Abstract
Chitosan derivates with varying degrees of polymerization (DP) have attracted great concern due to their excellent biological activities. Increasing the abundance of chitosanases with different degradation modes contributes to revealing their catalytic mechanisms and facilitating the production of chitosan derivates. However, the identification of endo-chitosanases capable of producing chitobiose and D-glucosamine (GlcN) from chitosan substrates has remained elusive. Herein, an endo-chitosanase (CsnCA) belonging to the GH46 family was identified based on structural analysis in phylogenetic evolution. Moreover, we demonstrate that CsnCA acts in a random endo-acting manner, producing chitosan derivatives with DP ≤ 2. The in-depth analysis of CsnCA revealed that (GlcN)3 serves as the minimal substrate, undergoing cleavage in the mode that occupies the subsites − 2 to + 1, resulting in the release of GlcN. This study succeeded in discovering a chitosanase with distinctive degradation modes, which could facilitate the mechanistic understanding of chitosanases, further empowering the production of chitosan derivates with specific DP.
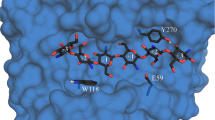
Graphical Abstract

Key points
• Structural docking and evolutionary analysis guide to mining the chitosanase.
• The endo-chitosanase exhibits a unique GlcN-producing cleavage pattern.
• The cleavage direction of chitosanase to produce GlcN was identified.





Similar content being viewed by others
Data availability
The data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Aktuganov GE, Galimzianova NF, Gilvanova EA, Pudova EA, Kuzmina LY, Melentiev AI, Safina VR (2019) Purification and characterization of exo-β-1,4-glucosaminidase produced by chitosan-degrading fungus, Penicillium sp. IB-37-2A. World J Microbiol Biotechnol 35:1–13
Ashkenazy H, Erez E, Martz E, Pupko T, Ben-Tal N (2010) ConSurf 2010: Calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res 38:529–533
Bhuvanachandra B, Sivaramakrishna D, Alim S, Preethiba G, Rambabu S, Swamy MJ, Podile AR (2021) New class of chitosanase from Bacillus amyloliquefaciens for the generation of chitooligosaccharides. J Agric Food Chem 69:78–87
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Chiang CL, Chang CT, Sung HY (2003) Purification and properties of chitosanase from a mutant of Bacillus subtilis IMR-NK1. Enzyme Microb Technol 32:260–267
Eide KB, Lindbom AR, Eijsink VGH, Norberg AL, Sørlie M (2013) Analysis of productive binding modes in the human chitotriosidase. FEBS Lett 587:3508–3513
Fukamizo T, Amano S, Yamaguchi K, Yoshikawa T, Katsumi T, Saito JI, Suzuki M, Miki K, Nagata Y, Ando A (2005) Bacillus circulans MH-K1 chitosanase: amino acid residues responsible for substrate binding. J Biochem 138:563–569
Gao L, Sun J, Secundo F, Gao X, Xue C, Mao X (2018) Cloning, characterization and substrate degradation mode of a novel chitinase from Streptomyces albolongus ATCC 27414. Food Chem 261:329–336
Guo N, Sun J, Wang W, Gao L, Liu J, Liu Z, Xue C, Mao X (2019) Cloning, expression and characterization of a novel chitosanase from Streptomyces albolongus ATCC 27414. Food Chem 286:696–702
Guo J, Wang Y, Zhang X, Gao W, Cai Z, Hong T, Man Z, Qing Q (2021) Improvement of the catalytic activity of chitosanase BsCsn46A from Bacillus subtilis by site-saturation mutagenesis of Proline121. J Agric Food Chem 69:11835–11846
Han Y, Guan F, Sun J, Wu N, Tian J (2020) Identification of a chitosanase from the marine metagenome and its molecular improvement based on evolution data. Appl Microbiol Biotechnol 104:6647–6657
Hekmat O, Lo Leggio L, Rosengren A, Kamarauskaite J, Kolenova K, Stålbrand H (2010) Rational engineering of mannosyl binding in the distal glycone subsites of Cellulomonas fimi endo-β-1, 4-mannanase: mannosyl binding promoted at subsite -2 and demoted at subsite -3. Biochemistry 49:4884–4896. https://doi.org/10.1021/bi100097f
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, Tunyasuvunakool K, Bates R, Žídek A, Potapenko A, Bridgland A, Meyer C, Kohl SAA, Ballard AJ, Cowie A, Romera-Paredes B, Nikolov S, Jain R, Adler J, Back T, Petersen S, Reiman D, Clancy E, Zielinski M, Steinegger M, Pacholska M, Berghammer T, Bodenstein S, Silver D, Vinyals O, Senior AW, Kavukcuoglu K, Kohli P, Hassabis D (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596:583–589
Kang LX, Chen XM, Fu L, Ma LX (2012) Recombinant expression of chitosanase from Bacillus subtilis HD145 in Pichia pastoris. Carbohydr Res 352:37–43
Liaqat F, Eltem R (2018) Chitooligosaccharides and their biological activities: a comprehensive review. Carbohydr Polym 184:243–259
Luo S, Qin Z, Chen Q, Fan L, Jiang L, Zhao L (2020) High level production of a Bacillus amlyoliquefaciens chitosanase in Pichia pastoris suitable for chitooligosaccharides preparation. Int J Biol Macromol 149:1034–1041
Ma C, Li X, Yang K, Li S (2020) Characterization of a new chitosanase from a marine Bacillus sp. and the anti-oxidant activity of its hydrolysate. Mar Drugs 18:1–13
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31:426–428
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791
Pechsrichuang P, Yoohat K, Yamabhai M (2013) Production of recombinant Bacillus subtilis chitosanase, suitable for biosynthesis of chitosan-oligosaccharides. Bioresour Technol 127:407–414
Pechsrichuang P, Lorentzen SB, Aam BB, Tuveng TR, Hamre AG, Eijsink VGH, Yamabhai M (2018) Bioconversion of chitosan into chito-oligosaccharides (CHOS) using family 46 chitosanase from Bacillus subtilis (BsCsn46A). Carbohydr Polym 186:420–428
Qin Z, Chen Q, Lin S, Luo S, Qiu Y, Zhao L (2018a) Expression and characterization of a novel cold-adapted chitosanase suitable for chitooligosaccharides controllable preparation. Food Chem 253:139–147. https://doi.org/10.1016/j.foodchem.2018.01.137
Qin Z, Luo S, Li Y, Chen Q, Qiu Y, Zhao L, Jiang L, Zhou J (2018b) Biochemical properties of a novel chitosanase from Bacillus amyloliquefaciens and its use in membrane reactor. Lwt- Food Sci Technol 97:9–16
Regel EK, Evers M, Liss M, Cord-Landwehr S, Moerschbacher BM (2020) High-throughput screening using UHPLC-MS to characterize the subsite specificities of chitosanases or chitinases. Anal Chem 92:3246–3252
Saito A, Ooya T, Miyatsuchi D, Fuchigami H, Terakado K, Nakayama S, Watanabe T, Nagata Y, Ando A (2009) Molecular characterization and antifungal activity of a family 46. FEMS Microbiol Lett 293:79–84
Seki K, Nishiyama Y, Mitsutomi M (2019) Characterization of a novel exo-chitosanase, an exo-chitobiohydrolase, from Gongronella butleri. J Biosci Bioeng 127:425–429
Singh R, Weikert T, Basa S, Moerschbacher BM (2019) Structural and biochemical insight into mode of action and subsite specificity of a chitosan degrading enzyme from Bacillus spec. MN Sci Rep 9:1–13
Su H, Sun J, Chu W, Yuan B, Mao X (2022a) Biochemical characterization and cleavage pattern analysis of a novel chitosanase with cellulase activity. Appl Microbiol Biotechnol 106:1979–1990
Su H, Sun J, Guo C, Jia Z, Mao X (2022b) New insights into bifunctional chitosanases with hydrolysis activity toward chito- and cello-substrates. J Agric Food Chem 70:6168–6176
Sun Y, Liu W, Han B, Zhang J, Liu B (2006) Purification and characterization of two types of chitosanase from a Microbacterium sp. Biotechnol Lett 28:1393–1399
Sun H, Cao R, Li L, Zhao L, Liu Q (2018a) Cloning, purification and characterization of a novel GH46 family chitosanase, Csn-CAP, from Staphylococcus capitis. Process Biochem 75:146–151
Sun H, Mao X, Guo N, Zhao L, Cao R, Liu Q (2018b) Discovery and characterization of a novel chitosanase from Paenibacillus dendritiformis by phylogeny-based enzymatic product specificity prediction. J Agric Food Chem 66:4645–4651
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461
Wang Y, Qin Z, Fan L, Zhao L (2020) Structure–function analysis of Gynuella sunshinyii chitosanase uncovers the mechanism of substrate binding in GH family 46 members. Int J Biol Macromol 165:2038–2048
Wang Y, Li D, Liu M, Xia C, Fan Q, Li X, Lan Z, Shi G, Dong W, Li Z, Cui Z (2021) Preparation of active chitooligosaccharides with a novel chitosanase AqCoA and their application in fungal disease protection. J Agric Food Chem 69:3351–3361
Wang J, Wang P, Zhu M, Chen W, Yu S, Zhong B (2022) Overexpression and biochemical properties of a GH46 chitosanase from marine Streptomyces hygroscopicus R1 suitable for chitosan oligosaccharides preparation. Front Microbiol 12:1–13
Watanabe T, Oyanagi W, Suzuki K, Tanaka H (1990) Chitinase system of Bacillus circulans WL-12 and importance of chitinase A1 in chitin degradation. J Bacteriol 172:4017–4022
Xie X, Ban X, Gu Z, Li C, Hong Y, Cheng L, Li Z (2020) Structure-based engineering of a maltooligosaccharide-forming amylase to enhance product specificity. J Agric Food Chem 68:838–844
Xu Y, Wang H, Zhu B, Yao Z (2023) Biochemical characterization and elucidation action mode of a new endolytic chitosanase for efficient preparation of chitosan oligosaccharides. Biomass Convers Biorefin. https://doi.org/10.1007/s13399-023-03870-1
Yang Y, Zheng Z, Xiao Y, Zhang J, Zhou Y, Li X, Li S, Yu H (2019) Cloning and characterization of a cold-adapted chitosanase from marine bacterium Bacillus sp. BY01. Molecules 24:1–11
Yuan X, Zheng J, Jiao S, Cheng G, Feng C, Du Y, Liu H (2019) A review on the preparation of chitosan oligosaccharides and application to human health, animal husbandry and agricultural production. Carbohydr Polym 220:60–70
Zhou J, Liu X, Yuan F, Deng B, Yu X (2020) Biocatalysis of heterogenously-expressed chitosanase for the preparation of desirable chitosan oligosaccharides applied against phytopathogenic fungi. ACS Sustain Chem Eng 8:4781–4791
Funding
This work was supported by the National Natural Science Foundation of China (U21A20271 and 31922072) and China Agriculture Research System of MOF and MARA (CARS-48).
Author information
Authors and Affiliations
Contributions
H.P.S and H.D conceived and designed the research. H.P.S and Y.Z.W conducted experiments. C.R.G contributed new reagents or analytical tools. H.P.S and J.S performed the literature search and analyzed the data. H.P.S and H.D wrote the manuscript. F.S reviewed and edited the manuscript. X.Z.M administrated the project and provided funds. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This article does not contain any studies with human participants or animals performed by any of the authors.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Su, H., Sun, J., Guo, C. et al. Structure-based mining of a chitosanase with distinctive degradation mode and product specificity. Appl Microbiol Biotechnol 107, 6859–6871 (2023). https://doi.org/10.1007/s00253-023-12741-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-023-12741-8