Abstract
Key message
The weighted gene co-expression network analysis and antisense oligonucleotide-mediated transient gene silencing revealed that CsAAP6 plays an important role in amino acid transport during tea shoot development.
Abstract
Nitrogen transport from source to sink is crucial for tea shoot growth and quality formation. Amino acid represents the major transport form of reduced nitrogen in the phloem between source and sink, but the molecular mechanism of amino acid transport from source leaves to new shoots is not yet clear. Therefore, the composition of metabolites in phloem exudates collected by the EDTA-facilitated method was analyzed through widely targeted metabolomics. A total of 326 metabolites were identified in the phloem exudates with the richest variety of amino acids and their derivatives (93), accounting for approximately 39.13% of the total metabolites. Moreover, through targeted metabolomics, it was found that the content of glutamine, glutamic acid, and theanine was the most abundant, and gradually increased with the development of new shoots. Meanwhile, transcriptome analysis suggested that the expression of amino acid transport genes changed significantly. The WGCNA analysis identified that the expression levels of CsAVT1, CsLHTL8, and CsAAP6 genes located in the MEterquoise module were positively correlated with the content of amino acids such as glutamine, glutamic acid, and theanine in phloem exudates. Reducing the CsAAP6 in mature leaves resulted in a significant decrease in the content of glutamic acid, aspartic acid, alanine, leucine, asparagine, glutamine, and arginine in the phloem exudates, indicating that CsAAP6 played an important role in the source to sink transport of amino acids in the phloem. The research results will provide the theoretical basis and genetic resources for the improvement of nitrogen use efficiency and tea quality.
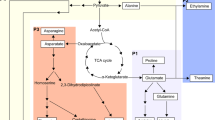
Graphical abstract








Similar content being viewed by others
Data availability
The datasets generated during and analyzed during the current study are available from the corresponding author on reasonable request.
References
Blicharz S, Beemster GTS, Ragni L, De Diego N, Spichal L, Hernandiz AE, Marczak L, Olszak M, Perlikowski D, Kosmala A, Malinowski R (2021) Phloem exudate metabolic content reflects the response to water-deficit stress in pea plants (Pisum sativum L.). Plant J 106:1338–1355. https://doi.org/10.1111/tpj.15240
Broussard L, Abadie C, Lalande J, Limami AM, Lothier J, Tcherkez G (2023) Phloem sap composition: what have we learnt from metabolomics? Int J Mol Sci 24:6917. https://doi.org/10.3390/ijms24086917
Chardon F, De Marco F, Marmagne A, Le Hir R, Vilaine F, Bellini C, Dinant S (2022) Natural variation in the long-distance transport of nutrients and photoassimilates in response to N availability. J Plant Physiol 273:153707. https://doi.org/10.1016/j.jplph.2022.153707
Dong C, Li F, Yang T, Feng L, Zhang S, Li F, Li W, Xu G, Bao S, Wan X, Lucas WJ, Zhang Z (2020) Theanine transporters identified in tea plants (Camellia sinensis L.). Plant J 101:57–70. https://doi.org/10.1111/tpj.14517
Doris R, Susanne S, Mechthild T (2007) Transporters for uptake and allocation of organic nitrogen compounds in plants. FEBS Lett 581:2281–2289. https://doi.org/10.1016/j.febslet.2007.04.013
Enrico M, Stefan M, Alexis DA, Réka N (2012) Vacuolar transporters in their physiological context. Annu Rev Plant Biol 63:183–213. https://doi.org/10.1146/annurev-arplant-042811-105608
Fan HF, Xu YL, Du CX, Wu X (2015) Phloem sap proteome studied by iTRAQ provides integrated insight into salinity response mechanisms in cucumber plants. J Proteomics 125:54–67. https://doi.org/10.1016/j.jprot.2015.05.001
Fan K, Zhang Q, Tang D, Shi Y, Ma L, Liu M, Ruan J (2020) Dynamics of nitrogen translocation from mature leaves to new shoots and related gene expression during spring shoots development in tea plants (Camellia sinensis L.). J Plant Nutr Soil Sci 183:180–191. https://doi.org/10.1002/jpln.201900268
Fischer WN, Loo DD, Koch W, Ludewig U, Boorer KJ, Tegeder M, Rentsch D, Wright EM, Frommer WB (2002) Low and high affinity amino acid H+-cotransporters for cellular import of neutral and charged amino acids. Plant J 29:717–731. https://doi.org/10.1046/j.1365-313X.2002.01248.x
Fraga CG, Clowers BH, Moore RJ, Zink EM (2010) Signature-discovery approach for sample matching of a nerve-agent precursor using liquid chromatography-mass spectrometry, XCMS, and chemometrics. Anal Chem 82:4165–4173. https://doi.org/10.1021/ac1003568
Fujiki Y, Teshima H, Kashiwao S, Kawano-Kawada M, Ohsumi Y, Kakinuma Y, Sekito T (2017) Functional identification of AtAVT3, a family of vacuolar amino acid transporters, in Arabidopsis. FEBS Lett 591:5–15. https://doi.org/10.1002/1873-3468.12507
Gourieroux AM, Holzapfel BP, Scollary GR, McCully ME, Canny MJ, Rogiers SY (2016) The amino acid distribution in rachis xylem sap and phloem exudate of Vitis vinifera ‘Cabernet Sauvignon’ bunches. Plant Physiol Biochem 105:45–54. https://doi.org/10.1016/j.plaphy.2016.04.010
Guo N, Gu M, Hu J, Qu H, Xu G (2020) Rice OsLHT1 functions in leaf-to-panicle nitrogen allocation for grain yield and quality. Front Plant Sci 11:1150. https://doi.org/10.3389/fpls.2020.01150
Hijaz F, Killiny N (2014) Collection and chemical composition of phloem sap from Citrus sinensis L. Osbeck (Sweet Orange). PLoS ONE 9:e101830. https://doi.org/10.1371/journal.pone.0101830
Hunt E, Gattolin S, Newbury HJ, Bale JS, Tseng HM, Barrett DA, Pritchard J (2010) A mutation in amino acid permease AAP6 reduces the amino acid content of the Arabidopsis sieve elements but leaves aphid herbivores unaffected. J Exp Bot 61:55–64. https://doi.org/10.1093/jxb/erp274
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360. https://doi.org/10.1038/nmeth.3317
King RW, Zeevaart JA (1974) Enhancement of Phloem exudation from cut petioles by chelating agents. Plant Physiol 53:96–103. https://doi.org/10.1104/pp.53.1.96
Kunio O, Shinya K, Kiyoshi M (1994) Remobilization of Nitrogen from vegetative parts to sprouting shoots of young tea (Camellia sinensis L.) plants. Jpn J Crop Sci 63:125–130. https://doi.org/10.1626/jcs.63.125
Li F, Dong C, Yang T, Bao S, Fang W, Lucas WJ, Zhang Z (2021) The tea plant CsLHT1 and CsLHT6 transporters take up amino acids, as a nitrogen source, from the soil of organic tea plantations. Horticult Res 8:178. https://doi.org/10.1038/s41438-021-00615-x
Li F, Lv C, Zou Z, Duan Y, Zhou J, Zhu X, Ma Y, Zhang Z, Fang W (2022) CsAAP7.2 is involved in the uptake of amino acids from soil and the long-distance transport of theanine in tea plants (Camellia sinensis L.). Tree Physiol 42:2369–2381. https://doi.org/10.1093/treephys/tpac071
Liang G, Hua Y, Chen H, Luo J, Xiang H, Song H, Zhang Z (2023) Increased nitrogen use efficiency via amino acid remobilization from source to sink organs in Brassica napus. Crop J 11:119–131. https://doi.org/10.1016/j.cj.2022.05.011
Liu XP, Gong CM, Fan YY, Eiblmeier M, Zhao Z, Han G, Rennenberg H (2013) Response pattern of amino compounds in phloem and xylem of trees to soil drought depends on drought intensity and root symbiosis. Plant Biol (Stuttg) 1:101–108. https://doi.org/10.1093/treephys/tpac071
Lough TJ, Lucas WJ (2006) Integrative plant biology: role of phloem long-distance macromolecular trafficking. Annu Rev Plant Biol 57:203–232. https://doi.org/10.1146/annurev.arplant.56.032604.144145
Lv YQ, Li D, Wu LY, Zhu YM, Ye Y, Zheng XQ, Lu JL, Liang YR, Li QS, Ye JH (2022) Sugar signal mediates flavonoid biosynthesis in tea leaves. Horticult Res 9:uhac049. https://doi.org/10.1093/hr/uhac049
Maillard A, Diquelou S, Billard V, Laine P, Garnica M, Prudent M, Garcia-Mina JM, Yvin JC, Ourry A (2015) Leaf mineral nutrient remobilization during leaf senescence and modulation by nutrient deficiency. Front Plant Sci 6:317. https://doi.org/10.3389/fpls.2015.00317
Martin A, Lee J, Kichey T, Gerentes D, Zivy M, Tatout C, Dubois F, Balliau T, Valot B, Davanture M, Terce-Laforgue T, Quillere I, Coque M, Gallais A, Gonzalez-Moro MB, Bethencourt L, Habash DZ, Lea PJ, Charcosset A, Perez P, Murigneux A, Sakakibara H, Edwards KJ, Hirel B (2006) Two cytosolic glutamine synthetase isoforms of maize are specifically involved in the control of grain production. Plant Cell 18:3252–3274. https://doi.org/10.1105/tpc.106.042689
Nam H, Gupta A, Nam H, Lee S, Cho HS, Park C, Park S, Park SJ, Hwang I (2022) JULGI-mediated increment in phloem transport capacity relates to fruit yield in tomato. Plant Biotechnol J 20:1533–1545. https://doi.org/10.1111/pbi.13831
Ruan J, Ma L, Yang Y (2012) Magnesium nutrition on accumulation and transport of amino acids in tea plants. J Sci Food Agric 92:1375–1383. https://doi.org/10.1002/jsfa.4709
Sakamoto S, Yoshikawa T, Teraishi M, Yoshinaga N, Ochiai K, Kobayashi M, Schmelz EA, Okumoto Y, Mori N (2022) A non-proteinogenic amino acid, β-tyrosine, accumulates in young rice leaves via long-distance phloem transport from mature leaves. Biosci Biotechnol Biochem 86:427–434. https://doi.org/10.1093/bbb/zbac012
Sevanto S (2014) Phloem transport and drought. Journal of Experimental Botany 65:1751-1759. https://doi.org/10.1093/jxb/ert467
Sevanto S (2018) Drought impacts on phloem transport. Curr Opin Plant Biol 43:76–81. https://doi.org/10.1016/j.pbi.2018.01.002
Taylor MR, Reinders A, Ward JM (2015) Transport Function of Rice Amino Acid Permeases (AAPs). Plant Cell Physiology 56:1355-1363. https://doi.org/10.1093/pcp/pcv053
Tegeder M, Masclaux-Daubresse C (2018) Source and sink mechanisms of nitrogen transport and use. New Phytol 217:35–53. https://doi.org/10.1111/nph.14876
Tegeder M, Ward JM (2012) Molecular evolution of plant AAP and LHT amino acid transporters. Front Plant Sci 3:21. https://doi.org/10.3389/fpls.2012.00021
Tegeder M (2014) Transporters involved in source to sink partitioning of amino acids and ureides: opportunities for crop improvement. Journal of Experimental Botany 65:1865–1878. https://doi.org/10.1093/jxb/eru012
Tetyuk O, Benning UF, Hoffmann-Benning S (2013) Collection and analysis of Arabidopsis phloem exudates using the EDTA-facilitated method. J vis Exp 80:e51111
Valim MF, Killiny N (2017) Occurrence of free fatty acids in the phloem sap of different citrus varieties. Plant Signal Behav 12:e1327497. https://doi.org/10.1080/15592324.2017.1327497
Wei C, Liang G, Zilong G, Wensheng W, Hongyan Z, Xianqing L, Sibin Y, Lizhong X, Jie L (2013) A novel integrated method for large-scale detection, identification, and quantification of widely targeted metabolites: application in the study of rice metabolomics. Mol Plant 6:1769–1780. https://doi.org/10.1093/mp/sst080
Xiong H, Ma H, Zhao H, Yang L, Hu B, Wang J, Shi X, Zhang Y, Rennenberg H (2022) Integrated physiological, proteome and gene expression analyses provide new insights into nitrogen remobilization in citrus trees. Tree Physiol 42:1628–1645. https://doi.org/10.1093/treephys/tpac024
Yao X, Nie J, Bai R, Sui X (2020) Amino acid transporters in plants: identification and function. Plants (basel) 9:973. https://doi.org/10.3390/plants9080972
Yesbergenova-Cuny Z, Dinant S, Martin-Magniette ML, Quillere I, Armengaud P, Monfalet P, Lea PJ, Hirel B (2016) Genetic variability of the phloem sap metabolite content of maize (Zea mays L.) during the kernel-filling period. Plant Sci 252:347–357. https://doi.org/10.1016/j.plantsci.2016.08.007
Yoshida M, Yoshida T, Yoshida-Takashima Y, Kashima A, Hiroishi S (2010) Real-time PCR detection of host-mediated cyanophage gene transcripts during infection of a natural Microcystis aeruginosa population. Microbes Environ 25:211–215. https://doi.org/10.1264/jsme2.ME10117
Zhang L, Tan Q, Lee R, Trethewy A, Lee YH, Tegeder M (2010) Altered xylem-phloem transfer of amino acids affects metabolism and leads to increased seed yield and oil content in Arabidopsis. Plant Cell 22:3603–3620. https://doi.org/10.1105/tpc.110.073833
Zhao M, Zhang N, Gao T, Jin J, Jing T, Wang J, Wu Y, Wan X, Schwab W, Song C (2020) Sesquiterpene glucosylation mediated by glucosyltransferase UGT91Q2 is involved in the modulation of cold stress tolerance in tea plants. New Phytol 226:362–372. https://doi.org/10.1111/nph.16364
Funding
This work was financially supported by the Natural Science Foundation of China (32002087), the Natural Science Foundation of Shandong Province (ZR2020QC171), the Agricultural Variety Improvement Project of Shandong Province (2020LZGC010), the Livelihood Project of Qingdao City (22-3-7-xdny-5-nsh) and the School Fund Project of Qingdao Agricultural University (1120096).
Author information
Authors and Affiliations
Contributions
KF and ZD conceived and designed the experiment. JZ, KS, and KF performed most of the experiments and drafted the paper. YW provided the tea seedlings used in the experiment. WQ, LS, and JS reviewed and modified the language of the paper. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest with the contents of this article.
Additional information
Communicated by Li Tian.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, J., Sun, K., Wang, Y. et al. Integrated metabolomic and transcriptomic analyses reveal the molecular mechanism of amino acid transport between source and sink during tea shoot development. Plant Cell Rep 43, 28 (2024). https://doi.org/10.1007/s00299-023-03110-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00299-023-03110-w