Abstract
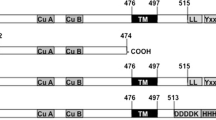
Current clinical laboratory assays are not sufficient for determining the activity of many specific human proteases yet. In this study, we developed a general approach that enables the determination of activities of caspase-3 based on the proteolytic activation of the engineered zymogen of the recombinant tyrosinase from Verrucomicrobium spinosum (Vs-tyrosinase) by detecting the diphenolase activity in an increase in absorbance at 475 nm. Here, we designed three different zymogen constructs of Vs-tyrosinase, including RSL-pre-pro-TYR, Pre-pro-TYR, and Pro-TYR. The active domain was fused to the reactive site loop (RSL) of α1-proteinase inhibitor and/or its own signal peptide (pre) and/or its own C-terminal domain (pro) via a linker containing a specific caspase-3 cleavage site. Further studies revealed that both RSL peptide and TAT signal peptide were able to inhibit tyrosinase diphenolase activity, in which RSL-pre-pro-TYR had the lowest background signals. Therefore, a specific protease activity such as caspase-3 could be detected when a suitable zymogen was established. Our results could provide a new way to directly detect the activities of key human proteases, for instance, to monitor the efficacy and safety of tumor therapy by determining the activity of apoptosis-related caspase-3 in patients.
Key points
• RSL inhibited the activity of Verrucomicrobium spinosum tyrosinase.
• N-pre and C-terminal domain exerted stronger dual inhibition on the Vs-tyrosinase.
• The activity of caspase-3 could be measured by the zymogen activation system.





Similar content being viewed by others

Data availability
The datasets generated and/or analyzed during this study are included in this manuscript (and its supplementary information file). Any additional data, if needed, will be available with the corresponding author upon reasonable request.
References
Agunbiade M, Le Roes-Hill M (2021) Application of bacterial tyrosinases in organic synthesis. World J Microbiol Biotechnol 38(1):2. https://doi.org/10.1007/s11274-021-03186-0
Al-Shobaili HA, Rasheed Z (2015) Oxidized tyrosinase: a possible antigenic stimulus for non-segmental vitiligo autoantibodies. J Dermatol Sci 79(3):203–213. https://doi.org/10.1016/j.jdermsci.2015.06.009
Axambayeva AS, Zhaparova LR, Shagyrova ZS, Ramankulov EM, Shustov AV (2018) Unusual stability of a recombinant Verrucomicrobium spinosum tyrosinase to denaturing agents and its use for a production of a protein with adhesive properties. Appl Biochem Biotechnol 185(3):736–754. https://doi.org/10.1007/s12010-017-2686-y
Buchanan G, Maillard J, Nabuurs SB, Richardson DJ, Palmer T, Sargent F (2008) Features of a twin-arginine signal peptide required for recognition by a Tat proofreading chaperone. FEBS Lett 582(29):3979–3984. https://doi.org/10.1016/j.febslet.2008.10.049
Claus H, Decker H (2006) Bacterial tyrosinases. Syst Appl Microbiol 29(1):3–14. https://doi.org/10.1016/j.syapm.2005.07.012
Decker H, Schweikardt T, Tuczek F (2006) The first crystal structure of tyrosinase: all questions answered? Angew Chem Int Ed Engl 45(28):4546–4550. https://doi.org/10.1002/anie.200601255
Djie MZ, Stone SR, Le Bonniec BF (1997) Intrinsic specificity of the reactive site loop of alpha1-antitrypsin, alpha1-antichymotrypsin, antithrombin III, and protease nexin I. J Biol Chem 272(26):16268–16273. https://doi.org/10.1074/jbc.272.26.16268
Espín JC, van Leeuwen J, Wichers HJ (1999) Kinetic study of the activation process of a latent mushroom (Agaricus bisporus) tyrosinase by serine proteases. J Agric Food Chem 47(9):3509–3517. https://doi.org/10.1021/jf9813539
Fairhead M, Thöny-Meyer L (2010) Role of the C-terminal extension in a bacterial tyrosinase. Febs j 277(9):2083–2095. https://doi.org/10.1111/j.1742-4658.2010.07621.x
Fairhead M, Thöny-Meyer L (2012) Bacterial tyrosinases: old enzymes with new relevance to biotechnology. N Biotechnol 29(2):183–191. https://doi.org/10.1016/j.nbt.2011.05.007
Fling M, Horowitz NH, Heinemann SF (1963) The isolation and properties of crystalline tyrosinase from Neurospora. J Biol Chem 238:2045–2053
Gurtu V, Kain SR, Zhang G (1997) Fluorometric and colorimetric detection of caspase activity associated with apoptosis. Anal Biochem 251(1):98–102. https://doi.org/10.1006/abio.1997.2220
Halaouli S, Asther M, Sigoillot JC, Hamdi M, Lomascolo A (2006) Fungal tyrosinases: new prospects in molecular characteristics, bioengineering and biotechnological applications. J Appl Microbiol 100(2):219–232. https://doi.org/10.1111/j.1365-2672.2006.02866.x
Inlow JK (2012) Homology models of four Agaricus bisporus tyrosinases. Int J Biol Macromol 50(1):283–293. https://doi.org/10.1016/j.ijbiomac.2011.11.010
Kanteev M, Goldfeder M, Fishman A (2015) Structure-function correlations in tyrosinases. Protein Sci 24(9):1360–1369. https://doi.org/10.1002/pro.2734
Lai X, Wichers HJ, Soler-Lopez M, Dijkstra BW (2018) Structure and function of human tyrosinase and tyrosinase-related proteins. Chemistry 24(1):47–55. https://doi.org/10.1002/chem.201704410
Mast AE, Enghild JJ, Salvesen G (1992) Conformation of the reactive site loop of alpha 1-proteinase inhibitor probed by limited proteolysis. Biochemistry 31(10):2720–2728. https://doi.org/10.1021/bi00125a012
Mauracher SG, Molitor C, Al-Oweini R, Kortz U, Rompel A (2014) Latent and active abPPO4 mushroom tyrosinase cocrystallized with hexatungstotellurate(VI) in a single crystal. Acta Crystallogr D Biol Crystallogr 70(Pt 9):2301–2315. https://doi.org/10.1107/s1399004714013777
Pan Y, Guo M, Nie Z, Huang Y, Peng Y, Liu A, Qing M, Yao S (2012) Colorimetric detection of apoptosis based on caspase-3 activity assay using unmodified gold nanoparticles. Chem Commun (camb) 48(7):997–999. https://doi.org/10.1039/c1cc15407a
Ramsden CA, Riley PA (2014) Tyrosinase: the four oxidation states of the active site and their relevance to enzymatic activation, oxidation and inactivation. Bioorg Med Chem 22(8):2388–2395. https://doi.org/10.1016/j.bmc.2014.02.048
Ren Q, Henes B, Fairhead M, Thöny-Meyer L (2013) High level production of tyrosinase in recombinant Escherichia coli. BMC Biotechnol 13:18. https://doi.org/10.1186/1472-6750-13-18
Sanrattana W, Sefiane T, Smits S, van Kleef ND, Fens MH, Lenting PJ, Maas C, de Maat S (2021) A reactive center loop-based prediction platform to enhance the design of therapeutic SERPINs. Proc Natl Acad Sci U S A 118(45). https://doi.org/10.1073/pnas.2108458118
Valanne A, Malmi P, Appelblom H, Niemelä P, Soukka T (2008) A dual-step fluorescence resonance energy transfer-based quenching assay for screening of caspase-3 inhibitors. Anal Biochem 375(1):71–81. https://doi.org/10.1016/j.ab.2007.12.032
Vuojola J, Riuttamäki T, Kulta E, Arppe R, Soukka T (2012) Fluorescence-quenching-based homogeneous caspase-3 activity assay using photon upconversion. Anal Chim Acta 725:67–73. https://doi.org/10.1016/j.aca.2012.03.010
Xavier Senra MV, Fonseca AL (2021) New tyrosinases with putative action against contaminants of emerging concern. Proteins 89(9):1180–1192. https://doi.org/10.1002/prot.26139
Xu DY, Yang Z (2013) Cross-linked tyrosinase aggregates for elimination of phenolic compounds from wastewater. Chemosphere 92(4):391–398. https://doi.org/10.1016/j.chemosphere.2012.12.076
Zhang LS, Xu HL, Xia Y, Bi JP, Zhang CZ, Xi Z, Li LY, Zhang ZS (2021) Real-time monitoring of caspase-3/8 activity by self-assembling nanofiber probes in living cells. Chem Commun (camb) 57(6):797–800. https://doi.org/10.1039/d0cc07821b
Acknowledgements
We are grateful to Professor Fei Xiao for his valuable suggestions on the experimental design. We are grateful to Professor Meng Wang for his valuable suggestions on the enzymatic activity analysis.
Funding
This study was funded by Beijing Hospital Project BJ-2020–134 and the National Natural Science Foundation of China (81871107).
Author information
Authors and Affiliations
Contributions
JZ and LZ conceived and designed research. JZ, WH, LZ, and XT conducted experiments. JZ and GS analyzed data. JZ and LZ wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, J., Huang, W., Zhang, L. et al. Modifying a bacterial tyrosinase zymogen for use in protease activity assays. Appl Microbiol Biotechnol 106, 8285–8294 (2022). https://doi.org/10.1007/s00253-022-12284-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-12284-4