Abstract
Polysialic acid (PSA) is a unique polysaccharide that plays critical roles in many bioprocesses, which makes it useful in a wide range of biomedical applications. The increased demand for PSA has led to considerable efforts to improve its production using bacteria, such as Escherichia coli. Bioprocess optimization and metabolic engineering have allowed the efficient production of PSA. This review aims to summarize the metabolism of PSA with an emphasis on the importance of the key pathway components. In addition, this review provides an update on state of the art PSA production using E. coli with a special emphasis on strategies of strain engineering and process development for the enhanced production of PSA.
Similar content being viewed by others
Introduction
Polysialic acid (PSA), a unique polymer of α-(2,8) and/or α-(2,9)-linked sialic acid (Neu5Ac) monomers, which is mainly attached onto glycoproteins, plays a major role in many biological processes (Maarouf et al. 2006; Ferrero and Aparicio 2010; Colley et al. 2014). Some bacterial pathogens possess a PSA capsule as a masking agent by mimicking the extracellular surface of mammalian cells (Muhlenhoff et al. 1998), such as Neisseria meningitidis (Bhattacharjee et al. 1975), Escherichia coli (Egan et al. 1977), Mannheimia hemolytica (Puente-Polledo et al. 1998). In mammals, PSA is involved in cell–cell and cell–matrix interactions, intermolecular interactions at cell surfaces, and interactions with other molecules in the cellular environment (Schnaar et al. 2014; Colley et al. 2014). PSA is present on the surface of numerous cells and contributes to the appropriate development, maintenance, and health of the nervous system (Sato and Kitajima 2013a, b; Schnaar et al. 2014; Colley et al. 2014). PSA accumulates on cancer cell surfaces in the later stages, which is associated with invasion and metastasis of cancer cells (Fukuda 1996; Falconer et al. 2012; Colley et al. 2014). PSA acts as an anti-adhesive molecule, plays important roles in normal mammalian organs and tissues, and promotes extraneural organ (e.g., lung, liver, testis, and placenta) development, repair, and regeneration (Stamatos et al. 2014; Ulm et al. 2013; Simon et al. 2013; Tsuchiya et al. 2014; Colley et al. 2014).
The various biological effects of PSA make it useful in a wide range of biomedical applications. PSA has been used in vaccine development to protect individuals from pathogenic meningitis (Tan et al. 2010; Colley et al. 2014). PSA can be a tumor-associated carbohydrate antigen to be used in cancer immunotherapy (Heimburg-Molinaro et al. 2011; Krug et al. 2012; Colley et al. 2014). Due to the poor immunogenicity, biodegradability, and biocompatibility of PSA, it has been proposed as the next generation for bioavailable products. Polysialylated enzymes are as effective as PEGylated enzymes in respect of prolonged activity and stability, but exhibit lower immunogenicity and antigenicity with significantly improved therapeutic functions (Gregoriadis et al. 2005). PSA has been used for the controlled release of drugs and as scaffolds in biomedical applications (Steinhaus et al. 2010; Colley et al. 2014). Polysialylation of therapeutic proteins and peptides such as insulin, asparaginase, antibody, and its fragment improved their stability and prolonged their half-life in body circulation, which may potentially improve the pharmacokinetics and pharmacodynamics of the drug molecules (Fernandes and Gregoriadis 2001; Jain et al. 2003; Constantinou et al. 2008, 2009). Antibody fragment (H17E2 Fab) with PSA conjugated had over a 5-fold increase in blood exposure and over a 3-fold higher tumor uptake compared to the unconjugated Fab (Constantinou et al. 2008). Polysialylation can increase the half-life of single-chain Fv antibody (scFv) 3.4- to 4.9-fold, resulting in a 10.6- to 15.2-fold increase in blood exposure (Constantinou et al. 2009). Polysialylation of insulin enhanced the therapeutic value of insulin by 2- to 3-fold (Jain et al. 2003). Polysialylation technology offers a promising strategy for the enhancement of the therapeutic value of peptide and protein drugs. Many polysialylated pharmaceutical proteins and peptides have undergone pre-clinical and clinical trials (Epenetos et al. 2002; Jain et al. 2003; Constantinou et al. 2008, 2009). Moreover, by treating PSA with enzymatic catalysis or chemical degradation, sialic acid has been derived from PSA which is subsequently used in pharmaceutical, food, and health-care industries. The resource availability of PSA is limited and the market price of PSA is as high as $200/g (Liu et al. 2010). In the last two decades, the feasibility of producing PSA from bacteria has focused on both industry and applied science. This review focuses on the metabolism and current state of biotechnological production of PSA by E. coli with a special emphasis on both stain engineering and process development strategies for the enhanced production of PSA.
Biosynthesis and metabolism of polysialic acid in E. coli
Many bacteria including N. meningitidis (Bhattacharjee et al. 1975), E. coli (Egan et al. 1977; Rodríguez-Aparicio et al. 1988), and M. hemolytica (previously Pasteurella haemolytica) (Puente-Polledo et al. 1998) produce PSA as their extracellular capsules consisting of linear homopolymers of N-acetyl-d-neuraminic acid (Neu5Ac) with α (2–8) or α (2–9) linkages or as linear copolymers with alternating α-(2–8)/α-(2–9)-linked polysialic acids (Ferrero and Aparicio 2010). These bacteria PSA as virulence factor mimic the mammalian PSA’s structure and can escape protection by the host’s immune system (Cress et al. 2014). E. coli is a model microorganism for investigating bacterial exopolysaccharide function and biosynthesis, especially in the case of PSA. The biosynthesis of PSA in E. coli involves the synthesis of sialic acid monomers, polymerization of PSA from sialic acid, and transportation of PSA to the cell surface (Ferrero and Aparicio 2010).
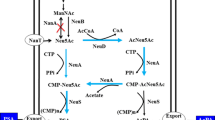
Neu5Ac is the precursor of PSA. The biosynthesis of Neu5Ac has been well characterized (Fig. 1). For the biosynthesis of sialic acid in bacteria, Neu5Ac aldolase (NanA) (Tao et al. 2011) or Neu5Ac synthase (NeuB) (Brody and Lundgren 2009; Ishikawa et al. 2010) is often used to catalyze the reaction from ManNAc to Neu5Ac. NeuB catalyzes the energy-dependent reaction of ManNAc and phosphoenolpyruvate (PEP) to form Neu5Ac (Lundgren and Boddy 2007; Boddy and Lundgren 2009). NanA catalyzes ManNAc and pyruvate to produce Neu5Ac (Li et al. 2008; Kang et al. 2012; Lin et al. 2013). Pyruvate is more readily available compared to PEP, making NanA a promising target in the engineered pathway for Neu5Ac synthesis. NanA was identified as the rate-controlling enzyme in the biosynthesis of Neu5Ac through ManNAc and pyruvate. With increased expression of NanA, a ninefold increase in Neu5Ac production was obtained (Lin et al. 2013).
Schematic representation of the polysialic acid metabolic pathway in E. coli. AGE GlcNAc 2-epimerase, NanA Neu5Ac aldolase, NanT sialic acid transporter, NanK ManNAc kinase, NanE ManNAc-6-P epimerase, Pgi, glucose-6-phosphate isomerase, GlmS l-glutamine:d-fructose-6-phosphate aminotransferase, GlmM phosphoglucosamine mutase, GlmU bifunctional UDP-N-acetylglucosamine pyrophosphorylase/glucosamine-1-phosphate N-acetyltransferase, NagA N-acetylglucosamine-6-phosphate deacetylase, NagB glucosamine-6-phosphate deaminase, NagK GlcNAc kinase, NagE GlcNAc-specific transporter, ManXYZ mannose transporter, NeuA CMP-Neu5Ac synthetase, NeuS α-Neu5Ac α-2,8-sialyltransferase, NeuB Neu5Ac synthase, NeuC UDP-GlcNAc-2-epimerase, NeuD Neu5Ac 7-O (or 9-O)-acetyltransferase
In E. coli, the kps gene cluster is involved in PSA biosynthesis, modification, and transport of the bacterial PSA chains (Fig. 2) (Barrett et al. 2002; Ferrero and Aparicio 2010). The kps cluster comprises three regions: (1) kpsFEDUCS, (2) neuDBACES, and (3) kpsMT The central neuDBACES region encodes the proteins involved in the biosynthesis, activation, and polymerization of Neu5Ac (Daines et al. 2000). NeuD, a Neu5Ac 7-O (or 9-O)-acetyltransferase, acetylates Neu5Ac residues at carbon position 7 or 9 (Steenbergen et al. 2006; Cress et al. 2014). NeuA, a bifunctional enzyme with both CMP-Neu5Ac synthetase (Vann et al. 1987; Zapata et al. 1989) and O-acetylesterase activity (Liu et al. 2004), catalyzes free sialic acid to cytidine 5′-monophosphate-sialic acid, thus generating the candidate donor form of sialic acid for all known sialyltransferases. NeuA also converts most of the CMP-O-acetyl-Neu5Ac to CMP-Neu5Ac before incorporation into the polymers, and only a small amount of CMP-O-acetyl-Neu5Ac is incorporated into the polymers (Song et al. 2011). NeuS, a α-Neu5Ac α-2,8-sialyltransferase, is responsible for polymerization of a homopolymer of α-2,8-linked Neu5Ac to form PSA (Steenbergen et al. 2006; Song et al. 2011; Ferrero and Aparicio 2010). The transcriptional antiterminator RfaH regulates PSA metabolism at a transcriptional level (Bailey et al. 1997). RfaH enhanced kps expression for the synthesis of polysialic acid capsule in E. coli K92 (Navasa et al. 2014).
The kps region 1 (kpsFEDUCS) and region 3 (kpsTM) genes participate in translocation of the polysaccharide across the periplasmic space and onto the cell surface (Willis and Whitfield 2013a; Colley et al. 2014). The E. coli kpsC and kspS genes in region 1 encode β-Kdo-transferases that are involved in the biosynthesis of a poly-Kdo linker at the reducing termini of PSA, which serves as a recognition signal for translocation (Willis and Whitfield 2013b). It was reported that KpsS could increase polysaccharide production by several folds (approximately 10 to 15 times) (Andreishcheva and Vann 2006). KpsT and KpsM are the components of an ABC transporter that enables PSA transfer from the cytoplasm to the cell surface. ABC transporter-dependent assembly also requires two other characteristic components: a member of the polysaccharide co-polymerase (PCP-3) family (KpsE) and an outer membrane polysaccharide (OPX) protein (KpsD) (Willis and Whitfield 2013a; Colley et al. 2014).
Process engineering for enhanced polysialic acid production in E. coli
Process engineering was used to improve PSA production through optimizing the medium and cultivation conditions. The production of PSA in E. coli was dependent on temperature, pH value, and composition of the culture mediums as well as the control strategy for fermentation cultivation (Table 1).
E. coli K92 synthesizes different capsular polysaccharides at different temperatures. At the cell growth temperature (37 °C), E. coli K92 synthesizes PSA, which belongs to the group 2 capsule responsible for bacterial virulence. At 20 °C, E. coli K92 accumulates colanic acid (CA), which provides protection against stressful conditions outside of the mammalian host (Navasa et al. 2009 and 2014). The production of PSA in P. haemolytica A2 is strictly regulated by the growth temperature, and above 40 °C, no production is detected. The activity of NanA, NeuA, and NeuS increased rapidly when bacteria grown at 43 °C were transferred to 37 °C; the enzymes decreased dramatically when the cells grown at 37 °C were transferred to 43 °C (Barrallo et al. 1999). PSA is optimally accumulated when bacteria are grown at 37 °C, and the accumulation is negligible when the microorganisms grow below 20 °C or above 40 °C (Barrallo et al. 1999; Navasa et al. 2009, 2011, 2013, 2014; Ferrero and Aparicio 2010).
The effects of medium compositions on PSA production were investigated. When bacteria grow in liquid medium, the PSA generated is largely broth liberated during cellular growth. The amount of PSA produced strongly depends on the applied carbon and nitrogen sources for the bacteria growth (Ferrero and Aparicio 2010). The optimal carbon and nitrogen source for PSA production was investigated in a batch culture (Revilla-Nuin et al. 1998, 2002; Ferrero and Aparicio 2010; Rode et al. 2008; Ezquerro-Sáenz et al. 2006). Sorbitol appears to induce PSA production (Honda et al. 1997) and has a significant effect on bacterial growth and PSA formation (Liu et al. 2010; Wu et al. 2010). The substrate xylose allowed E. coli K1 to produce more PSA (Wu et al. 2010). Using different nitrogen sources may result in a significant variation in the PSA level. When organic nitrogen sources were used, PSA production was higher than when inorganic nitrogen sources were used. Casamino acids were found to be the most effective nitrogen source for PSA production, and a high ammonia concentration inhibited PSA production (Honda et al. 1997). The catabolism of specific carbon and nitrogen sources causes a synergistic effect that facilitates the synthesis of the precursors (pyruvate and ManNAc) involved in the PSA biosynthesis pathway (Rodríguez-Aparicio et al. 1988; Rode et al. 2008).
PSA production and cell growth were closely related. The PSA concentration increased with a higher biomass in most cases. During PSA production by E. coli, the fermentation medium pH has a significant effect on the biosynthesis of the PSA. Liu et al. (2010) developed a novel strategy by controlling pH with ammonia water feeding coupled with sorbitol supplementation, and the resulting PSA level increased to 5.5 g/L. Wu et al. (2010) also used an ammonia water feeding strategy to control the pH at 6.4 in the bioreactor, and PSA production reached as high as 5.2 g/L. Zheng et al. (2013) developed a novel two-stage pH control fermentation process for production of high molecular weight PSA. The pH value was initially set at 6.4 to encourage cell growth and PSA production, and then the pH was set at 7.4 to promote the formation of higher molecular weight PSA. Their explanation for the impact of pH on the molecular weight of PSA was that cell growth and the activity of key PSA biosynthesis enzymes were affected by pH, thereby affecting its production rate. However, mildly acidic conditions affect the stability of the PSA molecules. A high pKa acts as a proton donor for general acid catalysis intramolecular self-cleavage of the glycosidic bonds of the internal sialic acid units adjacent to the carboxyl group. Another possible explanation is that rapid cell growth at the optimal conditions (37 °C and pH 6.4) would accelerate PSA biosynthesis. Thus, the molecular weight of PSA would be maintained at a relatively high level due to the PSA production rate significantly outpacing the degradation rate (Manzi et al. 1994; Zheng et al. 2013).
Compared with batch cultivation, fed-batch cultivations under a constant glucose supply were more efficient in PSA production. Chen and his colleagues compared fed-batch cultivation at a constant specific growth rate of 0.25/h with fed-batch cultivation at a constant glucose concentration of 50 mg/L. They found that both fed-batch cultivations provided higher yields than that of the batch cultivation and found that acetate formation was prevented. Moreover, the PSA yield on glucose resulted in higher PSA productivity. PSA formation was highly associated to the specific growth rate of the cells. The growth rate (0.32/h) in the fed-batch cultivation at a constant glucose concentration of 50 mg/L was 82 % higher than that of the fed-batch cultivation at a constant specific growth rate of 0.25/h, demonstrating a positive correlation between the specific growth rate and product formation (Chen et al. 2011).
Metabolic engineering for enhanced production of polysialic acid
With the advent of synthetic biology and metabolic engineering, many engineering tools have been developed for heterologous production of valuable chemicals. These tools create new opportunities for the efficient accumulation of PSA by designing and optimizing of the metabolic pathway in E. coli. N-Acetylneuraminic acid (Neu5Ac), the most ubiquitous species among the sialic acids, is the precursor of PSA biosynthesis and is widely distributed in the PSA of bacteria and mammals. Metabolic engineering was applied for enhanced Neu5Ac production. For the biosynthesis of sialic acid, the Neu5Ac synthase (NeuB) (Samain 2008; Brody and Lundgren 2009; Ishikawa et al. 2010) or the Neu5Ac aldolase (NanA) (Tao et al. 2011; Kang et al. 2012; Lin et al. 2013) is often employed to catalyze the reaction from ManNAc to Neu5Ac in the engineered strains.
Brody and Lundgren (2009) overexpressed glucosamine synthase (GlmS), NeuB, and UDPGlcNAc 2-epimerase (NeuC) from N. meningitidis in E. coli, and 1.5 g/L of Neu5Ac was produced from glucose after 98 h under shake-flask conditions. Samain (2008) reported that 39 g/L of Neu5Ac was produced through high cell density fermentation with glycerol as carbon source. Ishikawa et al. constructed a recombinant E. coli N18-14 strain by overexpressing genes of GlcNAc 2-epimerase (slr1975) and neuB resulting in a yield of 53 g/L of Neu5Ac (2.41 g/L/h) after 22 h (Ishikawa et al. 2010).
Neu5Ac aldolase (NanA), which catalyzes Neu5Ac synthesis using ManNAc and pyruvate as substrates, is used in Neu5Ac production (Lee et al. 2007; Wang et al. 2009; Hu et al. 2010; Tao et al. 2011; Kang et al. 2012; Lin et al. 2013). Lee et al. developed a Neu5Ac production system by coupling two E. coli strains that expressed GlcNAc 2-epimerase(AGE) and NanA individually; when co-cultured, 122.3 g/L of Neu5Ac was obtained with highly concentrated cells and highly concentrated substrates (1.2 M of GlcNAc and 1.2 M of pyruvate). Tao et al. (2011) constructed E. coli strains overexpressing slr1975 and nanA for Neu5Ac production, and Neu5Ac accumulated to 59 g/L after 36 h (1.64 g/L/h). Lin et al. (2013) engineered E. coli by assembling a two-step heterologous pathway consisting of AGE and NanA, resulting in a yield of 74.2 g/L Neu5Ac and a productivity of 6.2 g/L/h.
The nan operon (Fig. 2), which encodes the proteins for sialic acid catabolism, is found in hundreds of bacterial species (Vimr et al. 2004; Almagro-Moreno and Boyd 2009). The products of sialic acid catabolism are GlcNAc and pyruvate, which are the primary metabolism molecules (Almagro-Moreno and Boyd 2009; Li and Chen 2012).The removal of the nanA and nanT genes abolishes sialic acid catabolism and increases Neu5Ac biosynthesis using the NeuB pathway (Brody and Lundgren 2009). Knockout of both the nanK and nanA genes improved the sialylation efficiency by preventing ManNAc and Neu5Ac from being diverted from the biosynthesis pathway (Fierfort and Samain 2008). Knockout of the nanTEK genes blocked Neu5Ac uptake and the competing pathway, which forced the reactions toward the synthetic direction as the final product was secreted outside of the cells and enhanced the Neu5Ac production by 3-fold (Lin et al. 2013). NanA is the rate-controlling enzyme in the biosynthesis of Neu5Ac through ManNAc and pyruvate. With the increased expression of NanA, a ninefold increase in Neu5Ac production was obtained (Lin et al. 2013). By blocking Neu5Ac uptake and degradation and abolishing the ManNAc catabolic pathway by removing nanT and nanK, respectively, the efficiency of the whole-cell biocatalyst for Neu5Ac production was improved (Brody and Lundgren 2009; Lin et al. 2013).
For PSA biosynthesis, NeuD, NeuA, and NeuS are key components of the biosynthetic pathway (Fig. 1) (Steenbergen et al. 2006; Song et al. 2011; Ferrero and Aparicio 2010). To investigate the effects of the key enzymes of the PSA synthetic pathway on PSA production, NeuD, NeuA, and NeuS were overexpressed separately or in combination (Chen et al. 2015). The strain overexpressing NeuD produced threefold more PSA than that of the wild-type strain, while the strains overexpressing NeuA or/and NeuS only slightly increased PSA production. The results showed that NeuD played an important role in the biosynthesis of PSA. The strain harboring pDB1S-DA, which co-expressed NeuD and NeuA, enhanced the PSA biosynthetic pathway and improved PSA production significantly. Neu5Ac is the precursor of PSA, and blocking the Neu5Ac catabolic pathway by deleting NanA resulted in enhanced production of PSA. By overexpressing NeuD and NeuA in the nanA-knockout strain, the production of PSA increased by 85 % compared to the original strain, and 16.15 g/L PSA was obtained (Chen et al. 2015).
Purification of polysialic acid in E. coli
Studies on the isolation of PSA from fermentation broth have been reported. Rode et al. (2008) developed a process for purification of PSA. The major PSA fraction could be successfully isolated from the concentrated supernatant by consecutive precipitation steps with acetone, cetavlon, and ethanol. Final polishing of the purified fraction was achieved using size exclusion chromatography on a Sephacryl S-300 column. The overall amount of the isolated product was greater than 20 % of the total PSA that was produced (Rode et al. 2008).
Bacterial PSA is negatively charged due to the high content of carboxyl groups, which could be completely neutralized by the typical positively charged cationic surfactants, such as cetyl pyridinium chloride (CPC). Liu et al. (2010) developed a purification method for PSA and involved isolation from the broth by ethanol precipitation, filtration with perlite as a filter aid, CPC precipitation, and lyophilization. The final PSA product had 98.1 ± 1.6 % purity at a 56.1 ± 1.7 % recovery rate.
Future perspectives
In recent years, PSA has attracted considerable interest due to its important biological functions and has been a valuable resource in both the medical and biotechnological fields. Many research studies focus on the link between quality and quantity of PSA and their therapeutic applications. A vaccine using bacterial PSA with the α 2,8-polysialic structure is weakly immunogenic because it is identical to the polysialic acid expressed in mammals. As a consequence, modified PSA has been used in vaccine development to protect individuals from invasive meningitis (Tan et al. 2010). PSA as an anti-adhesive biological molecule is dependent on chain length (Colley et al. 2014). Therefore, the use of more uniform chain length PSA has a tremendous potential to improve the production of these therapeutic reagents, and may enable potential PSA strategies for nervous system tissue and other tissue repair.
For biotechnological production, E. coli strains have been process engineered and metabolically engineered for the efficient production of PSA. Metabolic engineering strategies have been successfully applied for improving Neu5Ac and PSA production. The combination of metabolic engineering and process engineering strategies, including enhancing the precursor (Neu5Ac) supply, promoting capsule polysaccharide exportation, and optimizing the fermentation process, will lead to new approaches to further improve the PSA production. These strategies will make the bioprocess a promising cost-effective resource for PSA production with not only higher titers, yields, and productivity but also regulated PSA chain lengths.
References
Almagro-Moreno S, Boyd EF (2009) Insights into the evolution of sialic acid catabolism among bacteria. BMC Evol Biol 9(1):118
Andreishcheva EN, Vann WF (2006) Gene products required for de novo synthesis of polysialic acid in Escherichia coli K1. J Bacteriol 188:1786–1797
Bailey MJ, Hughes C, Koronakis V (1997) RfaH and the ops element, components of a novel system controlling bacteria transcription elongation. Mol Microbiol 26:845–851
Barrallo S, Reglero A, Revilla-Nuin B, Martínez-Blanco H, Rodríguez-Aparicio LB, Ferrero MA (1999) Regulation of capsular polysialic acid biosynthesis by temperature in Pasteurella haemolytica A2. FEBS Lett 445(2–3):325–328
Barrett B, Ebah L, Roberts IS (2002) Genomic structure of capsular determinants. Curr Top Microbiol Immunol 264:137–155
Bhattacharjee AK, Jennings HJ, Kenny CP, Martin A, Smith IC (1975) Structural determination of the sialic acid polysaccharide antigens of Neisseria meningitides serogroups B and C with carbon 13 nuclear magnetic. J Biol Chem 250:1926–1932
Boddy CN, Lundgren BR (2009) Metabolically engineered Escherichia coli for enhanced production of sialic acid. (US Patent number: 8722365)
Chen F, Tao Y, Jin C, Xu Y, Lin BX (2015) Enhanced production of polysialic acid by metabolic engineering of Escherichia coli. Appl Microbiol Biotechnol 99(6):2603–2611
Chen R, John J, Rode B, Hitzmann B, Gerardy-Schahn R, Kasper C, Scheper T (2011) Comparison of polysialic acid production in Escherichia coli K1 during batch cultivation and fed-batch cultivationapplying two different control strategies. J Biotechnol 154(4):222–229
Colley KJ, Kitajima K, Sato C (2014) Polysialic acid: biosynthesis, novel functions and applications. Crit Rev Biochem Mol Biol 49(6):498–532
Constantinou A, Epenetos AA, Hreczuk-Hirst D, Deonarain MP (2008) Modulation of antibody pharmacokinetics by chemical polysialylation. Bioconjug Chem 19:643–650
Constantinou A, Epenetos AA, Hreczuk-Hirst D, Jain S, Wright M, Chester K, Deonarain M (2009) Site-specific polysialylation of an antitumor single-chain Fv fragment. Bioconjug Chem 20:924–931
Cress BF, Englaender JA, He W, Kasper D, Linhardt RJ, Koffas MA (2014) Masquerading microbial pathogens: capsular polysaccharides mimic host-tissue molecules. FEMS Microbiol Rev 38:660–697
Daines DA, Silver RP (2000) Evidence for multimerization of Neu proteins involved in polysialic acid synthesis in Escherichia coli K1 using improved LexA-based vectors. J Bacteriol 182:5267–5270
Egan W, Liu TY, Dorow D, Cohen JS, Robbins JD, Gotschlich EC, Robbins JB (1977) Structural studies on the sialic acid polysaccharide antigen of Escherichia coli strain Bos-12. Biochemistry 16:16989–16994
Epenetos AA, Hreczuk-Hirst DH, McCormack B, Gregoriadis G (2002) Polysialylated proteins: a potential role in cancer therapy. Clin Pharm 21:2186
Ezquerro-Sáenz C, Ferrero MA, Revilla-Nuín B, López-Velasco FF, Martínez-Blanco H, Rodríguez-Aparicio LB (2006) Transport of N-acetyl-D-galactosamine in Escherichia coli K92: effect of acetyl-amino sugar metabolism and polysialic acid production. Biochimie 88:95–102
Falconer RA, Errington RJ, Shnyder SD, Smith PJ, Patterson LH (2012) Polysialyltransferase: a new target in metastatic cancer. Curr Cancer Drug Targets 12:925–939
Fernandes AI, Gregoriadis G (2001) The effect of polysialylation on the immunogenicity and antigenicity of asparaginase: implication in its pharmacokinetics. Int J Pharm 217:215–224
Ferrero MA, Aparicio LR (2010) Biosynthesis and production of polysialic acids in bacteria. Appl Microbiol Biotechnol 86(6):1621–1635
Fierfort N, Samain E (2008) Genetic engineering of Escherichia coli or the economical production of sialylated oligosaccharides. J Biotechnol 134(3):261–265
Fukuda M (1996) Possible roles of tumor-associated carbohydrate antigens. Cancer Res 56:2237–2244
Gregoriadis G, Jain S, Papaioannou I, Laing P (2005) Improving the therapeutic efficacy of peptides and proteins: a role for polysialic acids. Int J Pharm 300:125–130
Heimburg-Molinaro J, Lum M, Vijay G, Jain M, Almogren A, Rittenhouse-Olson K (2011) Cancer vaccines and carbohydrate epitopes. Vaccine 29(48):8802–8826
Honda H, Nakazeko T, Ogiso K, Kawase Y, Aoki N, Kawase M, Kobayashi T (1997) Colominic acid production from Escherichia coli in a fed-batch culture under the control of ammonium ions using an FIA system. J Ferment Bioeng 83:59–63
Hu S, Chen J, Yang Z, Shao L, Bai H, Luo J, Jiang W, Yang Y (2010) Coupled bioconversion for preparation of N-acetyl-D-neuraminic acid using immobilized N-acetyl-D-glucosamine-2-epimerase and N-acetyl-D-neuraminic acid lyase. Appl Microbiol Biotechnol 85(5):1383–1391
Ishikawa M, Koizumi S (2010) Microbial production of N-acetyneuraminic acid by genetically engineered Escherichia coli. Carbohydr Res 345(18):2605–2609
Jain S, Hreczuk-Hirst DH, McCormack B, Mital M, Epenetos A, Laing P, Gregoriadis G (2003) Polysialylated insulin: synthesis, characterization and biological activity in vivo. Biochimica et Biophysica Acta (BBA)-General Subjects 1622(1):42–49
Kang J, Gu P, Wang Y, Li Y, Yang F, Wang Q, Qi Q (2012) Engineering of an N-acetylneuraminic acid synthetic pathway in Escherichia coli. Metab Eng 14(6):623–629
Krug LM, Ragupathi G, Hood C, George C, Hong F, Shen R, Abrey L, Jennings HJ, Kris MG, Livingston PO (2012) Immunization with N-propionyl polysialic acid-KLH conjugate in patients with small cell lung cancer is safe and induces IgM antibodies reactive with SCLC cells and bactericidal against group B meningococci. Cancer Immunol Immunother 61:9–18
Lee YC, Chien HC, Hsu WH (2007) Production of N-acetyl-D-neuraminic acid by recombinant whole cells expressing Anabaena sp. CH1 N-acetyl-D-glucosamine 2-epimerase and Escherichia coli N-acetyl-D-neuraminic acid lyase. J Biotechnol 129(3):453–460
Li Y, Yu H, Cao H, Lau K, Muthana S, Tiwari VK, Son B, Chen X (2008) Pasteurella multocida sialic acid aldolase: a promising biocatalyst. Appl Microbiol Biotechnol 79(6):963–970
Li Y, Chen X (2012) Sialic acid metabolism and sialyltransferases: natural functions and applications. Appl Microbiol Biotechnol 94:887–905
Lin BX, Zhang ZJ, Liu WF, Dong ZY, Tao Y (2013) Enhanced production of N-acetyl-D-neuraminic acid by multi-approach whole-cell biocatalyst. Appl Microbiol Biotechnol 97(11):4775–4784
Liu GC, Jin CS, Jin C (2004) CMP-N-acetylneuraminic acid synthetase from Escherichia coli K1 is a bifunctional enzyme: identification of minimal catalytic domain for synthetase activity and novel functional domain for platelet-activating factor acetylhydrolase activity. J Biol Chem 279:17738–17749
Liu JL, Zhan XB, Wu JR, Lin CC, Yu DF (2010) An efficient and large scale preparation process for polysialic acid by Escherichia coli CCTCC M208088. Biochem Eng J 53:97–103
Lundgren BR, Boddy CN (2007) Sialic acid and N-acyl sialic acid analog production by fermentation of metabolically and genetically engineered Escherichia coli. Org Biomol Chem 5(12):1903–1909
Maarouf AE, Petridis AK, Rutishauser U (2006) Use of polysialic acid in repair of the central nervous system. Proc Natl Acad Sci 103:16989–16994
Manzi AE, Higa HH, Diaz S, Varki A (1994) Intramolecular self-cleavage of polysialic acid. J Biol Chem 269:23617–23624
Muhlenhoff M, Eckhardt M, Schahn RG (1998) Polysialic acid: three-dimensional structure, biosynthesis and function. Curr Opin Struct Biol 8:558–564
Navasa N, Rodríguez-Aparicio LB, Martínez-Blanco H, Arcos M, Ferrero MA (2009) Temperature has reciprocal effects on colanic acid and polysialic acid biosynthesis in E. coli K92. Appl Microbiol Biotechnol 82(4):721–729
Navasa N, Rodríguez-Aparicio LB, Ferrero MA, Moteagudo-Mera A, Martínez-Blanco H (2011) Growth temperature regulation of some genes that define the superficial capsular carbohydrate composition of Escherichia coli K92. FEMS Microbiol Lett 320:135–141
Navasa N, Rodríguez-Aparicio LB, Ferrero MA, Monteagudo-Mera A, Martínez-Blanco H (2013) Polysialic acid and colania acids metabolism in Escherichia coli K92 is regulated by RcsA and RcsB. Biosci Rep 33(3):405–415
Navasa N, Rodríguez-Aparicio LB, Ferrero MA, Monteagudo-Mera A, Martínez-Blanco H (2014) Transcriptional control of RfaH on polysialic and colanic acid synthesis by Escherichia coli K92. FEBS Lett 588(6):922–928
Puente-Polledo L, Reglero A, González-Clemente C, Rodríguez-Aparicio LB, Ferrero MA (1998) Biochemical conditions for the production of polysialic acid by Pasteurella haemolytica A2. Glycoconj J 15(9):855–861
Revilla-NuiB R-ALB, FerreroMA RA (1998) Regulation of capsular polysialic acid biosynthesis by N-acetyl-D-mannosamine, an intermediate of sialic acid metabolism. FEBS Lett 426(2):191–195
Revilla-Nuin B, Reglero A, Martínez-Blanco H, Bravo IG, Ferrero MA, Rodríguez-Aparicio LB (2002) Transport of N-acetyl-D-mannosamine and N-acetyl-D-glucosamine in Escherichia coli K1: effect on capsular polysialic acid production. FEBS Lett 511(1–3):97–101
Rodríguez-Aparicio LB, Reglero A, Oritiz AI, Luengo JM (1988) Effect of physical and chemical conditions on the production of colominic acid by Escherichia coli in a defined medium. Appl Microbiol Biotechnol 27:474–483
Rode B, Endres C, Ran C, Stahl F, Beutel S, Kasper C, Galuska S, Geyer R, Mühlenhoff M, Gerardy-Schahn R, Scheper T (2008) Large-scale production and homogenous purification of long chain polysialic acids from E. coli K1. J Biotechnol 135(2):202–209
Samain, E (2008) High yield production of sialic acid (Neu5Ac) by fermentation. (US Patent number: 20110165626A1)
Sato C, Kitajima K (2013a) Impact of structural aberrancy of polysialic acid and its synthetic enzyme ST8SIA2 in schizophrenia. Front Cell Neurosci 7:61
Sato C, Kitajima K (2013b) Disialic, oligosialic and polysialic acids: distribution, functions and related disease. J Biochem 154:115–136
Schnaar RL, Gerardy-Schahn R, Hildebrandt H (2014) Sialic acids in the brain: gangliosides and polysialic acid in nervous system development, stability, disease, and regeneration. Physiol Rev 94:461–518
Simon P, Bäumner S, Busch O, Röhrich R, Kaese M, Richterich P, Wehrend A, Müller K, Gerardy-Schahn R, Mühlenhoff M, Geyer H, Geyer R, Middendorff R, Galuska SP (2013) Polysialic acid is present in mammalian semen as a post-translational modification of the neural cell adhesion molecule NCAM and the polysialyltransferase ST8SiaII. J Biol Chem 288:18825–18833
Song LL, Zhou H, Cai XH, Li CY, Liang JN, Jin C (2011) NeuA O-acetylesterase activity is specific for CMP-activated O-acetyl sialic acid in Streptococcus suis serotype 2. Biochem Biophys Res Commun 410:212–217
Stamatos NM, Zhang L, Jokilammi A, Finne J, Chen WH, El-Maarouf A, Cross AS, Hankey KG (2014) Changes in polysialic acid expression on myeloid cells during differentiation and recruitment to sites of inflammation: role in phagocytosis. Glycobiology 24:86
Steenbergen SM, Lee YC, Vann WF, Vionnet J, Wright LF, Vimr ER (2006) Separate pathway for O-acetylation of polymeric and monomeric sialic acid and identification of sialyl O-acetylesterase in Escherichia coli K1. J Bacteriol 188:6195–6620
Steinhaus S, Stark Y, Bruns S, Haile Y, Scheper T, Grothe C, Behrens P (2010) Polysialic acid immobilized on silanized glass surfaces: a test case for its use as a biomaterial for nerve regeneration. J Mater Sci Mater Med 21:1371–1378
Tao F, Zhang Y, Ma C, Xu P (2011) One-pot bio-synthesis: N-acetylneuraminic acid production by a powerful engineered whole-cell catalyst. Sci Rep 1:1–7
Tan LK, Carlone GM, Borrow R (2010) Advances in the development of vaccines against Neisseria meningitidis. N Engl J Med 362:1511–1520
Tsuchiya A, Lu WY, Weinhold B, Boulter L, Stutchfield BM, Williams MJ, Guest RV, Minnis-Lyons SE, AC MK, Schwarzer D, Ichida T, Nomoto M, Aoyagi Y, Gerardy-Schahn R, Forbes SJ (2014) PolySia-NCAM modulates the formation of ductular reactions in liver injury. Hepatology 60:1727–1740
Ulm C, Saffarzadeh M, Mahavadi P, Müller S, Prem G, Saboor F, Simon P, Middendorff R, Geyer H, Henneke I, Bayer N, Rinné S, Lütteke T, Böttcher-Friebertshäuser E, Gerardy-Schahn R, Schwarzer D, Mühlenhoff M, Preissner KT, Günther A, Geyer R, Galuska SP (2013) Soluble polysialylated NCAM: a novel player of the innate immune system in the lung. Cell Mol Life Sci 70:3695–3708
Vann WF, Silver RP, Abeijon C, Chang K, Aaronson W, Sutton A, Finn CW, Lindner W, Kotsatos M (1987) Purification, properties, and genetic location of Escherichia coli cytidine 5′-monophosphate N-acetyneuraminic acid synthetase. J Biol Chem 262:17556–17562
Vimr ER, Kalivoda KA, Deszo EL, Steenbergen SM (2004) Diversity of microbial sialic acid metabolism. Microbiol Mol Biol Rev 68:132–153
Wang TH, Chen YY, Pan HH, Wang FP, Cheng CH, Lee WC (2009) Production of N-acetyl-D-neuraminic acid using two sequential enzymes overexpressed as double-tagged fusion proteins. BMC Biotechnol 9:63
Willis LM, Whitfield C (2013a) Structure, biosynthesis, and function of bacterial capsular polysaccharides synthesized by ABC transporter dependent pathways. Carbohydr Res 378:35–44
Willis LM, Whitfield C (2013b) KpsC and KpsS are retaining 3-deoxy-D-manno-oct-2-ulosonic acid (Kdo) transferases involved in synthesis of bacterial capsules. Proc Natl Acad Sci U S A 110:20753–20758
Wu JR, Liu JL, Zhan XB, Lin CC, Zhao H (2010) Enhancement of polysialic acid yield by reducing initial phosphate and feeding ammonia water to Escherichia coli CCTCC M208088. Biotechnol Bioproc Eng 15:657–663
Zhan XB, Zhu L, Wu JR, Zhen ZY, Jia W (2002) Production of polysialic acid from fed-batch fermentation with pH control. Biochem Eng J 11:201–204
Zapata G, VannWF AW, Lewis MS, Moos M (1989) Sequence of the cloned Escherichia coli K1 CMP-N-acetylneuraminic acid synthetase gene. J Biol Chem 264:14769–14774
Zheng ZY, Wang SZ, Li GS, Zhan XB, Lin CC, Wu JR (2013) A new polysialic acid production process based on dual-stage pH control and fed-batch fermentation for higher yield and resulting high molecular weight product. Appl Microbiol Biotechnol 97:2405–2412
Acknowledgments
This work was supported by the Ministry of Science and Technology of China (grant 2013CB734003) and Key Research Program of the Chinese Academy of Sciences (grant no. KGZD-EW-606).
Conflict of interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
Bai-Xue Lin and Yu Qiao contributed equally to the paper.
Rights and permissions
About this article
Cite this article
Lin, BX., Qiao, Y., Shi, B. et al. Polysialic acid biosynthesis and production in Escherichia coli: current state and perspectives. Appl Microbiol Biotechnol 100, 1–8 (2016). https://doi.org/10.1007/s00253-015-7019-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-7019-x