Abstract
Key message
Nitrate-responsive transcriptomic, phenotypic and physiological analyses of rice RGA1 mutant revealed many novel RGA1-regulated genes/processes/traits related to nitrogen use efficiency, and provided robust genetic evidence of RGA1-regulation of NUE.
Abstract
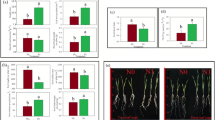
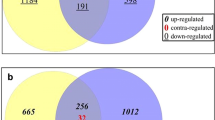
Nitrogen (N) use efficiency (NUE) is important for sustainable agriculture. G-protein signalling was implicated in N-response/NUE in rice, but needed firm genetic characterization of the role of alpha subunit (RGA1). The knock-out mutant of RGA1 in japonica rice exhibited lesser nitrate-dose sensitivity than the wild type (WT), in yield and NUE. We, therefore, investigated its genomewide nitrate-response relative to WT. It revealed 3416 differentially expressed genes (DEGs), including 719 associated with development, grain yield and phenotypic traits for NUE. The upregulated DEGs were related to photosynthesis, chlorophyll, tetrapyrrole and porphyrin biosynthesis, while the downregulated DEGs belonged to cellular protein metabolism and transport, small GTPase signalling, cell redox homeostasis, etc. We validated 26 nitrate-responsive DEGs across functional categories by RT-qPCR. Physiological validation of nitrate-response in the mutant and the WT at 1.5 and 15 mM doses revealed higher chlorophyll and stomatal length but decreased stomatal density, conductance and transpiration. The consequent increase in photosynthesis and water use efficiency may have contributed to better yield and NUE in the mutant, whereas the WT was N-dose sensitive. The mutant was not as N-dose-responsive as the WT in shoot/root growth, productive tillers and heading date, but equally responsive as WT in total N and protein content. The RGA1 mutant was less impacted by higher N-dose or salt stress in terms of yield, protein content, photosynthetic performance, relative water content, water use efficiency and catalase activity. PPI network analyses revealed known NUE-related proteins as RGA1 interactors. Therefore, RGA1 negatively regulates N-dose sensitivity and NUE in rice.









Similar content being viewed by others
Data availability/accession numbers
Our raw microarray data that support the findings of this study have been deposited in NCBI–GEO database (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE62164) with the accession number GSE62164 (GSM1520728, GSM1520729, GSM1520733, GSM1520732, GSM1520736 and GSM1520737). Additional transcriptome datasets were obtained from either the published supplementary materials and/or their authors cited in the article. All other datasets pertaining to the analyses are included in the supplementary information files.
Abbreviations
- NUE:
-
Nitrogen use efficiency
- N:
-
Nitrogen or nitrate
- WUE:
-
Water use efficiency
- PPI:
-
Protein–protein interaction
References
Abrol YP, Adhya TK, Aneja VP, Raghuram N, Pathak H, Kulshrestha U, Sharma C, Singh B (2017) The Indian nitrogen assessment: sources of reactive nitrogen, environmental and climate effects, management options, and policies. Elsevier, Amsterdam
Ali A, Sivakami S, Raghuram N (2007) Regulation of activity and transcript levels of NR in rice (Oryza sativa): roles of protein kinase and G-proteins. Plant Sci 172:406–413
Ashikari M, Wu J, Yano M, Sasaki T, Yoshimura A (1999) Rice gibberellin-insensitive dwarf mutant gene Dwarf 1 encodes the α-subunit of GTP-binding protein. Proc Natl Acad Sci 96:10284–10289
Bhatnagar N, Pandey S (2020) Heterotrimeric G-protein interactions are conserved despite regulatory element loss in some plants. Plant Physiol 184:1941–1954
Bloom AJ (2015) The increasing importance of distinguishing among plant nitrogen sources. Curr Opin Plant Biol 25:10–16
Boonburapong B, Buaboocha T (2007) Genome-wide identification and analyses of the rice calmodulin and related potential calcium sensor proteins. BMC Plant Biol 7:1–17
Cao W, Zhang H, Zhou Y, Zhao J, Lu S, Wang X, Chen X, Yuan L, Guan H, Wang G (2022) Suppressing chlorophyll degradation by silencing OsNYC3 improves rice resistance to Rhizoctonia solani, the causal agent of sheath blight. Plant Biotechnol. J. 20:335–349
Chakraborty N, Sharma P, Kanyuka K, Pathak RR, Choudhury D, Hooley R, Raghuram N (2015a) G-protein α-subunit (GPA1) regulates stress, nitrate and phosphate response, flavonoid biosynthesis, fruit/seed development and substantially shares GCR1 regulation in A. thaliana. Plant Mol Biol 89:559–576
Chakraborty N, Sharma P, Kanyuka K, Pathak RR, Choudhury D, Hooley RA, Raghuram N (2015b) Transcriptome analysis of Arabidopsis GCR1 mutant reveals its roles in stress, hormones, secondary metabolism and phosphate starvation. PLoS ONE 10:e0117819
Chakraborty N, Singh N, Kaur K, Raghuram N (2015c) G-protein signaling components GCR1 and GPA1 mediate responses to multiple abiotic stresses in Arabidopsis. Front Plant Sci 6:1000
Chakraborty N, Kanyuka K, Jaiswal DK, Kumar A, Arora V, Malik A, Gupta N, Hooley R, Raghuram N (2019) GCR1 and GPA1 coupling regulates nitrate, cell wall, immunity and light responses in Arabidopsis. Sci Rep 9:1–17
Choudhury SR, Pandey S (2016) Interaction of heterotrimeric G-protein components with receptor-like kinases in plants: an alternative to the established signaling paradigm? Mol Plant 9:1093–1095
Choudhury SR, Li M, Lee V, Nandety RS, Mysore KS, Pandey S (2020) Flexible functional interactions between G-protein subunits contribute to the specificity of plant responses. Plant J 102:207–221
Coneva V, Simopoulos C, Casaretto JA, El-Kereamy A, Guevara DR, Cohn J, Zhu T, Guo L, Alexander DC, Bi Y-M (2014) Metabolic and co-expression network-based analyses associated with nitrate response in rice. BMC Genomics 15:1–14
Chen M, Chen G, Di D, Kronzucker HJ, Shi W (2020) Higher nitrogen use efficiency (NUE) in hybrid “super rice” links to improved morphological and physiological traits in seedling roots. J. Plant Physiol. 251:153191
Chen Y, Liu Y, Ge J, Li R, Zhang R, Zhang Y, Huo Z, Xu K, Wei H, Dai Q (2022) Improved physiological and morphological traits of root synergistically enhanced salinity tolerance in rice under appropriate nitrogen application rate. Front Plant Sci 13:982637
Cui Y, Jiang N, Xu Z, Xu Q (2020) Heterotrimeric G protein are involved in the regulation of multiple agronomic traits and stress tolerance in rice. BMC Plant Biol 20:1–13
Damaris RN, Lin Z, Yang P, He D (2019) The rice alpha-amylase, conserved regulator of seed maturation and germination. Int. J. Mol. Sci. 20:450
Donaldson JG, Kahn RA, Lippincott-Schwartz J, Klausner RD (1991) Binding of ARF and β-COP to Golgi membranes: possible regulation by a trimeric G protein. Science 254:1197–1199
Ferrero-Serrano Á, Su Z, Assmann SM (2018) Illuminating the role of the Gα heterotrimeric G protein subunit, RGA1, in regulating photoprotection and photoavoidance in rice. Plant Cell Environ 41:451–468
Fredes I, Moreno S, Díaz FP, Gutiérrez RA (2019) Nitrate signaling and the control of Arabidopsis growth and development. Current Opin. Plant Biol. 47:112–118
Gao Y, Qi S, Wang Y (2022) Nitrate signaling and use efficiency in crops. Plant Commun. 100353
Gao L-L, Xue H-W (2012) Global analysis of expression profiles of rice receptor-like kinase genes. Mol Plant 5:143–153
Hoagland DR, Arnon DI (1950) The water-culture method for growing plants without soil, 2nd edn. Circular California Agricultural Experiment Station, p 347
Hooper CM, Castleden IR, Aryamanesh N, Jacoby RP, Millar AH (2016) Finding the subcellular location of barley, wheat, rice and maize proteins: the compendium of crop proteins with annotated locations (cropPAL). Plant Cell Physiol 57:e9–e9
Hou M, Yu M, Li Z, Ai Z, Chen J (2021) Molecular regulatory networks for improving nitrogen use efficiency in rice. Int J Mol Sci 22:9040
Jangam AP, Raghuram N (2015) Nitrogen and stress. In: Pandey G (ed) Elucidation of abiotic stress signaling in plants. Springer, New York, pp 323–339
Jangam AP, Pathak RR, Raghuram N (2016) Microarray analysis of rice d1 (RGA1) mutant reveals the potential role of G-protein alpha subunit in regulating multiple abiotic stresses such as drought, salinity, heat, and cold. Front Plant Sci 7:11
Jin J, Tian F, Yang D-C, Meng Y-Q, Kong L, Luo J, Gao G (2016) PlantTFDB 4.0: toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res. gkw982
Javed T, Indu I, Singhal RK, Shabbir R, Shah AN, Kumar P, Jinger D, Dharmappa PM, Shad MA, Saha D (2022) Recent advances in agronomic and physio-molecular approaches for improving nitrogen use efficiency in crop plants. Front. Plant Sci. 13
Joung J-G, Corbett AM, Fellman SM, Tieman DM, Klee HJ, Giovannoni JJ, Fei Z (2009) Plant MetGenMAP: an integrative analysis system for plant systems biology. Plant Physiol 151:1758–1768
Krouk G, Mirowski P, LeCun Y, Shasha DE, Coruzzi GM (2010) Predictive network modeling of the high-resolution dynamic plant transcriptome in response to nitrate. Genome Biol. 11:1–19
Kaur J, Roy Choudhury S, Vijayakumar A, Hovis L, Rhodes Z, Polzin R, Blumenthal D, Pandey S (2018) Arabidopsis type III Gγ Protein AGG3 is a positive regulator of yield and stress responses in the model monocot Setaria viridis. Front Plant Sci 9:109
Krapp A (2015) Plant nitrogen assimilation and its regulation: a complex puzzle with missing pieces. Curr Opin Plant Biol 25:115–122
Kronzucker HJ, Siddiqi MY, Glass AD, Kirk GJ (1999) Nitrate-ammonium synergism in rice. A subcellular flux analysis. Plant Physiol 119:1041–1046
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28:27–30
Kumari S, Sharma N, Raghuram N (2021) Meta-analysis of yield-related and N-responsive genes reveals chromosomal hotspots, key processes and candidate genes for nitrogen-use efficiency in rice. Front Plant Sci. https://doi.org/10.3389/fpls.2021.627955
Lichtenthaler H (1987) Chlorophyll and carotenoids–pigments of photosynthetic biomembrances za Colowick SP, Kaplan NO Methods in Enzymology. Vol. 148. In. Academic Press, San Diego
Li H, Hu B, Chu C (2017) Nitrogen use efficiency in crops: lessons from Arabidopsis and rice. J Exp Bot 68:2477–2488
Liang Y, Zhao X, Jones AM, Gao Y (2018) G proteins sculp root architecture in response to nitrogen in rice and Arabidopsis. Plant Sci 274:129–136
Loreto F, Velikova V (2001) Isoprene produced by leaves protects the photosynthetic apparatus against ozone damage, quenches ozone products, and reduces lipid peroxidation of cellular membranes. Plant Physiol. 127:1781–1787
Madan B, Malik A, Raghuram N (2022) Crop nitrogen use efficiency for sustainable food security and climate change mitigation. In: Kumar V, Srivastava AK, Suprasanna P (eds) Plant nutrition and food security in the era of climate change. Elsevier, Amsterdam, pp 47–72
Majumdar P, Torres Rodríguez MD, Pandey S (2023) Role of heterotrimeric G-proteins in improving abiotic stress tolerance of crop plants. J Plant Growth Regul. https://doi.org/10.1007/s00344-023-10965-6
Mandal VK, Sharma N, Raghuram N (2018) Molecular targets for improvement of crop nitrogen use efficiency: current and emerging options. In: Shrawat A, Zayed A, Lightfoot D (eds) Engineering nitrogen utilization in crop plants. Springer, Chamberland, pp 77–93
Mandal VK, Jangam AP, Chakraborty N, Raghuram N (2022) Nitrate-responsive transcriptome analysis reveals additional genes/processes and associated traits viz. height, tillering, heading date, stomatal density and yield in japonica rice. Planta 255:1–19
Maruta N, Trusov Y, Jones AM, Botella JR (2021) Heterotrimeric G proteins in plants: canonical and atypical Gα subunits. Int J Mol Sci 22:11841
Misyura M, Guevara D, Subedi S, Hudson D, McNicholas PD, Colasanti J, Rothstein SJ (2014) Nitrogen limitation and high density responses in rice suggest a role for ethylene under high density stress. BMC Genomics 15:1–14
Móring A, Hooda S, Raghuram N, Adhya TK, Ahmad A, Bandyopadhyay SK, Barsby T, Beig G, Bentley AR, Bhatia A (2021) Nitrogen challenges and opportunities for agricultural and environmental science in India. Front Sustain Food Syst. https://doi.org/10.3389/fsufs.2021.505347
Mudgil Y, Karve A, Teixeira PJ, Jiang K, Tunc-Ozdemir M, Jones AM (2016) Photosynthate regulation of the root system architecture mediated by the heterotrimeric G protein complex in Arabidopsis. Front Plant Sci 7:1255
Meng X, Wang X, Zhang Z, Xiong S, Wei Y, Guo J, Zhang J, Wang L, Ma X, Tegeder M (2021) Transcriptomic, proteomic, and physiological studies reveal key players in wheat nitrogen use efficiency under both high and low nitrogen supply. J. Exp. Bot. 72:4435–4456
Nazish T, Arshad M, Jan SU, Javaid A, Khan MH, Naeem MA, Baber M, Ali M (2021) Transporters and transcription factors gene families involved in improving nitrogen use efficiency (NUE) and assimilation in rice (Oryza sativa L.). Transgenic Res. 1-20
Neeraja CN, Barbadikar KM, Mangrauthia SK, Rao PR, Subrahmanayam D, Sundaram RM (2021) Genes for NUE in rice: a way forward for molecular breeding and genome editing. Plant Physiol Rep 26:587–599
O’Brien JA, Vega A, Bouguyon E, Krouk G, Gojon A, Coruzzi G, Gutiérrez RA (2016) Nitrate transport, sensing, and responses in plants. Mol Plant 9:837–856
Pandey S, Vijayakumar A (2018) Emerging themes in heterotrimeric G-protein signaling in plants. Plant Sci 270:292–300
Pathak RR, Ahmad A, Lochab S, Raghuram N (2008) Molecular physiology of plant nitrogen use efficiency and biotechnological options for its enhancement. Curr Sci 94:1394–1403
Pathak RR, Jangam AP, Malik A, Sharma N, Jaiswal DK, Raghuram N (2020) Transcriptomic and network analyses reveal distinct nitrate responses in light and dark in rice leaves (Oryza sativa Indica var. Panvel1). Sci Rep 10:1–17
Pathak RR, Mandal VK, Jangam AP, Sharma N, Madan B, Jaiswal DK, Raghuram N (2021) Heterotrimeric G-protein α subunit (RGA1) regulates tiller development, yield, cell wall, nitrogen response and biotic stress in rice. Sci Rep 11:1–19
Peng P, Gao Y, Li Z, Yu Y, Qin H, Guo Y, Huang R, Wang J (2019) Proteomic analysis of a rice mutant sd58 possessing a novel d1 allele of heterotrimeric G protein alpha subunit (RGA1) in salt stress with a focus on ROS scavenging. Int J Mol Sci 20:167
Prasad R, Shivay Y, Kumar D, Sharma S (2006) Learning by doing exercises in soil fertility (A practical manual for soil fertility). Division of Agronomy, IARI, New Delhi 68:
Qu C, Liu C, Gong X, Li C, Hong M, Wang L, Hong F (2012) Impairment of maize seedling photosynthesis caused by a combination of potassium deficiency and salt stress. Environ. Experim. Bot. 75:134–141
Raghuram N (2023) Reactive nitrogen in climate change, crop stress, and sustainable agriculture: a personal journey. In: Singh AK, Tuteja N (eds) Ansari MW. Global climate change and plant stress management, Wiley Online Library, pp 13–22
Raghuram N, Sharma N (2019) Improving crop nitrogen use efficiency. In: Moo-Young M (ed) Comprehensive biotechnology, 3rd edn. Elsevier, Pergamon
Raghuram N, Sopory SK (1999) Roles of nitrate, nitrite and ammonium ion in phytochrome regulation of nitrate reductase gene expression in maize. IUBMB Life 47:239–249
Raghuram N, Sutton MA, Jeffery R, Ramachandran R, Adhya TK (2021) From South Asia to the world: embracing the challenge of global sustainable nitrogen management. One Earth 4:22–27
Raghuram N, Aziz T, Kant S, Zhou J, Schmidt S (2022) Nitrogen use efficiency and sustainable nitrogen management in crop plants. Front Plant Sci 13:862091
Raudvere U, Kolberg L, Kuzmin I, Arak T, Adler P, Peterson H, Vilo J (2019) g: Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res 47:W191–W198
Sandhu N, Sethi M, Kumar A, Dang D, Singh J, Chhuneja P (2021) Biochemical and genetic approaches improving nitrogen use efficiency in cereal crops: a review. Front Plant Sci 12:657629
Sawaki N, Tsujimoto R, Shigyo M, Konishi M, Toki S, Fujiwara T, Yanagisawa S (2013) A nitrate-inducible GARP family gene encodes an auto-repressible transcriptional repressor in rice. Plant Cell Physiol 54:506–517
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
Sharma N, Sinha VB, Gupta N, Rajpal S, Kuchi S, Sitaramam V, Parsad R, Raghuram N (2018) Phenotyping for nitrogen use efficiency: rice genotypes differ in N-responsive germination, oxygen consumption, seed urease activities, root growth, crop duration, and yield at low N. Front Plant Sci 9:1452
Sharma N, Sinha VB, Prem Kumar NA, Subrahmanyam D, Neeraja C, Kuchi S, Jha A, Parsad R, Sitaramam V, Raghuram N (2021) Nitrogen use efficiency phenotype and associated genes: roles of germination, flowering, root/shoot length and biomass. Front Plant Sci 11:2329
Sharma N, Kumari S, Jaiswal DK, Raghuram N (2022) Comparative transcriptomic analyses of nitrate-response in rice genotypes with contrasting nitrogen use efficiency reveals common and genotype-specific processes, molecular targets and nitrogen use efficiency-candidates. Front Plant Sci 13:881204
Shin S-Y, Jeong JS, Lim JY, Kim T, Park JH, Kim J-K, Shin C (2018) Transcriptomic analyses of rice (Oryza sativa) genes and non-coding RNAs under nitrogen starvation using multiple omics technologies. BMC Genomics 19:1–20
Stateczny D, Oppenheimer J, Bommert P (2016) G protein signaling in plants: minus times minus equals plus. Curr Opin Plant Biol 34:127–135
Shanks CM, Huang J, Cheng C-Y, Shih H-J, Brooks M, Alvarez JM, Araus V, Swift J, Henry A, Coruzzi GM (2022) Validation of a high-confidence regulatory network for gene-to-NUE phenotype in field-grown rice. Front. Plant Sci. 4710
Sun H, Qian Q, Wu K, Luo J, Wang S, Zhang C, Ma Y, Liu Q, Huang X, Yuan Q (2014) Heterotrimeric G proteins regulate nitrogen-use efficiency in rice. Nat Genet 46:652–656
Supek F, Bošnjak M, Škunca N, Šmuc T (2011) REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6:e21800
Sutton M, Raghuram N, Adhya TK, Baron J, Cox C, de Vries W, Hicks K, Howard C, Ju X, Kanter D (2019) The nitrogen fix: from nitrogen cycle pollution to nitrogen circular economy-frontiers 2018/19: emerging issues of environmental concern chapter 4. In: Frontiers 2018/19: emerging issues of environmental concern
Swain DM, Sahoo RK, Srivastava VK, Tripathy BC, Tuteja R, Tuteja N (2017) Function of heterotrimeric G-protein γ subunit RGG1 in providing salinity stress tolerance in rice by elevating detoxification of ROS. Planta 245:367–383
The SV, Snyder R, Tegeder M (2021) Targeting nitrogen metabolism and transport processes to improve plant nitrogen use efficiency. Front Plant Sci 11:628366
Thimm O, Bläsing O, Gibon Y, Nagel A, Meyer S, Krüger P, Selbig J, Müller LA, Rhee SY, Stitt M (2004) MAPMAN: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J 37:914–939
Tian T, Liu Y, Yan H, You Q, Yi X, Du Z, Xu W, Su Z (2017) agriGO v2.0: a GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res 45:W122–W129
Tian W-j, Zhang X-q, Wang X-w, Xie J, Li Y-y, Sun Y, Tao Y-r, Xiong Y-z, Sang X-c (2018) Genetic mapping and salt tolerance of a novel D1-allelic mutant of rice (Oryza sativa L.). Acta Physiol Plant 40:1–10
Tiwari R, Bisht NC (2022) The multifaceted roles of heterotrimeric G-proteins: lessons from models and crops. Planta 255(4):88
Tseng KC, Li GZ, Hung YC, Chow CN, Wu NY, Chien YY, Zheng HQ, Lee TY, Kuo PL, Chang SB (2020) EXPath 2.0: an updated database for integrating high-throughput gene expression data with biological pathways. Plant Cell Physiol 61:1818–1827
Udvardi M, Below FE, Castellano MJ, Eagle AJ, Giller KE, Ladha JK, Liu X, Maaz TM, Nova-Franco B, Raghuram N (2021) A research road map for responsible use of agricultural nitrogen. Front Sustain Food Syst 5:660155
Urano D, Colaneri A, Jones AM (2014) Gα modulates salt-induced cellular senescence and cell division in rice and maize. J Exp Bot 65(22):6553–6561
Urano D, Leong R, Wu T-Y, Jones AM (2020) Quantitative morphological phenomics of rice G protein mutants portend autoimmunity. Dev Biol 457:83–90
Vidal EA, Alvarez JM, Araus V, Riveras E, Brooks MD, Krouk G, Ruffel S, Lejay L, Crawford NM, Coruzzi GM (2020) Nitrate in 2020: thirty years from transport to signaling networks. Plant Cell 32:2094–2119
Wang D, Qin B, Li X, Tang D, Ye Zhang, Cheng Z, Xue Y (2016) Nucleolar DEAD-box RNA helicase TOGR1 regulates thermotolerant growth as a pre-rRNA chaperone in rice. PLoS Genet. 12:e1005844
Wang R, Tischner R, Gutiérrez RA, Hoffman M, Xing X, Chen M, Coruzzi G, Crawford NM (2004) Genomic analysis of the nitrate response using a nitrate reductase-null mutant of Arabidopsis. Plant Physiol. 136:2512–2522
Wang K, Xu F, Yuan W, Sun L, Wang S, Aslam MM, Zhang J, Xu W (2021) G protein γ subunit qPE9-1 is involved in rice adaptation under elevated CO2 concentration by regulating leaf photosynthesis. Rice 14:1–10
Warpeha KM, Lateef SS, Lapik Y, Anderson M, Lee BS, Kaufman LS (2006) G-protein-coupled receptor 1, G-protein G α-subunit 1, and prephenate dehydratase 1 are required for blue light-induced production of phenylalanine in etiolated Arabidopsis. Plant Physiol 140:844–855
Warpeha KM, Upadhyay S, Yeh J, Adamiak J, Hawkins SI, Lapik YR, Anderson MB, Kaufman LS (2007) The GCR1, GPA1, PRN1, NF-Y signal chain mediates both blue light and abscisic acid responses in Arabidopsis. Plant Physiol 143:1590–1600
Xing J, Cao X, Zhang M, Wei X, Zhang J, Wan X (2022) Plant nitrogen availability and crosstalk with phytohormones signalings and their biotechnology breeding application in crops. Plant Biotechnol. J. 21:1320–1342
Xu G, Takahashi H (2020) Improving nitrogen use efficiency: From cells to plant systems. Oxford University Press, pp 4359–4364
Yadav DK, Shukla D, Tuteja N (2013) Rice heterotrimeric G-protein alpha subunit (RGA1): in silico analysis of the gene and promoter and its upregulation under abiotic stress. Plant Physiol Biochem 63:262–271
Yantong L, Ting L, Zhishu J, Chuihai Z, Rong H, Jiao Q, Xiaoli L, Limei P, Yongping S, Dahu Z, Yicong C, Changlan Z, Junru F, Haohua H, Jie X (2022) Characterization of a novel weak allele of RGA1/D1 and its potential application in rice breeding. Rice Sci 29(6):522–534
Yang X, Xia X, Zhang Z, Nong B, Zeng Y, Xiong F, Wu Y, Gao J, Deng G, Li D (2017) QTL mapping by whole genome re-sequencing and analysis of candidate genes for nitrogen use efficiency in rice. Front. Plant Sci. 8:1634
Yang SY, Hao DL, Song ZZ, Yang GZ, Wang L, Su YH (2015) RNA-Seq analysis of differentially expressed genes in rice under varied nitrogen supplies. Gene 555:305–317
Zait Y, Ferrero-Serrano Á, Assmann SM (2021) The α subunit of the heterotrimeric G protein regulates mesophyll CO2 conductance and drought tolerance in rice. New Phytol 232:2324–2338
Zhang L, Hu G, Cheng Y, Huang J (2008) Heterotrimeric G protein α and β subunits antagonistically modulate stomatal density in Arabidopsis thaliana. Dev Biol 324:68–75
Zhang H, Xie P, Xu X, Xie Q, Yu F (2021) Heterotrimeric G protein signalling in plant biotic and abiotic stress response. Plant Biol 23:20–30
Zhao Z, Stanley BA, Zhang W, Assmann SM (2010) ABA-regulated G protein signaling in Arabidopsis guard cells: a proteomic perspective. J Proteome Res 9:1637–1647
Zhu Y, Li T, Xu J, Wang J, Wang L, Zou W, Zeng D, Zhu L, Chen G, Hu J (2020) Leaf width gene LW5/D1 affects plant architecture and yield in rice by regulating nitrogen utilization efficiency. Plant Physiol Biochem 157:359–369
Acknowledgements
We thank Prof. T. Kumamaru from Kyushu University for providing the rice seeds and Regional Centre for Biotechnology (RCB), Faridabad for their help with the scanning electron microscopy.
Funding
This work was supported by research grants to NR from DST-SERB (F. No. CRG/2021/007467), ICAR (F. No. 2-2(60)/10-11/NICRA), Department of Biotechnology (DBT) [BT/IN/UK-VNC/44/NR/2015-16], UKRI GCRF South Asian Nitrogen Hub (SANH) [NE/S009019/1], GGSIPU[GGSIPU/DRC/Ph.D/Adm/2016/1549], [GGSIPU/DRC/FRGS/2018/22] and [GGSIPU/DRC/FRGS/2019/1553/24]. Fellowships were paid to VKM from DBT (DBT/JRF/14/AL/445) and GGSIPU (STRF: GGSIPU/DRC/2020/2049), APJ from CSIR (09/806(013)2008-EMR-I) and NC from UKRI GCRF-SANH [NE/S009019/1].
Author information
Authors and Affiliations
Contributions
JAP: planned the experiment, performed the microarray. VKM: performed the phenotypic, physiological, biochemical and qPCR validation. Performed part of in-silico analysis of microarray data. Assisted in writing and finalizing the draft of the manuscript. NC: performed most of the in-silico analysis of the microarray data, assisted in performing the phenotypic, physiological and qPCR validation. Wrote the initial draft of the manuscript. DK: Planned and supervised the experiments for total nitrogen and protein contents. NR: conceived, planned and supervised the transcriptome analyses, data interpretation, edited and finalized the manuscript. All authors have reviewed the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by Li Tian.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Prasanna, J.A., Mandal, V.K., Kumar, D. et al. Nitrate-responsive transcriptome analysis of rice RGA1 mutant reveals the role of G-protein alpha subunit in negative regulation of nitrogen-sensitivity and use efficiency. Plant Cell Rep 42, 1987–2010 (2023). https://doi.org/10.1007/s00299-023-03078-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-023-03078-7