Abstract
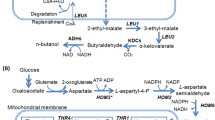
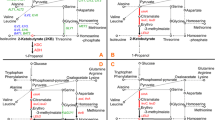
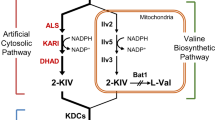
Saccharomyces cerevisiae naturally produces small amounts of isobutanol and 3-methyl-1-butanol via Ehrlich pathway from the catabolism of valine and leucine, respectively. In this study, we engineered CEN.PK2-1C, a leucine auxotrophic strain having a LEU2 gene mutation, for the production of isobutanol and 3-methyl-1-butanol. First, ALD6 encoding aldehyde dehydrogenase and BAT1 involved in valine synthesis were deleted to eliminate competing pathways. We also increased transcription of endogenous genes in the valine and leucine biosynthetic pathways by expressing Leu3Δ601, a constitutively active form of Leu3 transcriptional activator. For the production of isobutanol, genes involved in isobutanol production (ILV2, ILV3, ILV5, ARO10, and ADH2) were additionally overexpressed in ald6Δbat1Δ strain expressing LEU3Δ601, resulting in 376.9 mg/L isobutanol production from 100 g/L glucose. To increase 3-methyl-1-butanol production, leucine biosynthetic genes were additionally overexpressed in the final isobutanol-production strain. The resulting strain overexpressing LEU2 and LEU4 D578Y, a feedback inhibition-insensitive mutant of LEU4, showed a 34-fold increase in 3-methyl-1-butanol synthesis compared with CEN.PK2-1C control strain, producing 765.7 mg/L 3-methyl-1-butanol.





Similar content being viewed by others
References
Atsumi S, Hanai T, Liao JC (2008) Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature 451:86–89
Atsumi S, Wu TY, Eckl EM, Hawkins SD, Buelter T, Liao JC (2010) Engineering the isobutanol biosynthetic pathway in Escherichia coli by comparison of three aldehyde reductase/alcohol dehydrogenase genes. Appl Microbiol Biotechnol 85:651–657
Avalos JL, Fink GR, Stephanopoulos G (2013) Compartmentalization of metabolic pathways in yeast mitochondria improves the production of branched-chain alcohols. Nat Biotechnol 31:335–341
Baichwal VR, Cunningham TS, Gatzek PR, Kohlhaw GB (1983) Leucine biosynthesis in yeast: identification of two genes (LEU4, LEU5) that affect alpha-Isopropylmalate synthase activity and evidence that LEU1 and LEU2 gene expression is controlled by alpha-Isopropylmalate and the product of a regulatory gene. Curr Genet 7:369–377
Blombach B, Eikmanns BJ (2011) Current knowledge on isobutanol production with Escherichia coli, Bacillus subtilis and Corynebacterium glutamicum. Bioeng Bugs 2:346–350
Boer VM, Daran JM, Almering MJ, de Winde JH, Pronk JT (2005) Contribution of the Saccharomyces cerevisiae transcriptional regulator Leu3p to physiology and gene expression in nitrogen- and carbon-limited chemostat cultures. FEMS Yeast Res 5:885–897
Boles E, de Jong-Gubbels P, Pronk JT (1998) Identification and characterization of MAE1, the Saccharomyces cerevisiae structural gene encoding mitochondrial malic enzyme. J Bacteriol 180:2875–2882
Brat D, Weber C, Lorenzen W, Bode HB, Boles E (2012) Cytosolic re-localization and optimization of valine synthesis and catabolism enables increased isobutanol production with the yeast Saccharomyces cerevisiae. Biotechnol Biofuels 5:65
Buijs NA, Siewers V, Nielsen J (2013) Advanced biofuel production by the yeast Saccharomyces cerevisiae. Curr Opin Chem Biol 17:480–488
Chen X, Nielsen KF, Borodina I, Kielland-Brandt MC, Karhumaa K (2011) Increased isobutanol production in Saccharomyces cerevisiae by overexpression of genes in valine metabolism. Biotechnol Biofuels 4:21
Connor MR, Liao JC (2008) Engineering of an Escherichia coli strain for the production of 3-methyl-1-butanol. Appl Environ Microbiol 74:5769–5775
Connor MR, Liao JC (2009) Microbial production of advanced transportation fuels in non-natural hosts. Curr Opin Biotechnol 20:307–315
Dickinson JR, Harrison SJ, Hewlins MJ (1998) An investigation of the metabolism of valine to isobutyl alcohol in Saccharomyces cerevisiae. J Biol Chem 273:25751–25756
Eden A, Simchen G, Benvenisty N (1996) Two yeast homologs of ECA39, a target for c-Myc regulation, code for cytosolic and mitochondrial branched-chain amino acid aminotransferases. J Biol Chem 271:20242–20245
Friden P, Schimmel P (1987) LEU3 of Saccharomyces cerevisiae encodes a factor for control of RNA levels of a group of leucine-specific genes. Mol Cell Biol 7:2708–2717
Friden P, Schimmel P (1988) LEU3 of Saccharomyces cerevisiae activates multiple genes for branched-chain amino acid biosynthesis by binding to a common decanucleotide core sequence. Mol Cell Biol 8:2690–2697
Friden P, Reynolds C, Schimmel P (1989) A large internal deletion converts yeast LEU3 to a constitutive transcriptional activator. Mol Cell Biol 9:4056–4060
Gueldener U, Heinisch J, Koehler GJ, Voss D, Hegemann JH (2002) A second set of loxP marker cassettes for Cre-mediated multiple gene knockouts in budding yeast. Nucleic Acids Res 30:e23
Hazelwood LA, Daran JM, van Maris AJ, Pronk JT, Dickinson JR (2008) The Ehrlich pathway for fusel alcohol production: a century of research on Saccharomyces cerevisiae metabolism. Appl Environ Microbiol 74:2259–2266
Hohmann S (2002) Osmotic adaptation in yeast–control of the yeast osmolyte system. Int Rev Cytol 215:149–187
Horton CE, Huang KX, Bennett GN, Rudolph FB (2003) Heterologous expression of the Saccharomyces cerevisiae alcohol acetyltransferase genes in Clostridium acetobutylicum and Escherichia coli for the production of isoamyl acetate. J Ind Microbiol Biotechnol 30:427–432
Ihrig J, Hausmann A, Hain A, Richter N, Hamza I, Lill R, Muhlenhoff U (2010) Iron regulation through the back door: iron-dependent metabolite levels contribute to transcriptional adaptation to iron deprivation in Saccharomyces cerevisiae. Eukaryot Cell 9:460–471
Kispal G, Steiner H, Court DA, Rolinski B, Lill R (1996) Mitochondrial and cytosolic branched-chain amino acid transaminases from yeast, homologs of the myc oncogene-regulated Eca39 protein. J Biol Chem 271:24458–24464
Kohlhaw GB (2003) Leucine biosynthesis in fungi: entering metabolism through the back door. Microbiol Mol Biol Rev 67:1–15
Kondo T, Tezuka H, Ishii J, Matsuda F, Ogino C, Kondo A (2012) Genetic engineering to enhance the Ehrlich pathway and alter carbon flux for increased isobutanol production from glucose by Saccharomyces cerevisiae. J Biotechnol 159:32–37
Lee WH, Seo SO, Bae YH, Nan H, Jin YS, Seo JH (2012) Isobutanol production in engineered Saccharomyces cerevisiae by overexpression of 2-ketoisovalerate decarboxylase and valine biosynthetic enzymes. Bioprocess Biosyst Eng 35:1467–1475
Lilly M, Bauer FF, Styger G, Lambrechts MG, Pretorius IS (2006) The effect of increased branched-chain amino acid transaminase activity in yeast on the production of higher alcohols and on the flavour profiles of wine and distillates. FEMS Yeast Res 6:726–743
Matsuda F, Ishii J, Kondo T, Ida K, Tezuka H, Kondo A (2013) Increased isobutanol production in Saccharomyces cerevisiae by eliminating competing pathways and resolving cofactor imbalance. Microbiol Cell Fact 12:119
Mumberg D, Muller R, Funk M (1995) Yeast vectors for the controlled expression of heterologous proteins in different genetic backgrounds. Gene 156:119–122
Nevoigt E (2008) Progress in metabolic engineering of Saccharomyces cerevisiae. Microbiol Mol Biol Rev 72:379–412
Oba T, Nomiyama S, Hirakawa H, Tashiro K, Kuhara S (2005) Asp578 in LEU4p is one of the key residues for leucine feedback inhibition release in sake yeast. Biosci Biotechnol Biochem 69:1270–1273
Porro D, Gasser B, Fossati T, Maurer M, Branduardi P, Sauer M, Mattanovich D (2011) Production of recombinant proteins and metabolites in yeasts: when are these systems better than bacterial production systems? Appl Microbiol Biotechnol 89:939–948
Romagnoli G, Luttik MA, Kotter P, Pronk JT, Daran JM (2012) Substrate specificity of thiamine pyrophosphate-dependent 2-oxo-acid decarboxylases in Saccharomyces cerevisiae. Appl Environ Microbiol 78:7538–7548
Ryan ED, Kohlhaw GB (1974) Subcellular localization of isoleucine-valine biosynthetic enzymes in yeast. J Bacteriol 120:631–637
Schoondermark-Stolk SA, Tabernero M, Chapman J, Ter Schure EG, Verrips CT, Verkleij AJ, Boonstra J (2005) Bat2p is essential in Saccharomyces cerevisiae for fusel alcohol production on the non-fermentable carbon source ethanol. FEMS Yeast Res 5:757–766
Todd RB, Andrianopoulos A (1997) Evolution of a fungal regulatory gene family: the Zn(II)2Cys6 binuclear cluster DNA binding motif. Fungal Genet Biol 21:388–405
Vadali RV, Horton CE, Rudolph FB, Bennett GN, San KY (2004) Production of isoamyl acetate in ackA-pta and/or ldh mutants of Escherichia coli with overexpression of yeast ATF2. Appl Microbiol Biotechnol 63:698–704
Weber C, Farwick A, Benisch F, Brat D, Dietz H, Subtil T, Boles E (2010) Trends and challenges in the microbial production of lignocellulosic bioalcohol fuels. Appl Microbiol Biotechnol 87:1303–1315
Yamamoto S, Suda M, Niimi S, Inui M, Yukawa H (2013) Strain optimization for efficient isobutanol production using Corynebacterium glutamicum under oxygen deprivation. Biotechnol Bioeng 110:2938–2948
Zhou K, Brisco PR, Hinkkanen AE, Kohlhaw GB (1987) Structure of yeast regulatory gene LEU3 and evidence that LEU3 itself is under general amino acid control. Nucleic Acids Res 15:5261–5273
Zhou KM, Bai YL, Kohlhaw GB (1990) Yeast regulatory protein LEU3: a structure-function analysis. Nucleic Acids Res 18:291–298
Acknowledgments
This work was supported by the National Research Foundation (NRF) of Korea Grant funded by the Korean Government (2012-R1A1A-3011963).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Park, SH., Kim, S. & Hahn, JS. Metabolic engineering of Saccharomyces cerevisiae for the production of isobutanol and 3-methyl-1-butanol. Appl Microbiol Biotechnol 98, 9139–9147 (2014). https://doi.org/10.1007/s00253-014-6081-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-6081-0