Abstract
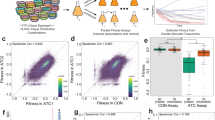
Phenotypic plasticity is the capacity to change the phenotype in response to different environments without alteration of the genotype. Despite sufficient evidence that microorganisms have a major role in the fitness and sickness of eukaryotes, there has been little research regarding microbial phenotypic plasticity. In this study, 45 strains of Staphylococcus aureus were grown for 12 days in both monoculture and in coculture with the same strain of Escherichia coli to create a competitive environment. Cell abundance was determined by quantitative PCR every 24 h, and growth curves of each S. aureus strain under the two sets of conditions were generated. Combined with whole-genome resequencing data, bivariate genome-wide association study (GWAS) was performed to analyze the growth plasticity of S. aureus in coculture. Finally, 20 significant single-nucleotide polymorphisms (eight annotated, seven unannotated, and five non-coding regions) were obtained, which may affect the competitive growth of S. aureus. This study advances genome-wide bacterial growth plasticity research and demonstrates the potential of bivariate GWAS for bacterial phenotypic plasticity research.
Key Points
• Growth plasticity of S. aureus was analyzed by bivariate GWAS.
• Twenty significant SNPs may affect the growth plasticity of S. aureus.





Similar content being viewed by others
References
Abu-Qatouseh LF, Chinni SV, Seggewiss J, Proctor RA, Brosius J, Rozhdestvensky TS, Peters G, von Eiff C, Becker K (2010) Identification of differentially expressed small non-protein-coding RNAs in Staphylococcus aureus displaying both the normal and the small-colony variant phenotype. J Mol Med (Berl) 88(6):565–575. https://doi.org/10.1007/s00109-010-0597-2
Abuin JM, Pichel JC, Pena TF, Amigo J (2016) SparkBWA: speeding up the alignment of high-throughput DNA sequencing data. PLoS One 11(5):e0155461. https://doi.org/10.1371/journal.pone.0155461
Anil R, Matthew S, Pritchard JK (2014) fastSTRUCTURE: variational inference of population structure in large SNP data sets. Genetics 197(2):573–589. https://doi.org/10.1534/genetics.114.164350
Bartoli C, Roux F (2017) Genome-wide association studies in plant pathosystems: toward an ecological genomics approach. Front Plant Sci 8:763. https://doi.org/10.3389/fpls.2017.00763
Beaudoin J, Ioannoni R, Labbe S (2012) Mfc1 is a novel copper transporter during meiosis. Commun Integr Biol 5(2):118–121. https://doi.org/10.4161/cib.18716
Beier S, Rivers AR, Moran MA, Obernosterer I (2015) Phenotypic plasticity in heterotrophic marine microbial communities in continuous cultures. ISME J 9(5):1141–1151. https://doi.org/10.1038/ismej.2014.206
Bowman L, Zeden MS, Schuster CF, Kaever V, Grundling A (2016) New insights into the cyclic di-adenosine monophosphate (c-di-AMP) degradation pathway and the requirement of the cyclic dinucleotide for acid stress resistance in Staphylococcus aureus. J Biol Chem 291(53):26970–26986. https://doi.org/10.1074/jbc.M116.747709
Camiolo S, Sablok G, Porceddu A (2016) Altools: a user friendly NGS data analyser. Biol Direct 11:8. https://doi.org/10.1186/s13062-016-0110-0
Charmantier A, McCleery RH, Cole LR, Perrins C, Kruuk LE, Sheldon BC (2008) Adaptive phenotypic plasticity in response to climate change in a wild bird population. Sci 320(5877):800–803. https://doi.org/10.1126/science.1157174
Chen R, Davydov EV, Sirota M, Butte AJ (2010) Non-synonymous and synonymous coding SNPs show similar likelihood and effect size of human disease association. PLoS One 5(10):e13574. https://doi.org/10.1371/journal.pone.0013574
Chevin LM, Lande R, Mace GM (2010) Adaptation, plasticity, and extinction in a changing environment: towards a predictive theory. PLoS Biol 8(4):e1000357. https://doi.org/10.1371/journal.pbio.1000357
Corrigan RM, Abbott JC, Burhenne H, Kaever V, Grundling A (2011) c-di-AMP is a new second messenger in Staphylococcus aureus with a role in controlling cell size and envelope stress. PLoS Path 7(9):e1002217. https://doi.org/10.1371/journal.ppat.1002217
Dicke M (2016) Plant phenotypic plasticity in the phytobiome: a volatile issue. Curr Opin Plant Biol 32:17–23. https://doi.org/10.1016/j.pbi.2016.05.004
Earle SG, Wu CH, Charlesworth J, Stoesser N, Gordon NC, Walker TM, Spencer CCA, Iqbal Z, Clifton DA, Hopkins KL, Woodford N, Smith EG, Ismail N, Llewelyn MJ, Peto TE, Crook DW, McVean G, Walker AS, Wilson DJ (2016) Identifying lineage effects when controlling for population structure improves power in bacterial association studies. Nat Microbiol 1:16041. https://doi.org/10.1038/nmicrobiol.2016.41
Emmons MF, Faiao-Flores F, Smalley KSM (2016) The role of phenotypic plasticity in the escape of cancer cells from targeted therapy. Biochem Pharmacol 122:1–9. https://doi.org/10.1016/j.bcp.2016.06.014
Feinberg AP (2007) Phenotypic plasticity and the epigenetics of human disease. Nature 447(7143):433–440. https://doi.org/10.1038/nature05919
Filice DCS, Long TAF (2017) Phenotypic plasticity in female mate choice behavior is mediated by an interaction of direct and indirect genetic effects in Drosophila melanogaster. Ecol Evol 7(10):3542–3551. https://doi.org/10.1002/ece3.2954
Fusco G, Minelli A (2010) Phenotypic plasticity in development and evolution: facts and concepts. Philos Trans R Soc Lond Ser B Biol Sci 365(1540):547–556. https://doi.org/10.1098/rstb.2009.0267
Goudey B, Rawlinson D, Wang Q, Shi F, Ferra H, Campbell RM, Stern L, Inouye MT, Ong CS, Kowalczyk A (2013) GWIS--model-free, fast and exhaustive search for epistatic interactions in case-control GWAS. BMC Genomics 14(Suppl 3):S10. https://doi.org/10.1186/1471-2164-14-s3-s10
Hayashi S, Wu HC (1990) Lipoproteins in bacteria. JBB 22(3):451–471. https://doi.org/10.1007/BF00763177
He X, Jin Y, Ye M, Chen N, Zhu J, Wang J, Jiang L, Wu R (2017) Bacterial genetic architecture of ecological interactions in co-culture by GWAS-taking Escherichia coli and Staphylococcus aureus as an example. Front Microbiol 8:2332. https://doi.org/10.3389/fmicb.2017.02332
Isidro-Sanchez J, Akdemir D, Montilla-Bascon G (2017) Genome-wide association analysis using R. Methods Mol Biol 1536:189–207. https://doi.org/10.1007/978-1-4939-6682-0_14
Kanaya S (2009) Ribonuclease H. FEBS J 276(6):1481. https://doi.org/10.1111/j.1742-4658.2009.06906.x
Kimchi-Sarfaty C, Oh JM, Kim IW, Sauna ZE, Calcagno AM, Ambudkar SV, Gottesman MM (2007) A “silent” polymorphism in the MDR1 gene changes substrate specificity. Sci 315(5811):525–528. https://doi.org/10.1126/science.1135308
Kummerli R, Jiricny N, Clarke LS, West SA, Griffin AS (2009) Phenotypic plasticity of a cooperative behaviour in bacteria. J Evol Biol 22(3):589–598. https://doi.org/10.1111/j.1420-9101.2008.01666.x
Lees JA, Vehkala M, Valimaki N, Harris SR, Chewapreecha C, Croucher NJ, Marttinen P, Davies MR, Steer AC, Tong SY, Honkela A, Parkhill J, Bentley SD, Corander J (2016) Sequence element enrichment analysis to determine the genetic basis of bacterial phenotypes. Nat Commun 7:12797. https://doi.org/10.1038/ncomms12797
Lele UN, Baig UI, Watve MG (2011) Phenotypic plasticity and effects of selection on cell division symmetry in Escherichia coli. PLoS One 6(1):e14516. https://doi.org/10.1371/journal.pone.0014516
Nadeau CP, Urban MC, Bridle JR (2017) Climates past, present, and yet-to-come shape climate change vulnerabilities. Trends Ecol Evol 32(10):786–800. https://doi.org/10.1016/j.tree.2017.07.012
Nowotny M, Gaidamakov SA, Ghirlando R, Cerritelli SM, Crouch RJ, Yang W (2007) Structure of human RNase H1 complexed with an RNA/DNA hybrid: insight into HIV reverse transcription. Mol Cell 28(2):264–276. https://doi.org/10.1016/j.molcel.2007.08.015
Okuda S, Tokuda H (2011) Lipoprotein sorting in bacteria. Annu Rev Microbiol 65(4):239–259. https://doi.org/10.1146/annurev-micro-090110-102859
Pigliucci M, Murren CJ, Schlichting CD (2006) Phenotypic plasticity and evolution by genetic assimilation. J Exp Biol 209(Pt 12):2362–2367. https://doi.org/10.1242/jeb.02070
Power RA, Parkhill J, Td O (2017) Microbial genome-wide association studies: lessons from human GWAS. Nat Rev Genet 18(1):41–50. https://doi.org/10.1038/nrg.2016.132
Prat C, Haas PJ, Bestebroer J, de Haas CJ, van Strijp JA, van Kessel KP (2009) A homolog of formyl peptide receptor-like 1 (FPRL1) inhibitor from Staphylococcus aureus (FPRL1 inhibitory protein) that inhibits FPRL1 and FPR. J Immunol 183(10):6569–6578. https://doi.org/10.4049/jimmunol.0801523
Ran S, Pei YF, Liu YJ, Zhang L, Han YY, Hai R, Tian Q, Lin Y, Yang TL, Guo YF, Shen H, Thethi IS, Zhu XZ, Deng HW (2013) Bivariate genome-wide association analyses identified genes with pleiotropic effects for femoral neck bone geometry and age at menarche. PLoS One 8(4):e60362. https://doi.org/10.1371/journal.pone.0060362
Rengefors K, Logares R, Laybourn-Parry J, Gast RJ (2015) Evidence of concurrent local adaptation and high phenotypic plasticity in a polar microeukaryote. Environ Microbiol 17(5):1510–1519. https://doi.org/10.1111/1462-2920.12571
Rhen T, Lang JW (1995) Phenotypic plasticity for growth in the common snapping turtle: effects of incubation temperature, clutch, and their interaction. Am Nat 146(5):726–747. https://doi.org/10.1086/285822
Robillard GT, Broos J (1999) Structure/function studies on the bacterial carbohydrate transporters, enzymes II, of the phosphoenolpyruvate-dependent phosphotransferase system. AcBB 1422(2):73–104. https://doi.org/10.1016/s0304-4157(99)00002-7
Rong M, Zheng X, Ye M, Bai J, Xie X, Jin Y, He X (2019) Phenotypic plasticity of Staphylococcus aureus in liquid medium containing vancomycin. Front Microbiol 10:809. https://doi.org/10.3389/fmicb.2019.00809
Saint-Pierre A, Kaufman JM, Ostertag A, Cohen-Solal M, Boland A, Toye K, Zelenika D, Lathrop M, de Vernejoul MC, Martinez M (2011) Bivariate association analysis in selected samples: application to a GWAS of two bone mineral density phenotypes in males with high or low BMD. Eur J Hum Genet 19(6):710–716. https://doi.org/10.1038/ejhg.2011.22
Santos JA, Pereira PJ, Macedo-Ribeiro S (2015) What a difference a cluster makes: the multifaceted roles of IscR in gene regulation and DNA recognition. AcBB 1854(9):1101–1112. https://doi.org/10.1016/j.bbapap.2015.01.010
Saxena R, Voight BF, Lyssenko V, Burtt NP, de Bakker PI, Chen H, Roix JJ, Kathiresan S, Hirschhorn JN, Daly MJ, Hughes TE, Groop L, Altshuler D, Almgren P, Florez JC, Meyer J, Ardlie K, Bengtsson Bostrom K, Isomaa B, Lettre G, Lindblad U, Lyon HN, Melander O, Newton-Cheh C, Nilsson P, Orho-Melander M, Rastam L, Speliotes EK, Taskinen MR, Tuomi T, Guiducci C, Berglund A, Carlson J, Gianniny L, Hackett R, Hall L, Holmkvist J, Laurila E, Sjogren M, Sterner M, Surti A, Svensson M, Svensson M, Tewhey R, Blumenstiel B, Parkin M, Defelice M, Barry R, Brodeur W, Camarata J, Chia N, Fava M, Gibbons J, Handsaker B, Healy C, Nguyen K, Gates C, Sougnez C, Gage D, Nizzari M, Gabriel SB, Chirn GW, Ma Q, Parikh H, Richardson D, Ricke D, Purcell S (2007) Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Sci 316(5829):1331–1336. https://doi.org/10.1126/science.1142358
Szurmant H, Mohan MA, Imus PM, Hoch JA (2007) YycH and YycI interact to regulate the essential YycFG two-component system in Bacillus subtilis. J Bacteriol 189(8):3280–3289. https://doi.org/10.1128/JB.01936-06
Turcotte MM, Levine JM (2016) Phenotypic plasticity and species coexistence. Trends Ecol Evol 31(10):803–813. https://doi.org/10.1016/j.tree.2016.07.013
Urban MC, Richardson JL, Freidenfelds NA (2014) Plasticity and genetic adaptation mediate amphibian and reptile responses to climate change. Evol Appl 7(1):88–103. https://doi.org/10.1111/eva.12114
Visscher PM, Brown MA, McCarthy MI, Yang J (2012) Five years of GWAS discovery. Am J Hum Genet 90(1):7–24. https://doi.org/10.1016/j.ajhg.2011.11.029
Williams WA, Zhang RG, Zhou M, Joachimiak G, Gornicki P, Missiakas D, Joachimiak A (2004) The membrane-associated lipoprotein-9 GmpC from Staphylococcus aureus binds the dipeptide GlyMet via side chain interactions. Biochemistry 43(51):16193–16202. https://doi.org/10.1021/bi048877o
Yan J, Nadell CD, Bassler BL (2017) Environmental fluctuation governs selection for plasticity in biofilm production.ISME. J 11(7):1569–1577. https://doi.org/10.1038/ismej.2017.33
Zhou X, Stephens M (2014) Efficient multivariate linear mixed model algorithms for genome-wide association studies. Nat Methods 11(4):407–409. https://doi.org/10.1038/nmeth.2848
Funding
This work was funded by the Fundamental Research Funds for the Central Universities (2017JC05, 2015ZCQ-SW-06), Beijing Municipal Funds for Talent Training (2017000020124G276), Science and technology foundation of Guizhou Province (NY[2012]3028), Natural Science Foundation of China (31971398, 31700633) and Science and Technology Service Network Initiative (KFJ-STS-ZDTP-036).
Author information
Authors and Affiliations
Contributions
Y.J. and X.H. conceived and designed the experiments. X.Z., J.B., and Y.L. performed the experiments. M.Y. and Y.J. analyzed the data. X.Z. and Y.J. wrote the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 6772 kb)
Rights and permissions
About this article
Cite this article
Zheng, X., Bai, J., Ye, M. et al. Bivariate genome-wide association study of the growth plasticity of Staphylococcus aureus in coculture with Escherichia coli. Appl Microbiol Biotechnol 104, 5437–5447 (2020). https://doi.org/10.1007/s00253-020-10636-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10636-6