Abstract
Hypoxia, or oxygen deficiency, is an abiotic stress that plants are subjected to during soil flooding. Therefore, plants have evolved adaptive mechanisms to sense oxygen deficiency and make coordinated changes at the transcriptional level. The results of this study show that the interplay between hydrogen peroxide and ethylene affected the transcriptional responses of ERF73/HRE1 and ADH1 during hypoxia signaling. H2O2 affected the abundance of ERF73/HRE1 and ADH1 mRNAs in both wild-type Arabidopsis and the ethylene-insensitive mutant, ein2-5. Promoter analysis was conducted using transgenic plants expressing an ERF73/HRE1 promoter–β-glucuronidase reporter gene construct. GUS staining observations and activity assays showed that GUS was regulated similarly to, and showed a similar accumulation pattern as, H2O2 during hypoxia. The transcript levels of ERF73/HRE1 and ADH1 were significantly decreased in the WT by combined hypoxia and diphenylene iodonium chloride (DPI, an NADPH oxidase inhibitor) treatment. In ein2-5, induction of ERF73/HRE1 was also reduced significantly by the combined hypoxia and DPI treatment. In contrast, ADH1 mRNA levels only slightly decreased after this treatment. When DPI was supplied at different time points during hypoxia treatment, H2O2 had critical effects on regulating the transcript levels of ERF73/HRE1 and ADH1 during the early stages of hypoxia signaling. The induction of hypoxia-inducible genes encoding peroxidases and cytochrome P450s was affected, and accumulation of H2O2 was reduced, in ein2-5 during hypoxic stress. Together, these results demonstrate that H2O2 plays an important role during primary hypoxia signaling to control the transcriptional responses of ERF73/HRE1 and ADH1 via modulation of ethylene signaling.
Similar content being viewed by others
Introduction
Under oxygen-deficient conditions, toxic products from anaerobic respiration and reactive oxygen species (ROS) can accumulate in plant cells. ROS such as superoxide radicals (·O2 −), hydroxyl radicals (·OH) and hydrogen peroxide (H2O2) are all very reactive, and can severely damage membranes (Mittler et al. 2004). Hypoxia leads to serious physical damage in plant cells but little is known about the interplay between H2O2 and ethylene during hypoxia signaling. H2O2 can function as a signaling molecule to induce responses to biotic or abiotic stresses, including ozone, salinity, chilling, wounding, and pathogen attack (Neill et al. 2002; Sagi et al. 2004; Valenzuela-Soto et al. 2011; Wang et al. 2012). It was reported that hypoxia triggers H2O2 production via a Rop-signal transduction pathway in Arabidopsis (Baxter-Burrell et al. 2002). A recent study showed that H2O2 and ethylene mediate the antioxidant responses of grapevines’ buds to hypoxia (Vergara et al. 2012). H2O2 production under O2 deprivation occurs during an early phase of anoxia, and ROS are required to regulate a set of genes belonging to heat-shock protein and ROS-mediated groups (Banti et al. 2010; Pucciariello et al. 2012). Both ethylene and H2O2 promote formation of aerenchyma in rice stems in a dose-dependent manner (Steffens et al. 2011). In Arabidopsis, the production of lysigenous aerenchyma requires both ethylene and H2O2 signals in hypoxic conditions (Muhlenbock et al. 2007). H2O2 is required for ethylene-induced epidermal cell death and is sufficient to promote it in Oryza sativa (Steffens and Sauter 2005, 2009).
Phytohormones participate in regulating stress-inducible genes to help plants to adapt to various environmental stresses. In particular, ethylene is not only involved in plant growth and development, but is also involved in hypoxic stress (Ophir et al. 2009; Yang et al. 2011). Under oxygen-deficient conditions, ethylene accumulation and components of the ethylene response factor group VII, ERF73/HRE1, which are involved in modulating ethylene responses, regulate the expression of alcohol dehydrogenase 1 (ADH1) (Peng et al. 2005; Yang et al. 2011). Arabidopsis ERF73/HRE1 is specifically induced during hypoxia and plays a key role in activating ADH1 and other genes related to fermentation and carbohydrate metabolism such as PDC1, PDC2, SUS1, SUS4, and LDH (Licausi et al. 2010; Yang et al. 2011). Hypoxia triggers ethylene-dependent and ethylene-independent pathways, both of which are required for increased transcript accumulation of ERF73/HRE1. Studies on rice showed that the SNORKEL and SUB1A proteins regulate opposite growth responses. These proteins play a major role in submergence tolerance, and all of them belong to the group VII ERF subfamily (Xu et al. 2006; Hattori et al. 2009; Pena-Castro et al. 2011).
Direct and indirect oxygen-sensing mechanisms may be responsible for acclimation responses that prolong survival when plants are oxygen deprived (Bailey-Serres et al. 2012). Recently, two independent studies suggested that the N-end rule pathway (NERP) for protein degradation may be involved in direct oxygen sensing, thereby regulating the low-O2 response in plants. The Met–Cys N-end has been found in HRE1, HRE2, RAP2.2, RAP2.3, and RAP2.12 of Arabidopsis, as well as the rice proteins SUB1A, SUB1B, and SK1/SK2 (Bailey-Serres et al. 2012). This Met–Cys motif permits the degradation of these proteins under oxygen-replete conditions, and their accumulation under hypoxia, providing a mechanism for homeostatic sensing of low oxygen levels in plants (Gibbs et al. 2011; Licausi et al. 2011). The production of ROS and nitric oxide (NO), adenylate levels (ATP, ADP, and/or AMP), consumable carbohydrates, pyruvate, cytosolic pH, and/or cytosolic Ca2+ may be involved in indirect oxygen-sensing mechanisms (Bailey-Serres et al. 2012).
The H2O2 generated under hypoxic conditions acts as a signal to trigger downstream responses that lead to activation of ADH1 (Baxter-Burrell et al. 2002; Peng et al. 2005). However, the connections between H2O2 and ethylene during hypoxic stress are still poorly understood. The results of this study demonstrate that cross-talk between H2O2 and ethylene under hypoxic conditions affects transcript accumulation of downstream hypoxia-inducible genes. The transcript levels of ERF73/HRE1 and ADH1 were increased by H2O2 treatment in wild-type Arabidopsis and ein2-5, an ethylene-signaling mutant. The results also show that H2O2 is involved in a primary hypoxia signaling pathway to regulate transcript levels of ERF73/HRE1 and ADH1. Moreover, ethylene signaling and its interplay with H2O2 partially modulate ADH1 in the hypoxia pathway.
Materials and methods
Plant materials, growth conditions and stress treatment
Arabidopsis thaliana ecotype Columbia (Col-0) seeds were obtained from the Arabidopsis Biological Resource Center, Ohio State University, USA. The ethylene-insensitive mutant ein2-5 (Col-0) seeds used in this study were obtained from Dr. L.-C. Wang (Institute of Plant and Microbial Biology, Academia Sinica, Taiwan). All seeds were sterilized and kept for 3 days at 4 °C in the dark to break dormancy. Seeds were germinated on 0.8 % (w/v) agar plates containing 1/2-strength MS medium (pH 5.7) supplemented with 0.5 % (w/v) sucrose. The plates were oriented vertically and incubated at 22 °C with a 16-h-light (81 mmol s−1 m−2)/8-h-dark cycle in a growth chamber. Subsequently, 7-day-old seedlings were transplanted onto fresh plates that were oriented vertically to prevent root growth into the medium. Seedlings were used in experiments when they were 14-day old. For the hypoxia treatment or hypoxia +DPI treatment, seedlings were placed on a floating platform with their roots immersed in 1/2 MS solution containing 0.5 % (w/v) sucrose supplemented with or without 30 μM DPI at pH 5.7. The solution was constantly supplied with 3 % (v/v) oxygen and 97 % (v/v) nitrogen in the dark for indicated times. For the H2O2 treatment, 14-day-old seedlings were placed on filter paper moistened with 1/2 MS solution supplemented with 3 mM H2O2 and kept in the dark for the indicated times.
Quantitative RT-PCR analyses
Root samples were collected from seedlings and frozen until analysis. RNA was isolated from frozen tissues using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). Total RNA samples from roots were first treated with DNase I and then reverse transcribed into cDNA by Moloney murine leukemia virus reverse transcriptase (Invitrogen). Quantitative RT-PCR was performed using a 7500 Real-Time PCR System (Applied Biosystems, Foster City, CA, USA) with Power SYBR Green PCR Master Mix (Applied Biosystems), according to the manufacturer’s instructions. The relative expression level of each gene was quantified using the comparative threshold cycle method, as described in the manufacturer’s instructions for the ABI PRISM 7500 Sequence Detection System (Applied Biosystems). The actin gene was used as an internal control to normalize the cDNA levels. Data were analyzed using the 7500 System SDS software, version 1.2.3 (Applied Biosystems). Amplification conditions were as follows: 95 °C for 10 min, and then 40 cycles at 95 °C for 15 s and 60 °C for 1 min. The primers used for quantitative RT-PCR analyses are listed in Table 1. Quantitative RT-PCR experiments were repeated three times independently in duplicate, and the data were averaged.
Construction of the ERF73/HRE1-GUS fusion gene
To generate ERF73/HRE1 pro -uidA transgenic plants, the 5′ upstream promoter region (−1108 to +91) relative to the putative initiation codon of the ERF73/HRE1 gene was PCR-amplified from Arabidopsis Col-0 genomic DNA and cloned into a binary vector (pMDC164) using the Gateway method for transformation. Primer sequences are listed in Table 1. The recombinant plasmid was transformed into Agrobacterium tumefaciens strain GV3101, and homozygous T3 plants were used for the GUS activity assay and histochemical staining.
GUS activity assay and histochemical staining
Protein extraction from 14-day-old seedlings, for use in GUS activity assays, was performed as described previously (Jefferson 1989). The protein concentration in the extract was determined using a Bio-Rad Protein Assay kit. Fluorescence was measured in a Microplate Spectrofluorometer (Wallac Victor3V 1420, Multilabel Counter; Perkin-Elmer, Turku, Finland) at excitation/emission 365/455 nm. Each assay was repeated four times. Data were obtained from at least seven independent lines. The GUS-positive seedlings were visualized under a Zeiss SteREO Lumar V12 Fluorescence Stereo Microscope.
H2O2 measurement
The 14-day-old seedlings of the wild type and ein2-5 were subjected to hypoxic conditions as described above. The production of H2O2 in roots was assessed using the Amplex Red Hydrogen Peroxide/Peroxidase Assay Kit (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. Roots were ground in liquid nitrogen, and then 500 ml phosphate buffer (20 mM K2HPO4, pH 6.5) was added to 50 mg ground frozen tissue. After centrifugation, protein concentrations were determined by conducting a dye-binding Bio-Rad protein assay according to the supplier’s instructions, and then adjusting each sample to the same protein concentration using a phosphate buffer. A 50-μl sample extract was incubated with 0.2 U ml−1 horseradish peroxidase and 100 μM Amplex Red reagent (10-acetyl-3,7-dihydrophenoxazine) at room temperature for 30 min in the dark. Fluorescence was measured with a Microplate Spectrofluorometer (Wallac Victor3V 1420, Multilabel Counter; Perkin-Elmer) at excitation/emission 535/590 nm. The experiments were repeated least five times independently, each time in duplicate.
Results
H2O2 induces transcript levels of ERF73/HRE1 and ADH1 in the wild type and ein2-5
Previously, we showed that transcript accumulation of ERF73/HRE1 and ADH1 was induced by ethylene under hypoxic conditions in Arabidopsis, and also demonstrated that ERF73/HRE1 negatively regulated a group of genes encoding peroxidases and cytochrome P450 under hypoxic stress (Peng et al. 2005; Yang et al. 2011). An increase in the H2O2 level was shown to regulate transcript accumulation of ADH1 (Baxter-Burrell et al. 2002). To further investigate the relationship between H2O2 and ethylene signals in regulating transcript levels of ERF73/HRE1 and ADH1, we quantified the transcript levels of ERF73/HRE1 and ADH1 mRNA after H2O2 treatment for 0, 0.5, 1, 3, and 6 h in wild-type Arabidopsis [ecotype Columbia (Col-0)] and ein2-5.
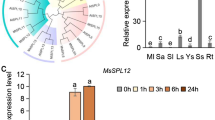
The quantitative RT-PCR results showed that transcript levels of both ERF73/HRE1 and ADH1 were induced by the 3 mM H2O2 treatment in the roots of wild-type and ein2-5 seedlings (Fig. 1a, b). The transcript levels of ERF73/HRE1 were highest after 1 and 6 h of the H2O2 treatment in Col and ein2-5, respectively. The transcript level of ADH1 was highest after 3 h of the H2O2 treatment in both the wild type and ein2-5. The ADH1 transcript level strongly decreased after 6 h of the H2O2 treatment in Col but only slightly decreased in ein2-5. These results showed that transcript accumulation of ERF73/HRE1 and ADH1 was induced by H2O2 in both the wild type and ein2-5, and also implied that H2O2 was still able to induce the transcript levels of ERF73/HRE1 and ADH1 in ein2-5, even though ethylene signaling is disrupted in this mutant.
Effects of H2O2 on expression levels of ERF73/HRE1 and ADH1 in Arabidopsis. Quantitative RT-PCR analyses of ERF73/HRE1 and ADH1 transcript levels in Col (a) and ein2-5 (b) in response to H2O2. Total RNAs were isolated from roots of 14-day-old seedlings after H2O2 (3 mM) treatment for 0, 0.5, 1, 3 and 6 h. The levels of ERF73/HRE1 and ADH1 transcripts were determined. Relative transcript amounts were calculated and normalized to actin mRNA levels. Values represent mean ± SD from three biologically independent experiments. *P < 0.05 versus value at time zero (one-way ANOVA, Dunnett’s t test)
Activity of the ERF73/HRE1 promoter under hypoxic and H2O2 conditions
To characterize the expression pattern of the ERF73/HRE1 promoter in response to hypoxia and H2O2, we generated transgenic plants harboring the ERF73/HRE1 promoter fused to the uidA gene, which encodes the β-glucuronidase (GUS) reporter enzyme. Seven independent homozygous transgenic lines for the ERF73/HRE1 pro -uidA construct were analyzed and consistent patterns were detected (Fig. 2). The 14-day-old ERF73/HRE1 pro -uidA seedlings were subjected to hypoxia or 3 mM H2O2 for 1–3 h. In roots, GUS expression was detected in the root tip and vasculature after hypoxia or H2O2 treatment. No GUS expression was observed in normoxic (control) conditions (Fig. 2a). The GUS activity assay showed that GUS was induced by both hypoxia and H2O2 treatment for 1 or 3 h; the highest GUS activity was detected after 3 h of the hypoxia treatment (Fig. 2c). GUS staining was also observed in the vascular bundle of the primary root after hypoxia or H2O2 treatment for 3 h (Fig. 2a). In leaves, significant GUS staining was observed in the trichomes and guard cells of plants under the hypoxia and H2O2 treatments (Fig. 2b). These results indicate that the promoter activity of ERF73/HRE1 was regulated by both H2O2 and hypoxia, which presented a similar induction pattern in the cellular response.
Histochemical analysis of the ERF73/HRE1 promoter by observations of GUS activity after treatment with hypoxia or H2O2 in transgenic Arabidopsis plants. a GUS expression in homozygous lines carrying an ERF73/HRE1 promoter-GUS construct after normoxic (Nor), hypoxic (Hyp), or H2O2 treatment for 1 and 3 h. b GUS expression in homozygous lines carrying an ERF73/HRE1 promoter–GUS construct in roots, trichomes, and stomata after normoxic, hypoxic, or H2O2 treatment for 3 h. c GUS activity in homozygous lines carrying an ERF73/HRE1 promoter–GUS construct after normoxic, hypoxic, or H2O2 treatment for 1 and 3 h. Data were collected using at least seven independent lines. Values represent mean ± SD from four biologically independent experiments. *P < 0.05, versus value in normoxic treatment (one-way ANOVA, Dunnett’s t test)
Cross-talk between ethylene and H2O2 is involved in regulating ERF73/HRE1 and ADH1 in hypoxia pathways
Previously, we reported that ethylene was accumulated during hypoxia in the wild type and involved in inducing ERF73/HRE1 and ADH1 (Yang et al. 2011). In addition, the production of H2O2 under hypoxia may be involved in indirect oxygen-sensing mechanisms (Bailey-Serres et al. 2012). In this study, our results showed that H2O2 influenced the accumulation of ERF73/HRE1 and ADH1 mRNA transcripts in both wild-type Arabidopsis and ein2-5 (Fig. 1). To further test whether H2O2 production is involved in regulating ERF73/HRE1 in hypoxia pathways, we analyzed the effects of diphenylene iodonium chloride (DPI) on transcript levels of ERF73/HRE1 and ADH1 during hypoxic stress. DPI binds strongly to flavoproteins and thus inhibits NADPH oxidases and nitric oxide synthase (NOS) activity, which blocks the production of H2O2 (Arasimowicz-Jelonek et al. 2012). We conducted quantitative RT-PCR to evaluate the transcript levels of ERF73/HRE1 and ADH1 in Col after hypoxia treatment (Fig. 3). The transcript levels of ERF73/HRE1 and ADH1 were significantly decreased in Col by the combined hypoxia and DPI treatment (Fig. 3). These results provided evidence that H2O2 participated in regulating ERF73/HRE1 and ADH1 transcript accumulation during hypoxic stress.
Repression of hypoxia-induced ERF73/HRE1 and ADH1 by DPI in Col and ein2-5. The effect of hypoxia combined with DPI treatment on transcript levels of ERF73/HRE1 and ADH1 as determined by quantitative RT-PCR analysis. Total RNAs were isolated from 14-day-old roots of Col and ein2-5 seedlings after hypoxia with or without DPI treatment for 0.5, 1, 3, 6 or 9 h. Levels of ERF73/HRE1 and ADH1 mRNA were determined. Relative amounts of mRNA were calculated and normalized to actin mRNA. Values represent mean ± SD from three biologically independent experiments
To further analyze the relationship between H2O2 and ethylene signaling under hypoxic stress, we analyzed the transcript levels of ERF73/HRE1 and ADH1 in ein2-5. Hypoxic induction of ERF73/HRE1 in ein2-5 was substantially reduced with the addition of DPI, but transcript levels of ADH1 were only slightly decreased (Fig. 3). This result showed that ERF73/HRE1 and ADH1 mRNA were regulated via H2O2 and ethylene signals under hypoxic stress. Taken together, these results suggested that the expression of ERF73/HRE1 and ADH1 in Arabidopsis was regulated by ethylene and H2O2 cross-talk during hypoxic stress.
The effects of H2O2 occur mainly at the early stages of hypoxia signaling pathways
To investigate the effect of H2O2 during hypoxia signaling, we supplied DPI after 0, 0.5, 1, and 2 h of hypoxia treatment. All samples were harvested after 3 h of the hypoxia treatment. The addition of DPI suppressed transcripts of ERF73/HRE1 and ADH1 under the hypoxia treatment. Compared with the transcript level under hypoxic conditions (without DPI), the level of ERF73/HRE1 mRNA was 10.6-fold lower when DPI was added at 0 h of the hypoxia treatment, but only 3.4-fold lower when DPI was added at 0.5 h of the hypoxia treatment (Fig. 4). A similar repression effect of DPI was detected for ADH1 mRNA. Comprehensive previous results have indicated that H2O2 has critical effects that regulate the expression of ERF73/HRE1 and ADH1 during the early stages of hypoxia signaling.
Inhibition effect of DPI on hypoxia-induced AtERF73/HRE1 and ADH1 in Col. Effect of H2O2 on transcript levels of AtERF73/HRE1 and ADH1 during hypoxia, as determined by quantitative RT-PCR. Total RNAs were isolated from 14-day-old roots of Col seedlings after hypoxia without DPI treatment for 3 h or after a 3-h hypoxia treatment with addition of DPI at 0, 0.5, 1, or 2 h during the hypoxia treatment. Levels of AtERF73/HRE1 and ADH1 mRNA were determined. Relative amounts of mRNA were calculated and normalized to actin mRNA. Values represent mean ± SD from three biologically independent experiments. *P < 0.05, versus 3 h hypoxia treatment without DPI (one-way ANOVA, Dunnett’s t test)
Induction of hypoxia-inducible genes was affected in ein2-5 during hypoxic stress
To determine how ethylene signaling affects hypoxia responses, we quantified the transcript levels of peroxidase (At3g03670, At1g14540, At1g14550, and At5g39580), cytochrome P450 CYP82C2 (At4g31970), and CYP81F2 (At5g57220) genes in ein2-5 during hypoxic stress. Quantitative RT-PCR analyses showed that the transcript levels of hypoxia-inducible peroxidase and cytochrome P450 genes (except for At5g39580) were reduced in ein2-5 during hypoxic stress (Fig. 5), compared with their respective levels in the wild type. This result suggested that ethylene signaling modulated these hypoxia-inducible genes during hypoxia signaling in Arabidopsis.
Transcript levels of hypoxia-inducible peroxidase and cytochrome P450 genes in ein2-5. Total RNAs were isolated from 14-day-old roots of Col (filled square) or ein2-5 (open square) seedlings after hypoxia treatment for 0, 0.5, 1, 3, 6 or 9 h. Quantitative RT-PCR analysis of the transcript levels of peroxidase (At3g03670, At1g14540, At1g14550, and At5g39580) and cytochrome P450 (At4g31970 and At5g57220) genes. Values represent mean ± SD from three biologically independent experiments. *P < 0.05, versus wild-type (one-way ANOVA, Dunnett’s t test)
Accumulation of H2O2 was reduced in ein2-5 under hypoxic stress
The transcript levels of ERF73/HRE1 and ADH1 were reduced after the combined hypoxia and DPI treatment (Fig. 3). To further clarify the effect of ethylene signaling on the H2O2 level during hypoxic stress, we quantified H2O2 during hypoxia in ein2-5. The H2O2 content was increased by 2.5-fold in the roots of 14-day-old wild-type seedlings after a 3-h hypoxia treatment. However, H2O2 was accumulated to lower levels in ein2-5 after hypoxia treatment for 1, 3 and 6 h, compared with its levels in the wild-type at the corresponding time points (Fig. 6). This result showed that ethylene signaling influenced H2O2 accumulation during hypoxic stress.
Effect of hypoxia-induced H2O2 accumulation in ein2-5. Levels of H2O2 in roots of 14-day-old Col and ein2-5 seedlings were determined after hypoxia treatment for 0, 0.5, 1, 3 and 6 h. Values represent mean ± SD from least five biologically independent experiments. *P < 0.05, versus wild type (one-way ANOVA, Dunnett’s t test)
Discussion
ROS production has been suggested to be a component of hypoxia signaling, and ROS are required for regulating a set of genes that belong to the heat-shock protein and ROS-mediated groups (Pucciariello et al. 2012). In contrast to other ROS, H2O2 has a relatively long half-life and can be produced in all cellular compartments. Increases in H2O2 accumulation under hypoxic conditions have been detected in several plant systems (Baxter-Burrell et al. 2002; Vergara et al. 2012). The activation of a ROP protein induces H2O2 accumulation during hypoxic stress via an NADPH oxidase mechanism, which was shown to be required for ADH1 expression and activity (Baxter-Burrell et al. 2002). During oxygen deprivation, the ubiquitin-dependent N-end rule pathway is involved in degrading the transcription factor ERF73/HRE1. The level of degraded proteins acts as a homeostatic sensor of hypoxia signaling in Arabidopsis (Gibbs et al. 2011; Licausi et al. 2011). In contrast, little is known about the transcriptional regulation upstream of ERF73/HRE1 during hypoxia signaling.
Previously, we reported that ethylene was involved in the hypoxia signal to regulate hypoxia marker genes such as ERF73/HRE1 and ADH1. The results of our previous studies showed that hypoxia-responsive ERF73/HRE1 expression is mediated by ethylene-dependent and ethylene-independent pathways, and that ethylene is necessary but not alone sufficient for the hypoxic induction of ERF73/HRE1 (Peng et al. 2005; Yang et al. 2011). In this study, our analyses showed that H2O2 affects the expression of ERF73/HRE1 and ADH1 mRNA in both wild-type Arabidopsis and ein2-5. The induction levels of ERF73/HRE1 and ADH1 mRNA were even persistent at 6 h in ein2-5 (Fig. 1). Our results imply that the effect of H2O2 under hypoxia is not completely dependent on ethylene signaling; therefore, whether H2O2 signaling cascades are related to the N-end rule pathway that modulates ERF73/HRE1 expression requires further investigation. We analyzed the transcript levels of ERF73/HRE1 and ADH1 after combined hypoxia and DPI treatment in ein2-5, and found that, although the induction of ERF73/HRE1 was substantially reduced, the level of ADH1 mRNA only slightly declined with the addition of DPI (Fig. 3). Moreover, the transcript level of ERF73/HRE1 remained at 2.8-fold induction when DPI was supplied at the beginning of hypoxia treatment for 3 h in the wild type (Fig. 4). This result demonstrated that H2O2 acted as a signaling molecule during primary hypoxia signaling, and had a critical effect on the regulation of ERF73/HRE1 and ADH1 transcript accumulation. The results also indicated that the control of downstream gene transcription by H2O2 did not occur entirely through ethylene signaling during hypoxic stress.
Our results also showed that accumulation of H2O2 was reduced in ein2-5 during hypoxic stress (Fig. 6). The transcript levels of most of the hypoxia-inducible peroxidase and cytochrome P450 genes were reduced in ein2-5 during hypoxic stress, except for the peroxidase gene At5g39580 (Fig. 5). This result indicated that any number of other perturbations in the ein2-5 mutant could account for the altered expression of genes such as the peroxidase and cytochrome P450 genes. Furthermore, hypoxia-induced H2O2 accumulation may be required for ethylene signaling and for modulating the transcription of hypoxia-inducible genes during hypoxic stress in Arabidopsis.
Hypoxic conditions that trigger H2O2 accumulation have been observed in plants (Baxter-Burrell et al. 2002; Vergara et al. 2012). Here, GUS expression exhibited a similar pattern after hypoxia or H2O2 treatment in ERF73/HRE1 pro -uidA transgenic plants. This result implies that ERF73/HRE1 is downstream of H2O2 during hypoxia. Interestingly, the histochemical analysis revealed GUS activity in the trichomes and guard cells after hypoxia and H2O2 treatments. In non-aquatic plants, guard cell responses to flooding have been demonstrated, and ROS production in guard cells is mediated redundantly by the NADPH oxidase subunits Rboh D and F (Kwak et al. 2003; Dat et al. 2004). Studies of trichomes suggest that these tissues contain sulfur- and glutathione (GSH)-dependent defense mechanisms against oxidative stress and for redox control (Gutierrez-Alcala et al. 2000; Harada et al. 2010). However, further research is required to determine whether ERF73/HRE1 is involved in stomatal or trichome regulatory mechanisms.
Taken together, our findings provide insight into the effects of H2O2 on the transcriptional responses of ERF73/HRE1 and ADH1 modulated via ethylene signaling during primary hypoxia signaling. This study may help us to clarify the cross-talk between H2O2 and ethylene, and determine how it is involved in regulating the ERF73/HRE1 network during hypoxic stress.
Abbreviations
- H2O2 :
-
Hydrogen peroxide
- ROS:
-
Reactive oxygen species
- ERF73/HRE1:
-
Ethylene-responsive factors73/hypoxia responsive1
- Ein2-5:
-
Ethylene insensitive 2-5
- GUS:
-
β-Glucuronidase
- DPI:
-
Diphenylene iodonium chloride
- PDC1:
-
Pyruvate decarboxylase1
- SUS1,4:
-
Sucrose synthase1,4
- LDH:
-
Lactate dehydrogenase
- ADH1:
-
Alcohol dehydrogenase 1
- ROP:
-
Rho-related GTPase proteins
- RT-PCR:
-
Reverse-transcriptase polymerase chain reaction
References
Arasimowicz-Jelonek M, Floryszak-Wieczorek J, Deckert J, Rucińska-Sobkowiak R, Gzyl J, Pawlak-Sprada S, Abramowski D, Jelonek T, Gwóźdź EA (2012) Nitric oxide implication in cadmiuminduced programmed cell death in roots and signaling response of yellow lupine plants. Plant Physiol Biochem 58:124–134
Bailey-Serres J, Fukao T, Gibbs DJ, Holdsworth MJ, Lee SC, Licausi F, Perata P, Voesenek LA, van Dongen JT (2012) Making sense of low oxygen sensing. Trends Plant Sci 17:129–138
Banti V, Mafessoni F, Loreti E, Alpi A, Perata P (2010) The heat-inducible transcription factor HsfA2 enhances anoxia tolerance in Arabidopsis. Plant Physiol 152:1471–1483
Baxter-Burrell A, Yang Z, Springer PS, Bailey-Serres J (2002) RopGAP4-dependent Rop GTPase rheostat control of Arabidopsis oxygen deprivation tolerance. Science 296:2026–2028
Dat JF, Capelli N, Folzer H, Bourgeade P, Badot PM (2004) Sensing and signalling during plant flooding. Plant Physiol Biochem 42:273–282
Gibbs DJ, Lee SC, Isa NM, Gramuglia S, Fukao T, Bassel GW, Correia CS, Corbineau F, Theodoulou FL, Bailey-Serres J, Holdsworth MJ (2011) Homeostatic response to hypoxia is regulated by the N-end rule pathway in plants. Nature 479:415–418
Gutierrez-Alcala G, Gotor C, Meyer AJ, Fricker M, Vega JM, Romero LC (2000) Glutathione biosynthesis in Arabidopsis trichome cells. Proc Natl Acad Sci USA 97:11108–11113
Harada E, Kim JA, Meyer AJ, Hell R, Clemens S, Choi YE (2010) Expression profiling of tobacco leaf trichomes identifies genes for biotic and abiotic stresses. Plant Cell Physiol 51:1627–1637
Hattori Y, Nagai K, Furukawa S, Song XJ, Kawano R, Sakakibara H, Wu J, Matsumoto T, Yoshimura A, Kitano H, Matsuoka M, Mori H, Ashikari M (2009) The ethylene response factors SNORKEL1 and SNORKEL2 allow rice to adapt to deep water. Nature 460:1026–1030
Jefferson RA (1989) The GUS reporter gene system. Nature 342:837–838
Kwak JM, Mori IC, Pei ZM, Leonhardt N, Torres MA, Dangl JL, Bloom RE, Bodde S, Jones JD, Schroeder JI (2003) NADPH oxidase AtrbohD and AtrbohF genes function in ROS-dependent ABA signaling in Arabidopsis. EMBO J 22:2623–2633
Licausi F, van Dongen JT, Giuntoli B, Novi G, Santaniello A, Geigenberger P, Perata P (2010) HRE1 and HRE2, two hypoxia-inducible ethylene response factors, affect anaerobic responses in Arabidopsis thaliana. Plant J 62:302–315
Licausi F, Kosmacz M, Weits DA, Giuntoli B, Giorgi FM, Voesenek LA, Perata P, van Dongen JT (2011) Oxygen sensing in plants is mediated by an N-end rule pathway for protein destabilization. Nature 479:419–422
Mittler R, Vanderauwera S, Gollery M, Van Breusegem F (2004) Reactive oxygen gene network of plants. Trends Plant Sci 9:490–498
Muhlenbock P, Plaszczyca M, Mellerowicz E, Karpinski S (2007) Lysigenous aerenchyma formation in Arabidopsis is controlled by LESION SIMULATING DISEASE1. Plant Cell 19:3819–3830
Neill SJ, Desikan R, Clarke A, Hurst RD, Hancock JT (2002) Hydrogen peroxide and nitric oxide as signalling molecules in plants. J Exp Bot 53:1237–1247
Ophir R, Pang X, Halaly T, Venkateswari J, Lavee S, Galbraith D, Or E (2009) Gene-expression profiling of grape bud response to two alternative dormancy-release stimuli expose possible links between impaired mitochondrial activity, hypoxia, ethylene–ABA interplay and cell enlargement. Plant Mol Biol 71:403–423
Pena-Castro JM, van Zanten M, Lee SC, Patel MR, Voesenek LA, Fukao T, Bailey-Serres J (2011) Expression of rice SUB1A and SUB1C transcription factors in Arabidopsis uncovers flowering inhibition as a submergence tolerance mechanism. Plant J 67:434–446
Peng HP, Lin TY, Wang NN, Shih MC (2005) Differential expression of genes encoding 1-aminocyclopropane-1-carboxylate synthase in Arabidopsis during hypoxia. Plant Mol Biol 58:15–25
Pucciariello C, Parlanti S, Banti V, Novi G, Perata P (2012) Reactive oxygen species-driven transcription in Arabidopsis under oxygen deprivation. Plant Physiol 159:184–196
Sagi M, Davydov O, Orazova S, Yesbergenova Z, Ophir R, Stratmann JW, Fluhr R (2004) Plant respiratory burst oxidase homologs impinge on wound responsiveness and development in Lycopersicon esculentum. Plant Cell 16:616–628
Steffens B, Sauter M (2005) Epidermal cell death in rice is regulated by ethylene, gibberellin, and abscisic acid. Plant Physiol 139:713–721
Steffens B, Sauter M (2009) Epidermal cell death in rice is confined to cells with a distinct molecular identity and is mediated by ethylene and H2O2 through an autoamplified signal pathway. Plant Cell 21:184–196
Steffens B, Geske T, Sauter M (2011) Aerenchyma formation in the rice stem and its promotion by H2O2. New Phytol 190:369–378
Valenzuela-Soto JH, Iruegas-Bocardo F, Martinez-Gallardo NA, Molina-Torres J, Gomez-Lim MA, Delano-Frier JP (2011) Transformed tobacco (Nicotiana tabacum) plants over-expressing a peroxisome proliferator-activated receptor gene from Xenopus laevis (xPPARα) show increased susceptibility to infection by virulent Pseudomonas syringae pathogens. Planta 233:507–521
Vergara R, Parada F, Rubio S, Perez FJ (2012) Hypoxia induces H2O2 production and activates antioxidant defence system in grapevine buds through mediation of H2O2 and ethylene. J Exp Bot 63:4123–4131
Wang H, Huang J, Liang X, Bi Y (2012) Involvement of hydrogen peroxide, calcium, and ethylene in the induction of the alternative pathway in chilling-stressed Arabidopsis callus. Planta 235:53–67
Xu K, Xu X, Fukao T, Canlas P, Maghirang-Rodriguez R, Heuer S, Ismail AM, Bailey-Serres J, Ronald PC, Mackill DJ (2006) Sub1A is an ethylene-response-factor-like gene that confers submergence tolerance to rice. Nature 442:705–708
Yang CY, Hsu FC, Li JP, Wang NN, Shih MC (2011) The AP2/ERF transcription factor AtERF73/HRE1 modulates ethylene responses during hypoxia in Arabidopsis. Plant Physiol 156:202–212
Acknowledgments
The author thanks Dr. Ming-Che Shih (Agricultural Biotechnology Research Center, Academia Sinica, Taiwan) for the suggestions in this work.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yang, CY. Hydrogen peroxide controls transcriptional responses of ERF73/HRE1 and ADH1 via modulation of ethylene signaling during hypoxic stress. Planta 239, 877–885 (2014). https://doi.org/10.1007/s00425-013-2020-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-013-2020-z