinson MD, McCarthy DJ, Smyth GK (2010) EdgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W et al (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178–183
Shen Y, Zhou Z, Feng S, Li J, Tan-Wilson A, Qu LJ, Wang H, Deng XW (2009a) Phytochrome a mediates rapid red light-induced phosphorylation of Arabidopsis far-red elongated hypocotyl1 in a low fluence response. Plant Cell 21:494–506
Shen Z, Li P, Ni RJ, Ritchie M, Yang CP, Liu GF et al (2009b) Label-free quantitative proteomics analysis of etiolated maize seedling leaves during greening. Mol Cell Proteomics 8:2443–2460
Shikata H, Hanada K, Ushijima T, Nakashima M, Suzuki Y, Matsushita T (2014) Phytochrome controls alternative splicing to mediate light responses in Arabidopsis. Proc Natl Acad Sci USA 111:18781–18786
Shin S, Song Y, Ahn J, Kim E, Chen P, Yang S, Suh Y, Lee K (2015) A novel mechanism of myostatin regulation by its alternative splicing variant during myogenesis in avian species. Am J Physiol Cell Physiol 309:C650–C659
Song J, Cao K, Hao Y, Song S, Su W, Liu H (2019) Hypocotyl elongation is regulated by supplemental blue and red light in cucumber seedling. Gene 707:117–125
Staiger D, Brown JW (2013) Alternative splicing at the intersection of biological timing, development, and stress responses. Plant Cell 25:3640–3656
Syed NH, Kalyna M, Marquez Y, Barta A, Brown JW (2012) Alternative splicing in plants - coming of age. Trends Plant Sci 17:616–623
Thatcher SR, Zhou W, Leonard A, Wang BB, Beatty M et al (2014) Genome-wide analysis of alternative splicing in Zea mays: landscape and genetic regulation. Plant Cell 26:3472–3487
Thatcher SR, Danilevskaya ON, Meng X, Beatty M et al (2016) Genome-wide analysis of alternative splicing during development and drought stress in maize. Plant Physiol 170:586–599
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ et al (2010) Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28:511–515
Trapnell C, Roberts A, Goff L, Pertea G, Kim D et al (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7:562–578
Walley JW, Shen Z, Sartor R, Wu KJ, Osborn J, Smith LG, Briggs SP (2013) Reconstruction of protein networks from an atlas of maize seed proteotypes. Proc Natl Acad Sci USA 110:E4808–E4817
Wang J, Delabie J, Aasheim HC, Smeland E, Myklebost O (2002) Clustering of the some easily reveals distinct gene expression patterns: results of a reanalysis of lymphoma study. BMC Bioinform 3:36
Wang Z, Xue Z, Wang T (2014) Differential analysis of proteomes and metabolomes reveals additively balanced networking for metabolism in maize heterosis. J Proteome Res 13:3987–4001
Wong MS, Wright WE, Shay JW (2014) Alternative splicing regulation of telomerase: a new paradigm? Trends Genet 30:430–438
Wu HP, Su YS, Chen HC, Chen YR, Wu CC, Lin WD, Tu SL (2014) Genome-wide analysis of light-regulated alternative splicing mediated by photoreceptors in Physcomitrella patens. Genome Biol 15:R10
Zhao Y, Sun J, Xu P, Zhang R, Li L (2014) Intron-mediated alternative splicing of wood-associated NAC transcription factor1b regulates cell wall thickening during fiber development in Populus species. Plant Physiol 164:765–776
Zheng L, Huang F, Narsai R, Wu J, Giraud E, He F et al (2009) Physiological and transcriptome analysis of iron and phosphorus interaction in rice seedlings. Plant Physiol 151:262–274
Acknowledgements
This work was supported by the “Strategic Priority Research Program” of the Chinese Academy of Sciences (Grant No. XDA24010203), and the National Key Research and Development Program of China (Grant No. 2016YFD0101003). We thank Yuxian Zhu and Qun He for critically reading the article and Zhen Xue for technical assistance. We thank Dr. Tiancong Lu (Protein World Biotechnology) for giving suggestions on proteomic data analysis. We thank Dr. Melissa Lehti-Shiu (Lehti Life Science Editing) for proofreading the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All the authors have declared no competing interests.
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
425_2020_3464_MOESM1_ESM.pdf
Fig. S1 PCR validation of alternative splicing events. Results of PCR validation of a, exon skipping b, intron retention, c, alternative 3’ splicing, d, alternative 5’ splicing and e, novel junction site alternative splicing events are shown in the figure. In a to e, the upper panel shows the PCR validation results and the lower panel shows the alignment between the sequenced and annotated sequence. Black boxes indicate exons, small black boxes indicate retained parts of the UTR, dark grey boxes indicate introns, light grey boxes indicate excluded parts of the UTR, orange boxes indicate parts excluded in novel transcripts and the blue blocks indicate parts retained in novel transcripts (PDF 6293 kb)
425_2020_3464_MOESM2_ESM.pdf
Fig. S2 PCR validation of novel transcripts. Four gel images of novel transcripts validated by PCR are shown in the upper panel. The transcript and genomic sequences for each transcript were aligned. The alignment result for TCONS_00098186 indicates that the novel transcript prediction was accurate (PDF 1428 kb)
425_2020_3464_MOESM3_ESM.pdf
Fig. S3 MapMan analysis of three DEU gene sets. Three DEU gene sets were enriched in 59 MapMan functional categories (FDR < 0.05). The colour shows the degree of significance, the size of the circle indicates the ratio of the number of DEU proteins to the total number of maize proteins assigned to each term. Horizontal axis shows the three DEU gene sets (PDF 256 kb)
425_2020_3464_MOESM4_ESM.pdf
Fig. S3 Correlation between expression level determined by qRT-PCR and RNA-seq. Horizontal axis and longitudinal axis show the expression levels of 20 tested transcripts from analysis of our RNA-seq data and qRT-PCR (PDF 210 kb)
425_2020_3464_MOESM5_ESM.pdf
Fig. S5 Expression profiles of each DEG cluster. K-means cluster analysis of DEGs in the blue light, white light, red light and blue red mixed light treatments (FDR < 0.05) (PDF 1441 kb)
425_2020_3464_MOESM6_ESM.pdf
Fig. S6 Level 1 MapMan category enrichment analysis of differentially expressed genes in the six main clusters identified for each light treatment. K-means cluster analysis of DEGs in the blue (B) light, red (R) light, blue red mixed (B7R) light and white (W) light treatments. The colour of each box shows the -log10(P) value. Grey to red, significant enrichment; white, not significant (PDF 750 kb)
425_2020_3464_MOESM7_ESM.pdf
Fig. S7 GO slim enrichment analysis of differentially expressed genes in the six main clusters identified for each light treatment. DEGs in clusters 1 to 6 in Fig.S6 were enriched in GO biological process (BP), cell component (CC) and molecular function (MF) categories (FDR < 0.05). The colour shows the degree of significance and the size of the circle indicates the protein ratio for each GO term. Horizontal axis shows the four types of light treatment (PDF 341 kb)
425_2020_3464_MOESM8_ESM.pdf
Fig. S8 Analysis of transcription factor enrichment among six main clusters for each light treatment. K-means cluster analysis of transcription factor family members in the blue (B) light, red (R) light, blue red mixed (B&R) light and white (W) light treatments (FDR < 0.05). The colour of each box shows the -log10(P) value. Grey to red, significant enrichment; white, not significant (PDF 829 kb)
425_2020_3464_MOESM9_ESM.pdf
Fig. S9 Clusters of genes differentially expressed among four light treatments after 6 hours of exposure. In total, 884 differentially expressed genes (DEGs) were identified and could be classified into 10 expression clusters. Clusters 9 and 10, corresponding to genes only differentially expressed in response to red light, contained the most (-72.9%) DEGs. The total number of DEGs in each cluster is shown on the top right corner (PDF 773 kb)
425_2020_3464_MOESM10_ESM.pdf
Fig. S10 Venn diagram of DEGs identified by pairwise comparison between blue light, red light , and blue and red mixed light after 6 hours of exposure. The number (in the blue circle) on the left shows the number of genes differentially expressed between the blue (B) light treatment and red (R) light treatment after 6 hours of exposure. The number in the red circle on the right shows the number of genes differentially expressed between the B light treatment and blue and red mixed (B&R) light treatment after 6 hours of exposure. The number in the yellow circle shows the number of genes differentially expressed between the R light treatment and B&R mixed light treatment after 6 hours of exposure. The total numbers of DEGs from the B, R and B&R mixed light treatments are 789, 143 and 610, respectively (PDF 393 kb)
425_2020_3464_MOESM11_ESM.pdf
Fig. S11 The non-additive effect of blue and red light on gene expression. Numbers in the big circle show the number of DEGs between blue (B) light treatment and red light treatment after 6 hours of exposure. Numbers in the small circle show the average expression levels under the between B and R mixed light (B&R) treatment and the average expression levels between the B and R light treatments ((B+R)/2) (PDF 210 kb)
425_2020_3464_MOESM12_ESM.pdf
Fig. S12 Models of three novel genes for the validation of alternative splicing in this study. a TCONS_00100007_19 is a novel transcript identified in the AC190628.4_FG007 locus and is longer than the annotated sequence. Four perfect matched peptides span the splicing junctions in TCONS_00100007_19 and the corresponding mass spectra are shown below the figure. b TCONS_00080402_23 is a novel transcript identified in the annotated GRMZM2G020865 locus. Peptide FQGISAAPHQFPESHLVK span the splicing junction in TCONS_00080402_23, and the corresponding peptide mass spectrum is shown under the figure. c TCONS_00152356_13 is a novel transcript identified in the AC234154.1_FG007 locus. Peptide LDGGEMLACGLATHFVQSNSLLSLEESLK span the splicing junction in TCONS_00152356_13, and the corresponding peptide mass spectra is shown under the figure. The green block represents an exon. Yellow lines represent the sequences translated before the junctions, and purple lines represent the sequences translated after the junction. Blue characters in mass spectra are b ions and red characters are y ions (PDF 1113 kb)
425_2020_3464_MOESM13_ESM.pdf
Fig. S13 Clusters of differentially expressed proteins among four light treatments after 6 hours of exposure. K-means clustering analysis of 1867 differentially expressed proteins (DEPs) in the blue light, red light and blue-red mixed light treatments after 6 hours of exposure (FDR < 0.05, fold-change ≥ ±1.5). Total number of DEPs in each cluster is shown on the top right corner (PDF 1535 kb)
425_2020_3464_MOESM14_ESM.pdf
Fig. S14 MapMan analysis of cluster 1 and cluster 2 proteins in Fig. S13. Differentially expressed proteins in cluster 1 and cluster 2 were enriched in 23 MapMan functional categories (FDR < 0.05). The colour shows the degree of significance, and the size of the circle shows the protein ratio for each term. Horizontal axis shows the total number of proteins annotated to each GO term (PDF 187 kb)
425_2020_3464_MOESM15_ESM.pdf
Fig. S15 GO slim analysis of cluster 1 and cluster 2 proteins in Fig. S13. Differentially expressed proteins in cluster 1 and cluster 2 were enriched in GO biological process (BP), cell component (CC) and molecular function (MF) categories (FDR < 0.05). The colour shows the degree of significance, and the size of the circle indicates the protein ratio. Horizontal axis shows the total number of proteins annotated to each GO term (PDF 150 kb)
425_2020_3464_MOESM16_ESM.pdf
Fig. S16 HY5 proteins differentially expressed in the proteome after 6h of light treatment. a Histogram of protein expression. Three HY5 proteins (GRMZM2G039828_P01, GRMZM2G137046_P01, and GRMZM2G171912_P02) were expressed at much lower levels in R6h than in B6h, B&R6h, and W6h. b to d Peptide spectra of three HY5 proteins (PDF 301 kb)
425_2020_3464_MOESM17_ESM.pdf
Fig. S17 MapMan analysis of genes encoding multiple protein isoforms. The genes encoding multiple protein isoforms were enriched in PS and TCA/org. transformation categories (FDR < 0.05). The colour shows the degree of significance and the size of the circle shows the ratio of the number of DEGs to the total number of maize genes assigned to each term. Horizontal axis shows the total number of genes annotated to each MapMan category (PDF 124 kb)
425_2020_3464_MOESM18_ESM.pdf
Fig. S18 GO slim analysis of the genes with protein isoforms. Genes with multiple protein isoforms were enriched in GO biological process (BP), cell component (CC) and molecular function (MF) categories (FDR < 0.05). The colour shows the degree of significance, and the size of the circle shows the protein ratio for each GO term. Horizontal axis shows the total number of genes annotated to each GO term (PDF 153 kb)
425_2020_3464_MOESM19_ESM.pdf
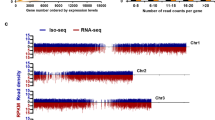
Fig. S19 Difference in RNA expression levels (FPKM values) between genes with multiple protein isoforms and genes with only one isoform. The density on the Y-axis indicates the area under the curve of a density function representing the probability of obtaining a given x value between a range of x values. Green, genes with multiple isoforms; Red, genes with a single isoform (PDF 968 kb)
425_2020_3464_MOESM20_ESM.pdf
Fig. S20 Photosynthesis proteins differentially expressed in response to one of the four light treatments at 6 hours of exposure. a The number of proteins identified in blue (B), red (R) and blue-red mixed (B&R) light treatment samples. b Venn diagram analysis of proteins identified in B, R and B&R treatment samples (PDF 347 kb)
Table S5 Mapman enrichment analysis of clusters of genes differentially expressed at 6 h (PDF 14 kb)
425_2020_3464_MOESM34_ESM.pdf
Table S14 Differences in the domain contents and expression levels of two GRMZM2G017957 isoforms resulting from alternative splicing (PDF 9 kb)
425_2020_3464_MOESM35_ESM.pdf
Table S15 Proteins involved in photosynthesis differentially expressed in response to the four light treatments at 6 hours of exposure (PDF 207 kb)
Rights and permissions
About this article
Cite this article
Yan, Z., Shen, Z., Li, Z. et al. Genome-wide transcriptome and proteome profiles indicate an active role of alternative splicing during de-etiolation of maize seedlings. Planta 252, 60 (2020). https://doi.org/10.1007/s00425-020-03464-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-020-03464-5