Abstract
Solid organ transplantation (SOT) is the standard of care for end-stage organ disease. The most frequent complication of SOT involves allograft rejection, which may occur via T cell– and/or antibody-mediated mechanisms. Diagnosis of rejection in the clinical setting requires an invasive biopsy as there are currently no reliable biomarkers to detect rejection episodes. Likewise, it is virtually impossible to identify patients who exhibit operational tolerance and may be candidates for reduced or complete withdrawal of immunosuppression. Emerging single-cell technologies, including cytometry by time-of-flight (CyTOF), imaging mass cytometry, and single-cell RNA sequencing, represent a new opportunity for deep characterization of pathogenic immune populations involved in both allograft rejection and tolerance in clinical samples. These techniques enable examination of both individual cellular phenotypes and cell-to-cell interactions, ultimately providing new insights into the complex pathophysiology of allograft rejection. However, working with these large, highly dimensional datasets requires expertise in advanced data processing and analysis using computational biology techniques. Machine learning algorithms represent an optimal strategy to analyze and create predictive models using these complex datasets and will likely be essential for future clinical application of patient level results based on single-cell data. Herein, we review the existing literature on single-cell techniques in the context of SOT.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Solid organ transplantation (SOT) has revolutionized the treatment of end-stage organ disease. SOT, which includes kidney, liver, heart, lung, pancreas, and intestine allografts, has prolonged survival in patients with otherwise terminal diseases and improved overall quality of life [1]. The volume of SOT performed in the USA has been progressively increased since the first successful kidney transplants performed in the 1950s (Fig. 1a), with almost 25,000 kidneys and 9000 livers transplanted in 2021 (Fig. 1b). Advances in surgical techniques, immunosuppression protocols, donor selection, and expansion of donor pool to include living donors (for kidney and liver transplant) and deceased organ donation after both circulatory and brain death have led to a concurrent improvement in patient and graft survival.
Despite excellent outcomes, allograft rejection remains the most frequent complication and important potential cause of graft failure, morbidity, and mortality after SOT [2,3,4,5,6]. Based on histological features, rejection episodes are classified into three major clinicopathological entities: acute cellular rejection (ACR), antibody-mediated rejection (AMR), and chronic rejection. ACR is mediated by the activation of T cells recognizing donor human leucocyte antigen (HLA) peptides and represents the most frequent form of rejection, with an incidence that varies based on the organ transplanted, donor type and recipient age, and most frequently occurring within 6 months of transplantation [6,7,8,9,10,11,12]. Within the first year after SOT, pancreas transplant, alone or in combination with kidney, has a rate of rejection between 11.2 and 18% [7]. The incidence of ACR in lung transplant recipients varies from 17.1 to 28% based on data source, while the rate of rejection in heart transplant recipients is 20–25% for both children and adults [6, 8]. A higher percentage of ACR is reported in liver transplant recipients, where 25–60% of children and up to 25% of adults can experience at least one episode of rejection in the first-year post-transplant, while the rate of rejection in kidney transplant has been reported to range from 5 to 20% in recent years [9,10,11,12,13,14]. Prevention and treatment of rejection requires the lifelong burden of immunosuppression that is inevitably associated with increased risk of infection, de novo malignancy, and cardiovascular and chronic kidney disease and thus directly impacts long-term survival and quality of life [15, 16]. On the other hand, a subset of SOT recipients can retain stable allograft function while receiving reduced doses of immunosuppressive drugs or after being completely weaned off immunosuppression, achieving a state of “operational tolerance” [17,18,19]. Unfortunately, despite many attempts, there are no reliable biomarkers that can predict which patients may be candidates for immunosuppression minimization or withdrawal [20,21,22].
Regardless of the type of SOT, the precise molecular underpinnings of rejection are poorly understood, particularly in the clinical setting. Improving the understanding of pathogenic processes involved in rejection, such as the interplay between innate and adaptive immune responses, and the role of certain leukocyte subpopulations in mediating both acute and chronic allograft pathology, is crucial to identifying therapeutic targets. An additional benefit of deep characterization of alloimmune responses in transplantation is the possibility of identifying biomarkers of rejection. In liver and kidney transplantation, clinicians monitor allograft function using routine lab tests, such as serum creatinine for the kidney or serum aspartate transaminase (AST) and alanine transaminase (ALT) for the liver. While these tests can be elevated during rejection episodes, it has long been recognized that these tests offer low sensitivity and specificity for differentiating rejection episodes from other etiologies of graft dysfunction including infection or ischemia [23,24,25,26]. The diagnosis of rejection in other solid organs is notoriously challenging and generally requires frequent surveillance biopsy procedures. Optimal management of maintenance and rescue immunosuppression is critical for allograft longevity. Currently, the gold standard for diagnosis of rejection involves biopsy of the allograft, which is often a small core (16–18G) of tissue used for scoring by a pathologist. Because of potential heterogeneity of pathologic severity across within tissue, raising the possibility for sampling error, multiple biopsy passes are typically performed, with a minimum of five for lung and between two and six for heart allografts. Six to eight portal triads and five to ten glomeruli with two arteries are required to provide an adequate sample in the case of liver and kidney allografts, respectively [27,28,29,30,31]. Tissue sections are stained with hematoxylin and eosin (H&E) to assess the presence of perivascular and interstitial inflammatory infiltrates, damage and degeneration of parenchymal cells, and the degree of endothelial inflammation. Supporting stains for C4d (AMR) and other architectural stains such as trichrome (fibrosis) can provide additional diagnostic utility [32,33,34,35]. However, exploratory research studies of allografts employing molecular and cellular techniques are limited by scarce tissue availability. In this scenario, highly dimensional and single-cell technologies, such as mass cytometry and single-cell and single nucleus RNA sequencing (scRNA-seq, snRNA-seq), represent a promising path for identification of different cell lineages, including rare subset of cells, and in-depth phenotyping which would be otherwise obscured from whole-tissue analysis [36, 37]; they also provide information on cell cycle states, epigenetic changes, signaling pathway activation, and cytokine production [38,39,40]. Additionally, multiplexed protein imaging technologies, including but not limited to imaging mass cytometry (IMC), multiplexed ion beam imaging by time-of-flight (MIBI-TOF), co-detection by indexing (CODEX), and cyclic Immunofluorescence (CycIF) enable examination of spatial relationships between immune cells and tissue resident cells (Fig. 2) [41,42,43,44,45]. To overcome the loss of spatial information in single-cell techniques, several new approaches referred to as “spatial transcriptomics” have been developed to preserve functional tissue architecture and provide the ability to resolve groups of interacting cells [46]. A summary of the advantages and disadvantages of the above techniques is illustrated in Table 1.
Schematic illustrations of single-cell technologies. Peripheral blood mononuclear cells (PBMC) and cells in suspension can be analyzed by either platforms protein based (CyTOF) or nucleic acid based (sc-RNAseq, sn-RNAseq). FFPE or frozen tissue samples obtained from biopsy or explant can be analyzed using protein-based assays including IMC, CODEX, MIBI, or CycIF. Whole transcriptome can be obtained via meFISH, seqFISH, Visium 10X, and CosMx. Data from single-cell characterization are used for phenotyping and cluster identification using dimensionality reduction analysis such as PhenoGraph, t-SNE, as well as neighborhood and differential gene expression analyses. (Figure created with Biorender.com)
The aim of this article is to review the current literature regarding application of single-cell technologies to the important immunological challenges in SOT. Specifically, this review focuses on cytometry by time-of-flight (CyTOF) or mass cytometry, scRNA-seq/snRNA-seq, and imaging mass cytometry (IMC) as principal technologies adopted in SOT studies, as well as experiments involving peripheral blood as a surrogate of tissue biopsy findings. We also discuss potential application of machine learning algorithms to develop predictive models of clinical outcomes in the transplantation and immunology fields.
I. Use of blood as surrogate of biopsy sample: concordance of immune milieu in tissue and peripheral blood.
I.A. Conventional techniques for examining biomarkers in blood in SOT recipients
There are several situations where the clinical utility of biomarkers of rejection and tolerance can be applied in SOT [47]. First, biomarkers of rejection identified in the peripheral blood can avoid the need for an invasive biopsy and could also be performed more frequently, allowing for closer monitoring of graft function. Ideally, biomarkers would be predictive of a patient’s status prior to development of clinical signs of rejection, such as abnormalities in liver function tests so that therapy could be initiated prior to any substantial graft inflammatory insult. Moreover, biomarkers of alloimmunity offer the potential to guide clinical decision-making and determine which patients may be candidates for minimization or withdrawal from immunosuppressive therapy. In patients with operational tolerance, biomarkers are needed to monitor and confirm that the tolerogenic milieu has been maintained. Flow cytometry, microarrays, and RT-PCR are the traditional methods that studies have been employed to characterize immune phenotypes in peripheral blood and identify new biomarkers of rejection. Flow cytometry uses antibody-conjugated fluorophores to identify distinct, known cellular subpopulations based on surface markers. Most studies that use flow cytometry focus on either the T cell or B cell responses, and this technique is favored for identifying and comparing proportions of leukocyte subsets. Microarrays utilize a chip with a series of wells with pre-determined cDNA sequences, where tagged RNA enriched from samples is incubated and hybridizes to detect a vast assortment of pre-determined transcripts. This technique is most useful for transcriptome-level quantification and defining cellular pathways and phenotypes.
In kidney transplantation, flow cytometry was used to examine tolerant patients, and increased proportions of regulatory T cells were found in PBMCs [48, 49]. In liver transplant patients, an expansion of CD8 + CD28-T-suppressor subpopulations was associated with lower rates of acute or chronic rejection [50]. The understanding of the humoral immune response in transplantation using flow cytometry is more limited. In 2010, Pallier et al. found that operationally tolerant kidney transplant patients had increased levels of activated memory and early memory B cells [51]. This study used flow cytometry for identification and quantification of B cell subsets, followed by microarray to characterize the B cell gene expression profile. Others have observed that regulatory B cells and transitional B cells were increased in the peripheral blood of tolerant kidney transplant recipients [52].
The main limitation to multichannel flow cytometry is spectral overlap between fluorophores, which limits the number of immune markers that can be used in combination within a given experiment. Using a higher number of leukocyte markers increases the ability to distinguish unique and potentially important subpopulations. Even when adopting the maximal number of fluorophores in flow cytometry studies, the use of established combinations of markers may not uncover relevant heterogeneity among subpopulations and could lead to bias in detection of certain subsets. Spectral overlap generally requires supervised clustering, i.e., compensation and gating to determine which cells are positive or negative for a given marker to identify specific populations. Gating is subjective and relies on the knowledge of parameters such as cell size, whether cell sizes change under different conditions as well as marker expression, thus limiting the ability to identify unrecognized or novel populations. These limitations have limited the application of flow cytometry to define the detailed mechanisms of humoral or innate immune components involved in rejection or tolerance.
Multiparametric analysis of PBMCs has been performed less frequently in SOT. One study examined peripheral blood from liver transplant patients using a combination of flow cytometry, microarray, and reverse transcription-polymerase chain reaction (RT-PCR) techniques to identify increases in regulatory T cell subsets in operationally tolerant patients [48]. In a different study, microarray analysis was completed to identify 49 genes of operationally tolerant kidney transplant patients using peripheral blood [53]. These studies demonstrated that there was a link between the immune response in transplant patients and circulating peripheral blood mononuclear cells (PBMCs) that could be used to create a noninvasive diagnostic signature for tolerance.
I.B. CyTOF analysis of blood for identification of biomarkers of rejection
CyTOF allows for deeper immunophenotyping of circulating leukocytes because combinations of more than 40 surface markers can be assessed at the single-cell level without the complications of spectral overlap seen in flow cytometry. This limitation is overcome by using heavy metal–tagged antibodies that are detected using mass spectrometry with a very discrete signal, rather than fluorophores. The highly multiplexed capacity of mass cytometry enables the detection of intracellular signaling markers in conjunction with cell surface markers, which allows simultaneous identification of a lymphocyte subset and characterization of its phenotype. Moreover, mass cytometry enables simultaneous assessment of innate, humoral, and cytotoxic immune responses to provide a more complete characterization of the immune response in the setting of transplantation. These highly dimensional datasets are visualized using dimensionality reduction techniques including t-distributed scholastic embedding (tSNE), vi-SNE, or Uniform Manifold Approximation and Projection (UMAP) plots. While with vi-SNE/t-SNE cells are shown as a continuum of phenotypes, with cells colored based on the level of expression of chosen parameters and pairwise distances are transformed into probabilities based on t-distribution retaining local data structure, UMAP is able to retain both local and global data structure [54]. PhenoGraph or FlowSOM, which are graph-based unsupervised clustering algorithms, are then used to identify individual immune cell subpopulations, allowing the exploration of unanticipated phenotypes. Many investigators using CyTOF data also use manual gating strategies, similar to flow cytometry, to correlate clusters identified by PhenoGraph or FlowSOM with well-established immune subsets. FlowSOM utilizes hierarchical clustering with no down sample data; however it requires the number of clusters to be predetermined, which can be difficult in exploratory experiments, where an estimation of cell-type diversity is hardly available. PhenoGraph relies on nearest-neighbor connectivity model and can be potentially impacted by large sample size (Fig. 2) [54, 55]. Of note, the same algorithms are also adopted for the analysis of data generated via imaging mass cytometry and scRNA-sequencing. A summary of the markers used in mass cytometry studies is listed in Supplemental Table 1.
CyTOF was first applied to transplantation immunology to characterize the immune profile of operationally tolerant pediatric liver transplant recipients compared to their counterparts on low-dose immunosuppression [56]. This study used samples from seven operationally tolerant patients defined as patients with stable allograft function and no rejection episodes for more than 1 year after complete immunosuppression discontinuation and eight control patients who were maintained on low-dose single-agent immunosuppression. The authors identified a specific T cell subpopulation CD4+ CD5+ CD25+ CD38−/lo CD45RA that correlated with tolerance, confirming previous flow cytometry studies that suggested a role for CD4 + regulatory T cells in determining allograft tolerance. Another study examining 26 patients identified distinct subsets of exhausted T cells in peripheral blood that correlated with improved outcomes in kidney transplant recipients. Mass cytometry was performed to define specific CD4 + T cell exhaustion phenotypes. Five CD4 + T cell clusters were identified, CD4+PD-1+TIGIT+TIM3−2B4−, CD4+PD1+TIGI+TIM3+2B4lo, CD4+Th1 exhausted, CD4+Th2 exhausted, and CD4+PD1−4-1BB+TIM3+2B4+. The percentage of these subpopulations was increased six months post-transplant and associated with lower rates of interstitial fibrosis and an improved glomerular filtration rate. Lymphocyte subsets were also compared between patients who were receiving a lymphocyte-depleting regimen, anti-thymocyte globulin (ATG), and patients who received a non-depleting induction therapy. This comparison identified an increased proportion of monocytes following ATG therapy while T-regs increased over time in all subjects regardless of the type of induction [57]. Li et al. further evaluated therapeutic changes in immune responses by comparing peripheral blood immune profiles using mass cytometry in kidney transplant patients receiving tacrolimus or rapamycin regimens [58]. This 29-marker immune panel identified different T cell suppressive patterns and functional differences in TCR signaling in patients taking rapamycin versus those taking tacrolimus. The most comprehensive application of mass cytometry in renal transplant patients involved a 40-marker immune panel with markers for lymphocyte and myeloid lineages in 5 patients [59]. In this study, immune subtype analysis of CD4 + , CD8 + , NK cells, γδ T cells, DCs, and B cells revealed phenotypically distinct subclusters within each cell type. The study compared the relative abundance of these cell clusters in kidney transplant patients before and after receiving immunosuppressive therapy. ROC analysis for each immune cluster to examine the feasibility of using peripheral immune subpopulations revealed that CD4 + T cells, monocytes/macrophages, and NK cell and γδ T cells were the strongest predictors for efficacy of immune treatment (AUC = 0.96, 1 and 1, respectively).
These studies demonstrate that CyTOF can be used for deep phenotyping of peripheral blood and has the potential to identify immune cell subpopulations that may be evaluated as biomarkers in transplant. It is possible that CyTOF analysis of blood samples collected from transplant recipients can be used to develop more personalized immunosuppressive strategies and ameliorate adverse effects of too much or too little medication. However, mass cytometry does have limitations that are important to consider. Data acquisition can be more difficult when using mass cytometry compared to flow cytometry, as more cells are lost during the experimental procedure and the data acquisition rate is slower [60]. Additionally, many of these early studies have relatively small sample cohort sizes, preventing broad interpretation of key findings.
II. Single-cell analysis of tissue-based immune populations
II.A. Conventional techniques to examine immune cells in allograft tissue
Prior to the introduction of single-cell techniques, the only options for tissue characterization included immunohistochemistry, immunofluorescence, and confocal microscopy. When using these approaches, it can be very challenging to optimize the staining protocol, minimize background, and detect more than 3–4 markers simultaneously. As mentioned previously, most biopsies performed in SOT are needle cores, which usually only produce 5–8 discrete tissue sections for both clinical diagnosis and research use. Thus, detailed analysis of tissue architecture and single-cell phenotypes, especially in the research setting, has been very limited and a deep characterization of the single-cell immune landscape of rejection in SOT remains a largely unstudied research domain.
II.B. Imaging mass cytometry to examine immune cells in allograft tissue
Prior studies have adopted mass cytometry to examine individual cells obtained from allograft tissue. These first attempts involved tissue dissociation to obtain single-cell suspensions, red blood cell removal, and in some cases cell stimulation. This approach enables deep cellular phenotyping and extended characterization of functional state, without preserving tissue architecture nor the mutual cellular interactions [61,62,63,64]. More recently, the development of imaging mass cytometry (IMC) has offered a powerful tool that allows for detailed single-cell analysis within a tissue section (such as a biopsy section) while preserving spatial relationships. Formalin-fixed paraffin-embedded (FFPE) or frozen tissue sections are stained with a panel of up to 50 antibodies that have been conjugated to metal ions from the lanthanide series to reduce signal background. Tissue is then ablated with a UV laser, vaporized, and analyzed by time-of-flight mass spectrometer that quantifies each metal ion present in as little as 1 μm2 of tissue at 1000-nm resolution, which is fivefold higher than standard immunohistochemistry techniques. The isotope abundance is then used for image reconstruction and downstream analysis, including at the single-cell level (Fig. 2) [65]. Most IMC data has been reported using the Fluidigm hyperion system; however, another metal-based multiplex immunohistochemistry/immunofluorescence platform has been also developed. This is represented by multiplexed ion beam imaging by time-of-flight (MIBI-TOF), which uses oxygen-based primary ion beam for tissue ablation and metal isotopes are liberated from antibodies as secondary ions which are then delivered to a time-of-flight mass spectrometer for quantification of ion abundance [65]. An alternative to IMC involves DNA barcoding-based multiplexed tissue imaging technology called Co-detection by indexing (CODEX) which can detect > 50 antigens simultaneously. CODEX uses primary antibodies conjugated with oligonucleotides that are incubated to either fresh-frozen or FFPE tissue. Fluorophore-labeled oligonucleotides that are complementary to the oligonucleotide-conjugated primary antibodies are serially added and removed for image acquisition and reconstruction. When compared to MIBI and IMC, CODEX does not degrade or destroy the tissue specimen [67, 68]. Cyclic immunofluorescence (CycIF) is based on cyclic staining and imaging, with bleaching for signal removal and re-labeling for multiplexed imaging detection, enabling examination of up to 60 antigens [69, 70]. CycIF uses commercially available antibodies conjugated with fluorophores (e.g., Alexa Fluor Dyes) and image acquisition via conventional microscopy. CycIF has several variants; however, three-channel immunofluorescence combined with a fourth channel for DNA visualization is most commonly used. This multiplex technology provides information on cell morphology and protein translocation essential in studying the cytoskeleton, membrane receptor, and nuclear foci [70].
Thus far, most studies using IMC have focused on cancer immunobiology, inflammation, and autoimmune disease. Our group has recently published the first application of IMC in SOT, focusing on chronic rejection in liver transplantation [71]. Given the paucity of IMC data in SOT, we will review studies using IMC in SOT and in tissue samples obtained from organs that are used in SOT (Tables 2 and 3). The most frequent markers used in IMC studies is shown in Supplemental Table 1. The subcellular resolution of IMC allows localization of antigens within the cell (nucleus vs cytoplasm), and the large number of antibodies available that can be used simultaneously on a single section offers the ability to define cell type and phenotype, activation state, signaling pathway activation, cell-to-cell interaction, as well as proliferation state, features essential for understanding disease pathophysiology, and/or identification of novel biomarkers. For these reasons, IMC (and other multiplexed tissue analysis techniques) is ideally suited for the study of transplant immunology.
II.C Review of studies using IMC in SOT allografts
The first IMC application in the kidney transplantation was conducted by Avigan et al. to examine the molecular mechanism underlying delayed graft function (DGF) [72]. Preimplantation biopsy of 11 deceased donor kidney transplants (DDKT) at high risk for DGF were compared with 10 biopsies from living donor kidney transplants (LDKT). DDKT showed a significant reduction in tubular cells, mainly proximal tubule cells, and higher proportion of macrophage infiltration compared to LDKT. A subgroup analysis of the DDKT group showed that patients who ultimately developed DGF exhibited a reduction in overall tubular cell number, mostly in the connecting tubules/collecting ducts, with a compensatory increase of stromal cells when compared to high-risk patients with immediate graft function. This analysis using IMC also revealed the presence of a small cell population in the proximal tubule that was vimentin + , megalinlow, and co-expressing Ki67 and KIM-1, both typical markers of injury, in close association with macrophage-enriched infiltrates. These data suggested that preimplantation tubular cell dropout can play an important role in the development of DGF (Table 2) [72].
To our knowledge, our group has been the first to apply IMC to characterize rejection in SOT by investigating the alloimmunity landscape in patients with chronic rejection (CR) post-liver transplantation [71]. A workflow of the entire study is illustrated in Fig. 3. Using a 10-marker pilot IMC panel, a total of 92,791 cell from 18 CR (14 adult and 4 children) and 16,454 cells from 5 normal liver tissues without rejection (no rejection, NR) resulted in 29 meta-clusters, including 11 subpopulations with immune markers. More than 90% of the immune subpopulations identified were present in CR, which demonstrated a greater proportion of macrophages, cytotoxic T cells, and two other CD3 + T cell subpopulations when compared to NR. Interesting, a distinct population of CD68 + macrophages was completely absent, and CD66 + neutrophils were more prominent in NR. Spatial relationships examined using neighborhood analysis demonstrated only rare interactions among immune subpopulations in NR tissue, while strong interactions were observed in CR, particularly within the two CD68 + macrophage subpopulations and the CD3 + CD8 + cytotoxic T cell populations (Fig. 3). Strong interactions were also observed between macrophages and cytotoxic T cells and macrophages and other T and B cells, strongly correlating with the upregulation of the alloimmune response observed in the CR (Table 2). Principal component analysis (PCA) showed that CD20, CD3, CD45, CD66, CD8, and HLA-DR contributed similarly to principal component 1 (PC1), and explained almost 52% of the difference in cell distribution between NR and CR. Additionally, logistic regression model demonstrated that PC1 could predict CR with a probability above 75%(Fig. 3) [71].
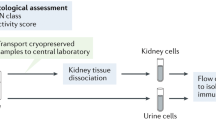
Schematic representation of IMC workflow for investigation of chronic rejection in liver transplantation recipients. Liver tissue sections were obtained from patients who underwent re-transplantation for chronic rejection (CR) of the primary graft. Pre-implantation liver biopsies from donors were used as control for liver with no rejection (NR). Staining was performed using a cocktail of 11 antibodies coupled with metal tags targeting immune cells. Tissue ablation and measurement of the metal ion abundance by time-of-flight mass spectrometry was performed using Hyperion Imaging System. Multidimensional images were segmented using Ilastik and Cell Profiler. t-SNE plots were used for dimensionality reduction to visualize level of expression of individual markers for each cell. PhenoGraph was used to establish immune meta-cluster identification. Cell subpopulation proportions were compared between chronic CR and NR and difference in macrophage and neutrophil proportions were observed between the two cohorts. Principal component and regression analyses were performed and revealed that PC1 had a high accuracy in rejection prediction. (Created with Biorender.com)
II.D. Review of studies using IMC in organs used for SOT
II.D.1 Kidney
The complex histopathologic organization of the human kidney makes IMC a well-suited technique to study cellular arrangement and interaction between parenchymal and immune cells with high resolution and essential spatial relationships. Using a 23-marker IMC and kidney-MAPPS (multiplexed antibody-based profiling with preservation of spatial context) pipeline on 16 histologically “normal” kidney samples, it was possible to recapitulate the cellular architecture of tubule, interstitium, and glomeruli of both renal cortex and medulla and identify 22 distinct cell populations, including a rare population of megalin + , aquaporin-1 + , and vimentin + cells that could represent an injured, fibrotic, or regenerative cell type, respectively [41]. Three tumor-remote specimens with immune infiltrate or interstitial nephritis visible on H&E were compared with healthy tissues, demonstrating fewer tubular, stromal, and endothelial cells as well as increase in immune cells mostly represented by CD4+ and CD8+ T cells and CD68+ macrophages [41]. Abundance of CD11b+ macrophages, CD11c+ dendritic cells, CD19+ B cells, and overproduction of TNFα were observed in single-cell analysis using IMC on kidney tissue from patients who died from SARS-CoV2 infection, suggesting a possible role of those cells in mediating multiorgan injuries, while the unequal B cell infiltration among various organs could be responsible for a different susceptibility and tissue-specific immune response to COVID-19 [73].
II.D.2 Liver
The liver is considered an immunologically privileged site, thus an attentive study of its sophisticated immune microenvironment remains essential to understand its response to chronic infection, tumor promoting/suppressing function, and tolerogenic state in transplantation. A 30-marker antibody panel was used to study patients with chronic HBV, showing greater immune cell enrichment, particularly CD4+ and CD8+ T cells, CD20+ B cells, and CD68+ Kupffer cells in the portal tracts of subjects with immune active disease compared to immunotolerant subjects [74]. Investigation of the functional state showed a greater activation (increased expression of HLA-DR, CD45RO, CD38) of immune cells in portal tracts of patients with immune active HBV infection. As well, both cell density and phenotype correlated with serum ALT level, inflammation, and fibrosis scores [74]. In a separate study examining the TME in hepatocellular carcinoma, spatial information gathered with IMC allowed identification of 22 meta-clusters and three distinct regions, with different distribution of stromal and immune cells [42]. Based on neighborhood analysis and identification of different interaction partners, Kupffer cells were hypothesized to be immunosuppressive and induce T cell exhaustion via PD-1-PD1L interaction, while infiltrating macrophages had stronger antigen presentation function, suggesting a tumor suppression role. Interestingly, the IMC data on cellular neighborhood correlated with patient prognosis, specifically that Kupffer cell enriched cellular neighborhoods were associated with worse patient and disease-free survival, while infiltrating macrophages were associated with prolonged survival. Thus, IMC analysis suggested that Kupffer cells may represent a potential target for hepatocellular carcinoma treatment [42].
II.D.3 Lung
The recent COVID-19 pandemic represented a further clinical setting where IMC has been adopted for a comprehensive examination of immune cells and cytokines involved in the infection response within the human lung. Using an IMC panel with 36 markers, Rendeiro et al. demonstrated that lungs from COVID-19 patients had significant macrophage infiltration, composed of predominantly interstitial macrophages, and a higher number of fibroblasts and mesenchymal cells were strongly correlated with increased lung fibrosis scores reported by the clinical pathologist. The observation of an augmented interaction between macrophages and fibroblasts or dendritic cells in the alveolar walls might represent an attempt to repair the tissue damage that results in fibrosis and thickening of the alveolar walls. The hyperinflammatory state as well as apoptosis initiation induced by SARS-CoV2 within alveolar epithelial cells was demonstrated by higher level of IL-6, CASP3 and C5b-C9 in infected cells compared to not infected cells [75]. In a multiorgan study by Wang et al. that included tissues from the lung, intestine, spleen, kidney, and liver obtained from patients with COVID-19 disease, the pulmonary immune infiltrates were constituted by CD11b+ macrophages producing IL-10 and CD11c+ dendritic cells, both characterized by low expression of HLA-DR. Thus, use of IMC highlights the pathophysiology and the dysregulated immune response observed in SARS-CoV2 infection, which may be the result of the activation of immunosuppressive pathway mediated by IL-10 and inhibition of HLA-DR activation on these two antigens presenting cell types, ultimately preventing the adaptive immune response [73].
IMC has also been used to investigate the heterogeneity of the tumor microenvironment (TME) in the study of both lung adenocarcinoma and squamous cell carcinoma, showing a strong interplay between cancer-associated fibroblasts and monocytic myeloid cells which were polarized to adopt a suppressor phenotype [76,77,78]. IMC also revealed recruitment and interaction of effector CD8+ and CD4+ T cells in the TME and their transition to a memory T cell phenotype, as well as the presence of CD33+ immunosuppressive cells. As well, a novel population of CD3−CD4+Foxp3+ cells negative for CD25 and CD127 with an unclear function was identified using IMC, which was hypothesized to have a proinflammatory role due to the high level of TNFα expression [78].
II.D.4. Pancreas
IMC has been used in two studies using human pancreata, aiming to better understand the pathophysiology of type 1 diabetes [79, 80]. Pseudotime analysis, an algorithm that can measure the progression of individuals along a certain biological process of interest based on a collection of high-dimensional molecular data, of 1,581 pancreatic islets was used by Damond et al., showing that patients with recent onset of diabetes had a near-normal fraction of β cells, although their phenotype was altered in the period preceding their autoimmune destruction [79]. More importantly, IMC analysis of spatial interactions between immune cells and islets showed that T-helper and cytotoxic T cells were more abundant at the onset stage, likely driving the immune-mediated destruction of islet cells. Conversely, these cells were less abundant in long duration diabetes, suggesting that T cell inflammation resolves after islet destruction [79]. Wang et al. reported progressive changes in the islet architecture, including reduction in vascular area and modification of the extracellular matrix, associated with ongoing reduction in the number of β cells until their complete disappearance. Analysis of 2,191 islets with a panel of 33 markers showed molecular and phenotype alterations in α cells, suggesting a possible role for transdifferentiation of α cells to β cells as a compensatory response to hyperglycemia. Echoing the previous study, enrichment in CD8+ T cells was found in those islets that still contained β cells, where upregulation of HLA-ABC on both endocrine and ductal cell types was associated with CD8+ T cell recruitment [80]. Analysis of more than 2 million cells from 13 nondiabetic and 16 type II diabetic donors using an IMC panel composed of 33 markers demonstrated a remodeling of the endocrine pancreas, with loss of β cells and simultaneous gain in α cells and accumulation of type I collagen [81]. Moreover, the intra-islet immune infiltration of patients with diabetes showed a twofold increase in HLA-DRhigh macrophages and HLA-DRhigh CD8 T cells when compared to healthy individuals, providing important insights in the phenotype of immune cells involved in type II diabetes [81].
III. Tissue and blood-based analysis in transplantation using single-cell RNA sequencing (scRNA-seq)
III.A. Single-cell RNA sequencing to examine immune cells in allograft tissue
Advances in deep sequencing have led to the development of bulk RNA seq, which enables tissue level characterization of the entire transcriptomes [82]. Pooled cell populations, typically comprising thousands of cells, can be evaluated to provide insights on contrasting gene expression signatures (differential expression) between experimental groups (Fig. 2). The first whole transcriptome single-cell analysis emerged in 2009 and since that time, a multitude of new scRNA-seq technologies have been developed [83]. Similar to other single-cell technologies, scRNA-seq techniques have led to rapid advances in our understanding of cellular phenotypes mediating disease processes that would be otherwise obscured by whole-tissue analyses. A major drawback is that studies involving scRNA-seq are expensive, however as more high-throughput technologies emerge, more cells can be analyzed with lower cost per cell. Ultimately, these techniques have further delineated novel cell subpopulations and signaling pathways mediated in disease processes and have already begun to show great promise in SOT (Table 4).
III.B. Review of studies using sc-RNA-seq in organs used for SOT
III.B.1. Liver
The first study on human liver transplantation and rejection has been recently published [84]. Li and colleagues analyzed 55 liver allograft biopsies, 7 livers (3 = health and 4 = liver transplantation), and 4 PBMC specimens. Liver biopsies obtained from transplanted patients, especially those with ACR, showed a higher percentage of exhausted CD8 T cells, suggesting that those cells may play a role during in the allograft tolerance. Additionally, LDLR + myeloid-derived suppressor cells (MDSC) expressing high level of TIGIT and Nectin2 were increased in liver transplantation tissues, especially in rejection tissues, suggesting their role in immune tolerance induction. Finally, the increased presence of CCR6+CD4+ T cell in the graft might suggest the presence of a higher proportion of memory T cells along with DC cells in the liver after transplantation. Interestingly, the author described the presence of a novel CD4+CD8+FOXP3+ T cell population in the context of liver transplantation [84].
For the study of the normal immune environment in liver, Zhao et al. used specimens from the liver, blood, and spleen from organ donors to represent “healthy human subjects” and isolated CD45 + immune cells within each and then performed scRNA-seq on over 70,000 immune cells [85]. With unsupervised clustering, they identified 30 discrete immune cell subpopulations across all sample types and given that scRNA-seq aims to characterize a cell’s entire transcriptome; this is likely a good representation of the true heterogeneity across immune cells in human. They also noted reproducibility across individual donors but important differences in the immune compartment across blood liver and spleen. The characterization of the specific cell subpopulations which were represented in the liver revealed novel heterogeneity among monocytes, NK and T cells, and B cells. Rocque et al. extracted this dataset as well as datasets from two additional high-quality scRNA-seq studies of human liver to increase the number of patients included and describe a meta-atlas of liver immune homeostasis [86,87,88]. This study comprising immune subsets within the liver from 17 unique human subjects described key pathways involved in liver immune homeostasis within subpopulation compartments such as upregulated apoptosis signaling in NK and T cells in the liver immune compartment relative to the peripheral blood immune compartment. These results provide a detailed account of the liver immune homeostasis which includes mechanisms of immunotolerance and are a branch points for the study of alloimmunity, immunosuppression, and rejection in SOT.
Yang and colleagues produced an experimental model of liver transplantation in rats where recipients either received fatty donor liver (FDL) which is more susceptible to ischemia–reperfusion injury or control donor liver (CDL) [89]. ScRNA-seq was used to identify a subset of inflammation-associated Kupffer cells which were characteristic of FCL, had a pro-inflammatory phenotype, and likely mediated ischemic injury following transplantation with FDL. Additional analysis of T cell subpopulations revealed that CCR7+ CD8+ T cells in FDL were noted to have a pro-apoptotic and pro-inflammatory phenotype. Further inflammatory pathways highlighted a role for multiple IL2 receptor subunits suggesting that IL2 might mediate this inflammation. The authors posited that early anti-IL2R induction therapy may be considered as an effective treatment for acute rejection in human patients who underwent transplantation with steatotic livers. This study again highlights how the deep characterization of immune profiles using scRNA-seq can generate clinical hypotheses that may lead to improvements in clinical treatment algorithms.
III.B.2. Kidney
To date, there is a larger body of literature employing scRNA-seq in kidney transplant immunology. The first study to emerge was in 2018 which described a patient with Banff 1B acute T cell–mediated rejection of their kidney allograft compared with a discarded donor kidney of a 70-year-old male with a serum creatinine of 1.1 [90]. Rejection appeared to be associated with the presence of a pro-inflammatory nonclassical monocyte subpopulation, and these cells expressed DC maturation markers suggesting a differentiation toward a DC fate. Characterization of epithelial cells showed pro-inflammatory gene signatures via TGFβ/BMP indicating that epithelial damage may further augment inflammatory signaling [90]. Another interesting study by Malone and colleagues took the analysis of the rejecting kidney transplant a step further to characterize leukocyte chimerism (i.e., roles of donor and recipient immune subsets) in patients with antibody-mediated rejection [91]. Whole exome sequencing of recipient and donor samples was used to identify single nucleotide polymorphisms between donor and recipient and to identify the origin of the cells in the scRNA-seq dataset. Interestingly, donor origin macrophages and lymphocytes comprised a major proportion of these cell subpopulations and donor-derived macrophages had a wound healing phenotype and did not appear to diminish linearly over time, but rather changed with rejection status. Recipient macrophages were enriched in rejection and had an activated phenotype. Ultimately, this study suggests that donor macrophages may play an important role in the response to alloimmune graft injury, and these complex and unique chimeric immune responses warrant further study to understand transplant immunology more fully.
When scRNA-seq was applied to chronic kidney transplant rejection (CKTR), it was found that CKTR had increases in mature and memory B cell markers and CKTR-associated monocytes appeared to have upregulation of inflammatory pathways [92]. Myofibroblasts were also expanded on the CKTR tissue relative to normal kidney and showed increases in activation, migration, and collagen expression which correlates with increased allograft fibrosis seen with CKTR.
III.B.3. Lung
The single-cell transcriptomics literature is limited across other types of SOT in humans and in animal models, but a study of macrophages and monocytes in sex mismatched lung transplant recipients identified that donor-derived airway macrophages are largely replaced by recipient-derived donor precursors [93]. Further studies are warranted to delineate how these cells that have the potential to mediate robust inflammatory responses are implicated in acute lung allograft rejection and/or bronchiolitis obliterans which pose major threats to allograft and patient survival. Most scRNA-seq studies in SOT have emerged within the past 4 years, and thus it is likely that more studies will soon uncover novel and important cell populations and pathways that mediate rejection which could lead to more sophisticated modeling of patient outcomes and the discovery of new or optimized therapeutic targets.
IV. Application of machine learning techniques to examine potential associations between single-cell signatures and clinical outcomes in SOT
Currently, predictive models for outcome trajectory and assessment of risk of rejection in transplant recipients are limited. The recent introduction of single-cell techniques that offer the potential to identify predictive biomarkers of rejection come with the challenge of analyzing large highly dimensional datasets which require additional skills in computational biology to analyze and model these data. The complexity of these data has necessitated development of new computational tools for dimensionality reduction, data normalization, visualization, cell population identification, and sample classification [94,95,96]. While most experiments using single-cell analysis have not been performed in a large cohort of patients, the granularity of the single-cell data can overcome a lower sample size and enable exploration of biomarkers and their potential association with clinical outcomes including post-transplant including delayed graft function, rejection, and infection. Advanced statistical approaches such as machine learning (ML) and regression analysis are well suited for these analyses. ML is already routinely used to enable visualization of high dimensional data in 2- or 3-dimensional graphs using dimensionality reduction techniques such as PCA, UMAP, and t-SNE for mass cytometry and scRNA-seq datasets [94, 95]. These tools facilitate data visualization, interpretation, and later statistical analyses. Unsupervised and supervised ML methods have also been developed for cell identification. While unsupervised methods such as PhenoGraph are considered unbiased, as they do not require previous model training, the cell population identity must be assigned by researchers based on marker expression, and it may be not possible to compare cellular phenotypes generated using this strategy across multiple datasets. Supervised clustering requires manual training of classifiers that can be subsequently applied to new data. While these models can be generalized to new datasets, they have implicit bias due to manual labeling of cells and are limited by pre-existing knowledge of certain cell types.
In terms of predictive modeling, regression analysis has traditionally been the “gold standard” for statistical modeling, but this approach is limited by the assumption of linear variable relationships. ML techniques allow for deeply complex data analysis on not only individual variable levels, but also on the nuanced relationships between any number of variables, including integration of single-cell data, relative to a specific outcome [97, 98]. It is clear that the application of ML is expanding, as illustrated by a recent review of these techniques in the field of liver transplantation [99].
ML algorithms have been widely tested to identify links between biomarkers and clinical outcomes in kidney transplantation [100,101,102,103]. Using recursive feature elimination (RFE), eight clinical explanatory features were identified as having a strong correlation with severe pneumonia in patients after DDKT. Those variables were used in combination with five different ML algorithms, and random forest displayed the best predictive performance, showing a specificity of 0.97, sensitivity of 0.67, and area under the receiver operating characteristic curve (AUROC) of 0.91 [102]. In another study, archetypal analysis models were applied to molecular profiles obtained from tissue microarray of kidney allograft biopsies to classify rejection episodes in six archetypes. For each biopsy, a score for each archetype was assigned, resulting in a final probability to belong to the highest scored archetype; however, there was a variable disagreement with the histological diagnosis [101]. Two separate studies examining rejection-associated transcripts previously identified in kidney biopsies explored archetypal analysis models demonstrating that it was possible to classify four different phenotypes of rejection in heart and lung transplant tissue [104, 105]. Although molecular classifiers could predict molecular diagnosis of rejection, discrepancy still remain in their correlation with the histological diagnosis of rejection [104, 105]. Lee et al. compared the predictive accuracy of multivariable logistic regression to ML models for the development of acute kidney injury following liver transplant [106]. This study showed that gradient boosting ML resulted in the best accuracy prediction for acute kidney injury and that random forest model performed better than logistic regression [106]. Wadhwani et al. utilized ML analyses to evaluate registry data for pediatric liver transplant recipients at 1-year post-transplant, to successfully identify favorable factors predisposing a patient to ideal outcome at 3 years, as well as predictive factors for non-ideal outcomes [107]. ML represents a useful tool for risk stratification and outcome prediction in SOT recipients and can ultimately assist clinicians in delivering more targeted therapies and interventions as well as optimize resource allocation.
V. Conclusions
In summary, by utilizing the increased dimensionality, CyTOF, scRNA-seq, and snRNA-seq applied to peripheral blood can help to identify the peripheral modification of the immune system during episode of SOT rejection. Analysis of the tissue allograft by IMC or RNA-seq offers a detailed description of the entire spectrum of immune cells involved in the rejection process within the transplanted organ and the relationship among different parties. Thus, the implementation of these techniques in the identification of rejection, immunotolerance, and biomarkers could transform transplant medicine. Not only would clinicians be able to use more data points than currently available, but they also could possibly limit the procedural risks associated with obtaining tissue biopsy samples. As demonstrated by Zamora et al., computational analysis of an array of biomarkers can identify markers of disease progression as well as possible therapeutic targets for future development [108]. By combining machine learning and single-cell data, promising predictive models could likely be generated which in turn, could lead to novel therapies, improve management strategies, and optimize patient outcomes.
References
Dean PG, Kukla A, Stegall MD, Kudva YC (2017) “Pancreas transplantation,” BMJ 357, https://doi.org/10.1136/BMJ.J1321
Jalalzadeh M, Mousavinasab N, Peyrovi S, Ghadiani MH (2015) The impact of acute rejection in kidney transplantation on long-term allograft and patient outcome. Nephrourol Mon 7(1):24439. https://doi.org/10.5812/NUMONTHLY.24439
Choudhary NS, Saigal S, Bansal RK, Saraf N, Gautam D, Soin AS (2017) Acute and chronic rejection after liver transplantation: what a clinician needs to know. J Clin Exp Hepatol 7(4):358–366. https://doi.org/10.1016/J.JCEH.2017.10.003
Patil DT, Yerian LM (2010) Pancreas transplant: recent advances and spectrum of features in pancreas allograft pathology. Adv Anat Pathol 17(3):202–208. https://doi.org/10.1097/PAP.0B013E3181D97635
Potena L, Zuckermann A, Barberini F, Aliabadi-Zuckermann A (2018) “Complications of cardiac transplantation,” Curr Cardiol Rep 20(9) https://doi.org/10.1007/S11886-018-1018-3
Parulekar AD, Kao CC (2019) Detection, classification, and management of rejection after lung transplantation. J Thorac Dis 11:S1732–S1739. https://doi.org/10.21037/JTD.2019.03.83
“Pancreas.” https://srtr.transplant.hrsa.gov/annual_reports/2019/Pancreas.aspx (accessed Feb. 03, 2022)
“Heart.” https://srtr.transplant.hrsa.gov/annual_reports/2019/Heart.aspx#HR_tx_ped_inc_AR_age_b64 (accessed Feb. 03, 2022).
Clayton PA, McDonald SP, Russ GR, Chadban SJ (2019) Long-term outcomes after acute rejection in kidney transplant recipients: an ANZDATA analysis. J Am Soc Nephrol 30(9):1697–1707. https://doi.org/10.1681/ASN.2018111101/-/
Cornell LD, Smith RN, Colvin RB (2008) Kidney transplantation: mechanisms of rejection and acceptance. Annu Rev Pathol Mech Dis 3:189–220. https://doi.org/10.1146/ANNUREV.PATHMECHDIS.3.121806.151508
Ng VL et al (2008) Outcomes of 5-year survivors of pediatric liver transplantation: report on 461 children from a North American multicenter registry. Pediatrics 122(6):e1128–e1135. https://doi.org/10.1542/peds.2008-1363
Levitsky J et al (2017) Acute rejection increases risk of graft failure and death in recent liver transplant recipients. Clin Gastroenterol Hepatol 15(4):584-593.e2. https://doi.org/10.1016/j.cgh.2016.07.035
Hart A et al (2017) OPTN/SRTR 2015 Annual Data Report: Kidney. Am J Transplant 17:21–116. https://doi.org/10.1111/AJT.14124
Kwong A et al “OPTN/SRTR 2018 Annual Data Report: Liver.”
Noble J, Terrec F, Malvezzi P, Rostaing L (2021) Adverse effects of immunosuppression after liver transplantation. Best Pract Res Clin Gastroenterol 54–55:101762. https://doi.org/10.1016/J.BPG.2021.101762
Claeys E, Vermeire K (2019) Immunosuppressive drugs in organ transplantation to prevent allograft rejection: mode of action and side effects. J Immunol Sci 3(4):14–21. https://doi.org/10.29245/2578-3009/2019/4.1178
Orlando G et al (2010) Clinical operational tolerance after renal transplantation: current status and future challenges. Ann Surg 252(6):915. https://doi.org/10.1097/SLA.0B013E3181F3EFB0
Massart A, Ghisdal L, Abramowicz M, Abramowicz D (2017) Operational tolerance in kidney transplantation and associated biomarkers. Clin Exp Immunol 189(2):138. https://doi.org/10.1111/CEI.12981
Feng S et al (2020) Efficacy and safety of immunosuppression withdrawal in pediatric liver transplant recipients: moving towards personalized management. https://doi.org/10.1002/hep.31520
Appenzeller-Herzog C, Hartleif S, Vionnet J (2021) Clinical parameters and biomarkers predicting spontaneous operational tolerance after liver transplantation: a scoping review. Am J Transplant 21(10):3312–3323. https://doi.org/10.1111/AJT.16585
Labarrere CA, Jaeger BR (2012) Biomarkers of heart transplant rejection: the good, the bad, and the ugly! Transl Res 159(4):238–251. https://doi.org/10.1016/J.TRSL.2012.01.018
Anglicheau D, Naesens M, Essig M, Gwinner W, Marquet P (2016) Establishing biomarkers in transplant medicine: a critical review of current approaches. Transplantation 100(10):2024–2038. https://doi.org/10.1097/TP.0000000000001321
Fedoravicius A, Charlton M (2016) Abnormal liver tests after liver transplantation. Clin Liver Dis 7(4):73–79. https://doi.org/10.1002/CLD.540
Santos J, Martins LS (2015) Estimating glomerular filtration rate in kidney transplantation: still searching for the best marker. World J Nephrol 4(3):345. https://doi.org/10.5527/WJN.V4.I3.345
Waikar SS, Betensky RA, Emerson SC, Bonventre JV (2012) Imperfect gold standards for kidney injury biomarker evaluation. J Am Soc Nephrol 23(1):13–21. https://doi.org/10.1681/ASN.2010111124
Abraham SC, Furth EE (1995) Receiver operating characteristic analysis of serum chemical parameters as tests of liver transplant rejection and correlation with histology. Transplantation 59(5):740–746. https://doi.org/10.1097/00007890-199503150-00018
Racusen LC (2006) Protocol transplant biopsies in kidney allografts: why and when are they indicated? Clin J Am Soc Nephrol 1(1):144–147. https://doi.org/10.2215/CJN.01010905
Van Ha TG (2004) Liver biopsy in liver transplant recipients. Semin Intervent Radiol 21(4):271. https://doi.org/10.1055/S-2004-861561
Strecker T, Rösch J, Weyand M, Agaimy A (2013) Endomyocardial biopsy for monitoring heart transplant patients: 11-years-experience at a German Heart Center. Int J Clin Exp Pathol 6(1):55. Available: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3515982/. Accessed 3 June 2022
Sakai K, Oguchi H, Muramatsu M, Shishido S (2018) Protocol graft biopsy in kidney transplantation. Nephrology 23:38–44. https://doi.org/10.1111/NEP.13282
Inaty H et al (2016) The optimal number of transbronchial biopsies needed to evaluate rejection in the lung allograft. J Hear Lung Transplant 35(4):S230. https://doi.org/10.1016/J.HEALUN.2016.01.654
Demetris AJ et al (2016) comprehensive update of the Banff working group on liver allograft pathology: introduction of antibody-mediated rejection. Am J Transplant 16(10):2816–2835. https://doi.org/10.1111/AJT.13909
Colvin MM et al (2015) Antibody-mediated rejection in cardiac transplantation: emerging knowledge in diagnosis and management: a scientific statement from the American Heart Association: Endorsed by the international society for heart and lung transplantation. Circulation 131(18):1608–1639. https://doi.org/10.1161/CIR.0000000000000093
Stewart S et al (2007) Revision of the 1996 working formulation for the standardization of nomenclature in the diagnosis of lung rejection. J Hear Lung Transplant 26(12):1229–1242. https://doi.org/10.1016/J.HEALUN.2007.10.017
Roufosse C et al (2018) A 2018 reference guide to the Banff classification of renal allograft pathology. Transplantation 102(11):1795–1814. https://doi.org/10.1097/TP.0000000000002366
Simoni Y et al (2018) “Bystander CD8 + T cells are abundant and phenotypically distinct in human tumour infiltrates,” https://doi.org/10.1038/s41586-018-0130-2.
David BA et al (2016) Combination of mass cytometry and imaging analysis reveals origin, location, and functional repopulation of liver myeloid cells in mice. Gastroenterology 151(6):1176–1191. https://doi.org/10.1053/J.GASTRO.2016.08.024
Galli E et al (2019) GM-CSF and CXCR4 define a T helper cell signature in multiple sclerosis. Nat Med 25(8):1290–1300. https://doi.org/10.1038/s41591-019-0521-4
Behbehani GK, Bendall SC, Clutter MR, Fantl WJ, Nolan GP (2012) Single-cell mass cytometry adapted to measurements of the cell cycle. Cytom Part A 81A(7):552–566. https://doi.org/10.1002/CYTO.A.22075
Bodenmiller B et al (2012) Multiplexed mass cytometry profiling of cellular states perturbed by small-molecule regulators. Nat Biotechnol 30(9):858–867. https://doi.org/10.1038/NBT.2317
Singh N et al (2019) Development of a 2-dimensional atlas of the human kidney with imaging mass cytometry. JCI Insight 4(12) https://doi.org/10.1172/JCI.INSIGHT.129477
Sheng J et al (2021) Topological analysis of hepatocellular carcinoma tumour microenvironment based on imaging mass cytometry reveals cellular neighbourhood regulated reversely by macrophages with different ontogeny. Gut. https://doi.org/10.1136/GUTJNL-2021-324339
Keren L et al (2019) MIBI-TOF: A multiplexed imaging platform relates cellular phenotypes and tissue structure. Sci Adv 5(10) https://doi.org/10.1126/SCIADV.AAX5851/SUPPL_FILE/AAX5851_TABLES_S1_TO_S3.XLSX
Phillips D et al (2021) Immune cell topography predicts response to PD-1 blockade in cutaneous T cell lymphoma. Nat Commun 12(1):1–18. https://doi.org/10.1038/s41467-021-26974-6
Lin JR et al (2018) Highly multiplexed immunofluorescence imaging of human tissues and tumors using t-CyCIF and conventional optical microscopes. Elife 7 https://doi.org/10.7554/ELIFE.31657.
Rao A, Barkley D, França GS, Yanai I (2021) Exploring tissue architecture using spatial transcriptomics. Nat 596(7871):211–220. https://doi.org/10.1038/s41586-021-03634-9
Vionnet J, Sánchez-Fueyo A (2018) Biomarkers of immune tolerance in liver transplantation. Hum Immunol 79(5):388–394. https://doi.org/10.1016/J.HUMIMM.2018.02.010
Martínez-Llordella M et al (2007) Multiparameter immune profiling of operational tolerance in liver transplantation. Am J Transplant 7(2):309–319. https://doi.org/10.1111/j.1600-6143.2006.01621.x
Braza F et al (2015) Central role of CD45RA- Foxp3hi memory regulatory T cells in clinical kidney transplantation tolerance. J Am Soc Nephrol 26(8):1795–1805. https://doi.org/10.1681/ASN.2014050480
Lin YX et al (2009) A significant expansion of CD8+ CD28- T-suppressor cells in adult-to-adult living donor liver transplant recipients. Transplant Proc 41(10):4229–4231. https://doi.org/10.1016/J.TRANSPROCEED.2009.09.072
Pallier A et al (2010) Patients with drug-free long-term graft function display increased numbers of peripheral B cells with a memory and inhibitory phenotype. Kidney Int 78(5):503–513. https://doi.org/10.1038/KI.2010.162
Nova-Lamperti E et al (2017) Increased CD40 ligation and reduced BCR signalling leads to higher IL-10 production in B-cells from tolerant kidney transplant patients. Transplantation 101(3):541. https://doi.org/10.1097/TP.0000000000001341
Brouard S et al (2007) Identification of a peripheral blood transcriptional biomarker panel associated with operational renal allograft tolerance. Proc Natl Acad Sci U S A 104(39):15448–15453. https://doi.org/10.1073/PNAS.0705834104
Kimball AK, Oko LM, Bullock BL, Nemenoff RA, van Dyk LF, Clambey ET (2018) A Beginner’s guide to analyzing and visualizing mass cytometry data. J Immunol 200(1):3–22. https://doi.org/10.4049/JIMMUNOL.1701494
Liu X et al A comparison framework and guideline of clustering methods for mass cytometry data. https://doi.org/10.1186/s13059-019-1917-7
Lau AH et al (2016) Mass cytometry reveals a distinct immunoprofile of operational tolerance in pediatric liver transplantation. Pediatr Transplant 20(8):1072–1080. https://doi.org/10.1111/PETR.12795
Fribourg M et al (2019) T-cell exhaustion correlates with improved outcomes in kidney transplant recipients. Kidney Int 96(2):436–449. https://doi.org/10.1016/J.KINT.2019.01.040
Li Y et al (2021) Deep phenotyping of T cell populations under long-term treatment of tacrolimus and rapamycin in patients receiving renal transplantations by mass cytometry. Clin Transl Med 11(11):e629. https://doi.org/10.1002/CTM2.629
Liu Y et al (2021) Single-cell profiling of kidney transplant recipients with immunosuppressive treatment reveals the dynamic immune characteristics. Front Immunol 12 https://doi.org/10.3389/FIMMU.2021.639942/FULL
Simoni Y, Chng MHY, Li S, Fehlings M, Newell EW (2018) Mass cytometry: a powerful tool for dissecting the immune landscape. Current Opinion in Immunology 51 Elsevier Ltd 187–196 https://doi.org/10.1016/j.coi.2018.03.023
Hanidziar D et al (2020) Characterization of pulmonary immune responses to hyperoxia by high-dimensional mass cytometry analyses. Sci Reports 10(1):1–10. https://doi.org/10.1038/s41598-020-61489-y
Chew V et al (2017) Delineation of an immunosuppressive gradient in hepatocellular carcinoma using high-dimensional proteomic and transcriptomic analyses. Proc Natl Acad Sci U S A 114(29):E5900–E5909. https://doi.org/10.1073/PNAS.1706559114/-/DCSUPPLEMENTAL
Wang YJ et al (2016) Single-cell mass cytometry analysis of the human endocrine pancreas. Cell Metab 24(4):616. https://doi.org/10.1016/J.CMET.2016.09.007
Zhang Q et al (2019) Integrated multiomic analysis reveals comprehensive tumour heterogeneity and novel immunophenotypic classification in hepatocellular carcinomas. Gut 68:2019–2031. https://doi.org/10.1136/gutjnl-2019-318912
Baharlou H, Canete NP, Cunningham AL, Harman AN, Patrick E (2019) Mass cytometry imaging for the study of human diseases—applications and data analysis strategies. Front Immunol 0:2657. https://doi.org/10.3389/FIMMU.2019.02657
Chattopadhyay PK, Gierahn TM, Roederer M, Love JC (2014) Single-cell technologies for monitoring immune systems. Nat Immunol 15(2):128–135. https://doi.org/10.1038/NI.2796
Black S et al (2021) CODEX multiplexed tissue imaging with DNA-conjugated antibodies. Nat Protoc 16(8):3802–3835. https://doi.org/10.1038/s41596-021-00556-8
Neumann EK et al (2022) Highly multiplexed immunofluorescence of the human kidney using co-detection by indexing. Kidney Int 101(1):137–143. https://doi.org/10.1016/J.KINT.2021.08.033
Lin J-R et al (2018) Highly multiplexed immunofluorescence imaging of human tissues and tumors using t-CyCIF and conventional optical microscopes. Elife 7 https://doi.org/10.7554/eLife.31657
Lin JR, Fallahi-Sichani M, Sorger PK (2015) Highly multiplexed imaging of single cells using a high-throughput cyclic immunofluorescence method. Nat Commun 6(1):1–7. https://doi.org/10.1038/ncomms9390
Ung N et al Adaptation of imaging mass cytometry to explore the single cell alloimmune landscape of liver transplant rejection. Front Immunol 0: 1363, 1AD, https://doi.org/10.3389/FIMMU.2022.831103
Avigan ZM, Singh N, Kliegel JA, Weiss M, Moeckel GW, Cantley LG (2021) Tubular cell dropout in preimplantation deceased donor biopsies as a predictor of delayed graft function. Transplant Direct. https://doi.org/10.1097/TXD.0000000000001168
Wang C et al (2020) Imaging mass cytometric analysis of postmortem tissues reveals dysregulated immune cell and cytokine responses in multiple organs of COVID-19 patients. Front Microbiol 11:3279. https://doi.org/10.3389/FMICB.2020.600989/BIBTEX
D. Traum et al (2021) Highly multiplexed 2-dimensional imaging mass cytometry analysis of HBV-infected liver. JCI Insight 6(7). https://doi.org/10.1172/JCI.INSIGHT.146883
Rendeiro AF et al (2021) The spatial landscape of lung pathology during COVID-19 progression. Nat 593(7860):564–569. https://doi.org/10.1038/s41586-021-03475-6
Flint LE et al (2020) Characterization of an aggregated three-dimensional cell culture model by multimodal mass spectrometry imaging. Anal Chem 92(18):12538–12547. https://doi.org/10.1021/ACS.ANALCHEM.0C02389/SUPPL_FILE/AC0C02389_LIVESLIDES.MP4
Xiang H et al (2020) Cancer-associated fibroblasts promote immunosuppression by inducing ROS-generating monocytic MDSCs in lung squamous cell carcinoma. Cancer Immunol Res 8(4):436–450. https://doi.org/10.1158/2326-6066.CIR-19-0507
Li R et al (2021) Characterization of the tumor immune microenvironment in lung squamous cell carcinoma using imaging mass cytometry. Front Oncol 11:1034. https://doi.org/10.3389/FONC.2021.620989/BIBTEX
Damond N, Engler S, Zanotelli VRT, Herrera PL, Atkinson MA, Correspondence BB (2019) A map of human type 1 diabetes progression by imaging mass cytometry. Cell Metab 29:755-768.e5. https://doi.org/10.1016/j.cmet.2018.11.014
Wang YJ et al (2019) Multiplexed in situ imaging mass cytometry analysis of the human endocrine pancreas and immune system in type 1 diabetes. Cell Metab 29(3):769-783.e4. https://doi.org/10.1016/j.cmet.2019.01.003
Wu M et al (2021) Single-cell analysis of the human pancreas in type 2 diabetes using multi-spectral imaging mass cytometry. Cell Rep 37(5):109919. https://doi.org/10.1016/J.CELREP.2021.109919
Wang Z, Gerstein M, Snyder M (2009) RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10(1):57–63. https://doi.org/10.1038/nrg2484
Tang F et al (2009) mRNA-Seq whole-transcriptome analysis of a single cell. Nat Methods 6(5):377–382. https://doi.org/10.1038/nmeth.1315
Li X et al (2022) Landscape of immune cells heterogeneity in liver transplantation by single-cell RNA sequencing analysis. Front Immunol 0:2065. https://doi.org/10.3389/FIMMU.2022.890019
Zhao J et al (2020) Single-cell RNA sequencing reveals the heterogeneity of liver-resident immune cells in human. Cell Discov. https://doi.org/10.1038/s41421-020-0157-z
Rocque B et al (2021) Creation of a single cell RNASeq meta-atlas to define human liver immune homeostasis. Front Immunol 12:679521. https://doi.org/10.3389/fimmu.2021.679521
MacParland SA et al (2018) Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat Commun 9(1):1–21. https://doi.org/10.1038/s41467-018-06318-7
Ramachandran P et al (2019) Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature 575(7783):512–518. https://doi.org/10.1038/s41586-019-1631-3
Yang X et al (2021) Single-cell profiling reveals distinct immune phenotypes that contribute to ischaemia-reperfusion injury after steatotic liver transplantation. Cell Prolif 54(10):e13116. https://doi.org/10.1111/cpr.13116
Wu H et al (2018) Single-cell transcriptomics of a human kidney allograft biopsy specimen defines a diverse inflammatory response. J Am Soc Nephrol 29(8):2069–2080. https://doi.org/10.1681/ASN.2018020125
Malone AF, Wu H, Fronick C, Fulton R, Gaut JP, Humphreys BD (2020) Harnessing expressed single nucleotide variation and single cell RNA sequencing to define immune cell chimerism in the rejecting kidney transplant. J Am Soc Nephrol 31(9):1977–1986. https://doi.org/10.1681/ASN.2020030326
Liu Y et al (2020) Single-cell analysis reveals immune landscape in kidneys of patients with chronic transplant rejection. Theranostics 10(19):8851
Byrne AJ et al (2020) Dynamics of human monocytes and airway macrophages during healthy aging and after transplant. J Exp Med 217(3). https://doi.org/10.1084/jem.20191236
Hu Z, Bhattacharya S, Butte AJ (2022) Application of machine learning for cytometry data. Front Immunol 0:5703. https://doi.org/10.3389/FIMMU.2021.787574
Krams SM, Schaffert S, Lau AH, Martinez OM (2017) Applying mass cytometry to the analysis of lymphoid populations in transplantation. Am J Transplant 17(8):1992–1999. https://doi.org/10.1111/AJT.14145
Aghaeepour N, Nikolic R, Hoos HH, Brinkman RR (2011) Rapid cell population identification in flow cytometry data. Cytom Part A 79A(1):6–13. https://doi.org/10.1002/CYTO.A.21007
Gaudillière B et al (2014) Clinical recovery from surgery correlates with single-cell immune signatures. Sci Transl Med 6(255). https://doi.org/10.1126/SCITRANSLMED.3009701
Rumer KK et al (2022) Integrated single-cell and plasma proteomic modeling to predict surgical site complications: a prospective cohort study. Ann Surg 275(3):582–590. https://doi.org/10.1097/SLA.0000000000005348
Tran J, Sharma D, Gotlieb N, Xu W, Bhat M (2022) Application of machine learning in liver transplantation: a review. Hepatol Int. https://doi.org/10.1007/S12072-021-10291-7
Reeve J et al (2019) Generating automated kidney transplant biopsy reports combining molecular measurements with ensembles of machine learning classifiers. Am J Transplant 19(10):2719–2731. https://doi.org/10.1111/AJT.15351
Reeve J et al (2017) Assessing rejection-related disease in kidney transplant biopsies based on archetypal analysis of molecular phenotypes. JCI Insight 2(12). https://doi.org/10.1172/JCI.INSIGHT.94197
Luo Y et al (2020) Machine learning for the prediction of severe pneumonia during posttransplant hospitalization in recipients of a deceased-donor kidney transplant. Ann Transl Med 8(4):82–82. https://doi.org/10.21037/ATM.2020.01.09
Costa SD et al (2020) The impact of deceased donor maintenance on delayed kidney allograft function: a machine learning analysis. PLoS One 15(2):e0228597. https://doi.org/10.1371/JOURNAL.PONE.0228597
Parkes MD et al (2019) An integrated molecular diagnostic report for heart transplant biopsies using an ensemble of diagnostic algorithms. J Hear Lung Transplant 38(6):636–646. https://doi.org/10.1016/J.HEALUN.2019.01.1318
Halloran KM et al (2019) Molecular assessment of rejection and injury in lung transplant biopsies. J Hear Lung Transplant 38(5):504–513. https://doi.org/10.1016/J.HEALUN.2019.01.1317/ATTACHMENT/591929E2-565F-4502-B504-E0A453B1995E/MMC3.DOCX
Lee HC et al (2018) Prediction of acute kidney injury after liver transplantation: machine learning approaches vs. logistic regression model. J Clin Med 7(11). https://doi.org/10.3390/JCM7110428
Wadhwani SI, Hsu EK, Shaffer ML, Anand R, Ng VL, Bucuvalas JC (2019) Predicting ideal outcome after pediatric liver transplantation: an exploratory study using machine learning analyses to leverage Studies of Pediatric Liver Transplantation Data. Pediatr Transplant 23(7). https://doi.org/10.1111/PETR.13554
Zamora R et al (2016) Data-driven modeling for precision medicine in pediatric acute liver failure. Mol Med 22:821. https://doi.org/10.2119/MOLMED.2016.00183
Funding
No funding was received to assist with the preparation of this manuscript. BR has received research support from a Broad Clinical Research Fellowship and One Legacy Fellowship Training Grant. JE was supported by a K08 from National Cancer Institute (K08CA245220), American Society of Transplant Surgeons Faculty Development Grant, Society for the Study of Liver Diseases, Clinical, Translational, and Outcomes Research Award, and Liver Scholar Award from Gilead Research Foundation. JAB is supported by Broad Clinical Research Fellowship.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is a contribution to the special issue on: Single-cell and spatial multi-omics in clinical outcomes studies - Guest Editor: Brice Gaudillière
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Barbetta, A., Rocque, B., Sarode, D. et al. Revisiting transplant immunology through the lens of single-cell technologies. Semin Immunopathol 45, 91–109 (2023). https://doi.org/10.1007/s00281-022-00958-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00281-022-00958-0